Bifurcation analysis of logical models in systems biology
Sam Pastva


Modelling gene interactions
Gene X @DNA
Gene X @mRNA
Protein X
Gene Y @DNA
Gene Y @mRNA
Protein Y
\(\oplus\)
\(\ominus\)
Gene X
Gene Y
Gene Regulatory Network (GRN)
Gene X
Gene A
Gene B
Gene C
Do we need both A and C, or just one? Does B interact with A and C equally or not?
A
C
B
Boolean Network (BN)
000
100
111
011
001
Synchronous update: all updates happen at the same time.
000
100
110
101
001
111
011
Asynchronous update: better captures unknown process time-scales.
What "outcomes" are possible in our model? (Attractors - bottom SCCs)
- Single state (fixed-point attractor)
- Larger region (complex attractor)
- cycle (oscillation)
- other (disorder)
110
110
110
110
110
110
110
110
110
Assumption: chemistry is fair. Only terminal states/components can be attractors.
Different attractors can represent different biological functionality (phenotype).
Addressing model uncertainty
- Logical parameters (network inputs)
- Functional parameters: (partially) unknown dynamics
Gene X
Gene Y
Input A
Exponentially many "outcomes" for logical parameters, double exponential for functional parameters.
Benes, N., Brim, L., Pastva, S., Polácek, J., & Šafránek, D. (2019). Formal Analysis of Qualitative Long-Term Behaviour in Parametrised Boolean Networks. ICFEM.
Depending on system's parameters, which outcomes are "similar" and which are "different"?
Bifurcation analysis:
Bifurcation theory of continuous dynamical systems
Cont. dynamical system: (simplified)
A set of deterministic trajectories in \(\mathbb{R}^n\)
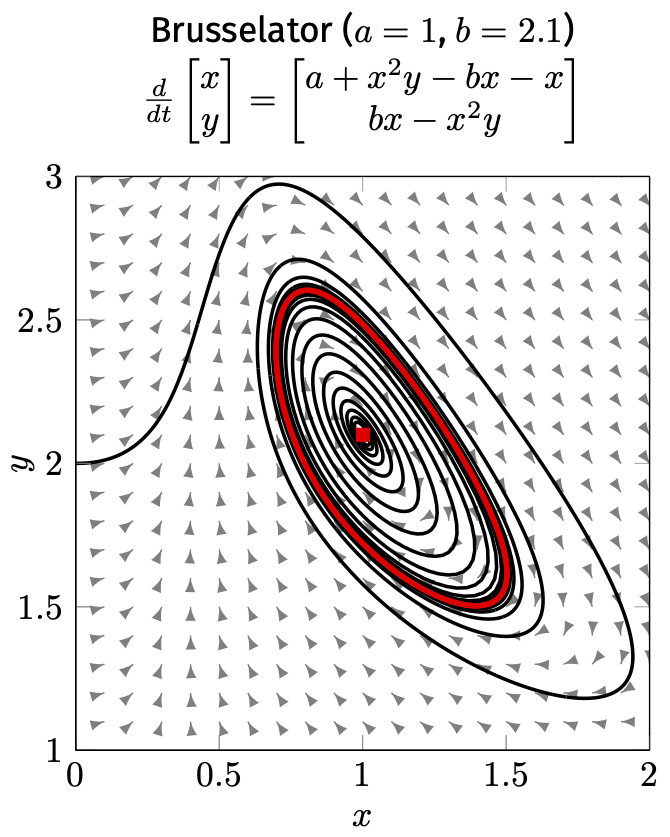
"Similar" vs. "different":
Topological equivalence (continuous transformation of trajectories)
Topological equivalence recognises similar patterns (e.g. attractors, repellers) that we saw in the discrete world:

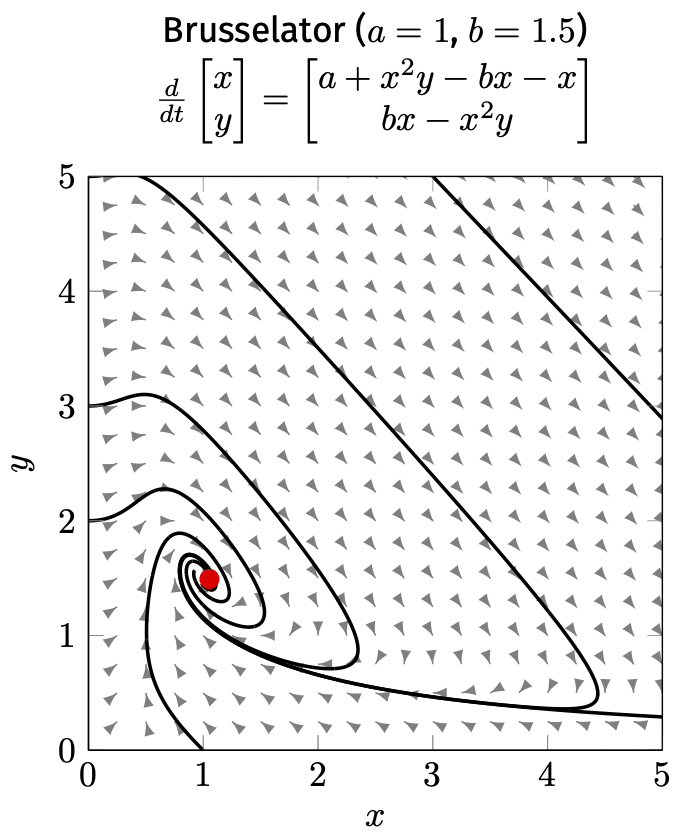
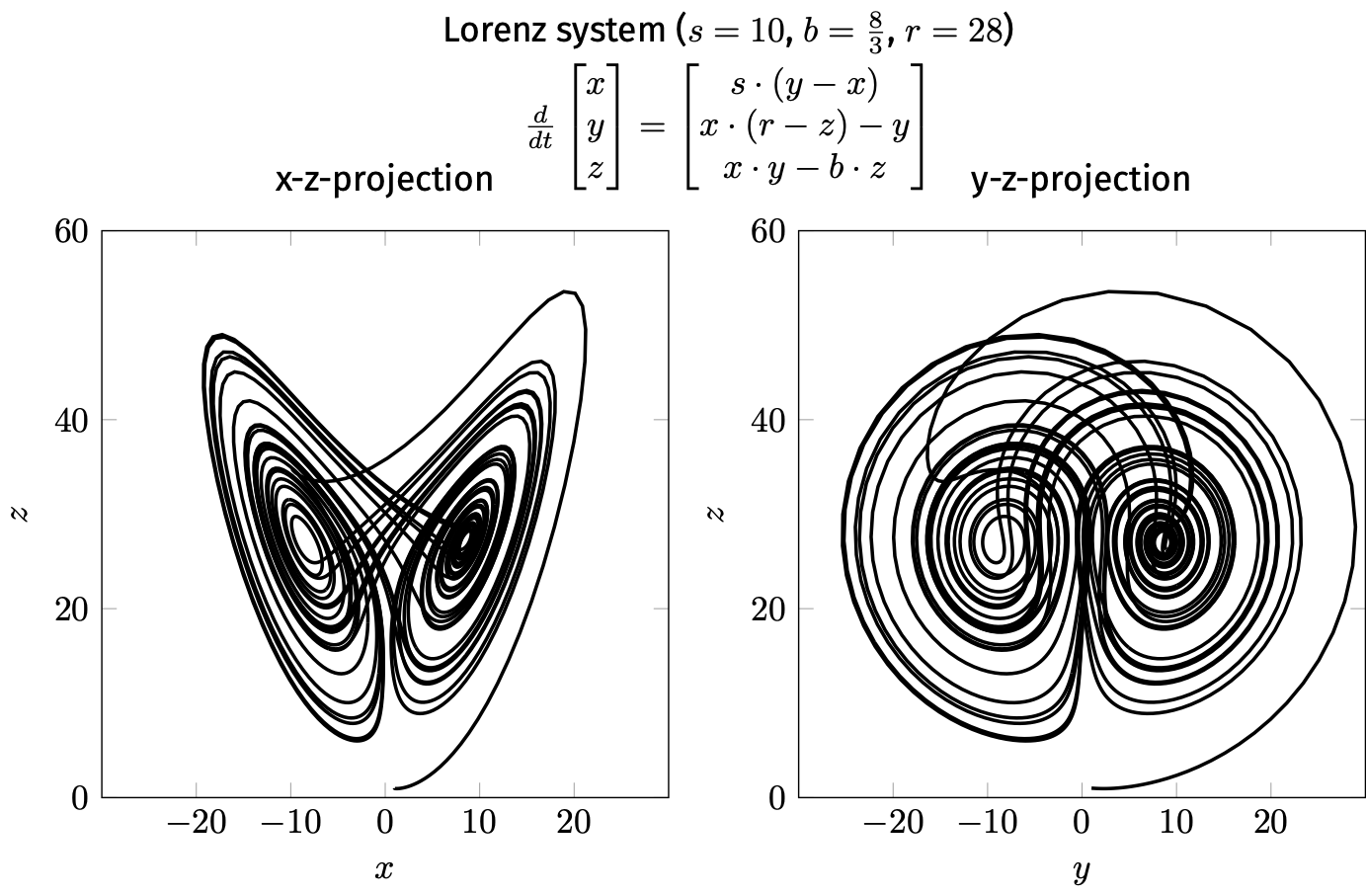
fixed-point
cycle
chaos
Bifurcation diagram: A (visual) partitioning of the parameter space into topologically equivalent classes.
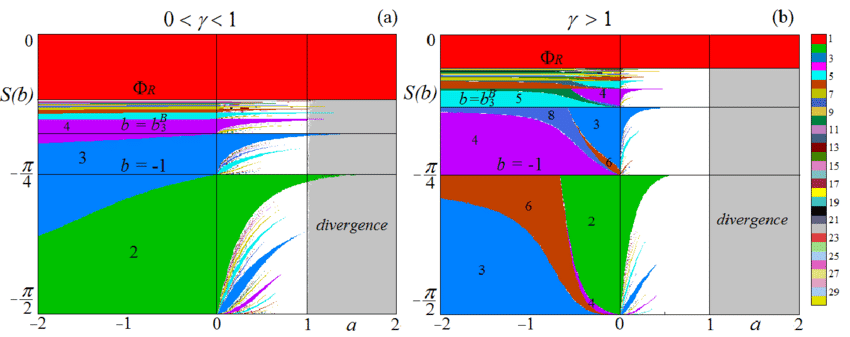
There is clearly an intuitive link between what we consider "similar long-term behaviour" in continuous and logical models.
Can we take inspiration from the "classical" bifurcation theory to enrich analysis of logical models?
Is there a possible counterpart to the topological equivalence?
How do we compute the parameter space decomposition?
What about "logical" bifurcation diagrams?
Equivalence of behaviours
* Shahrad Jamshidi, Heike Siebert, and Alexander Bockmayr. “Preservation of dynamic properties in qualitative modeling frameworks for gene regulatory networks”. In: Biosystems 112.2 (2013)
- Topological equivalence on transition systems is an isomorphism (not interesting).
- Classical bifurcation theory has no concept of non-determinism or fairness.
- Logical and continuous models of the same system often have fundamental differences in behaviour.*
(all models are wrong, some models are useful)
Alternative equivalence
- Take all valid runs of your system. (reflecting fairness, finiteness, time domain, initial states, etc.)
Alternative equivalence
- Take all valid runs of your system.
- Pick the smallest set of states that partitions these runs into finite segments.
Alternative equivalence
- Take all valid runs of your system.
- Pick the smallest set of states that partitions these runs into finite segments.
- Make a new graph from these states, based on the reachability in the original system.
Alternative equivalence
- Take all valid runs of your system.
- Pick the smallest set of states that partitions these runs into finite segments.
- Make a new graph from these states, based on the reachability in the original system.
- Two systems are equivalent if these condensed graphs are isomorphic.
\(\sim\)
In a fair system (e.g. BN), one representative state per attractor (bottom SCC) and repeller (top SCC) is selected. As fairness "decreases", more SCCs need to be included.
In a fair system, one representative state per SCC is sufficient. As fairness "decreases", more states need to be selected, up to the number of independent cycles.
Back to basics: (Bottom) SCC detection for asynchronous Boolean networks
Binary Decision Diagrams
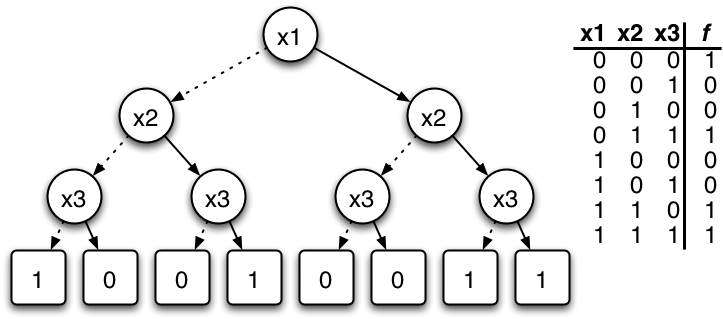
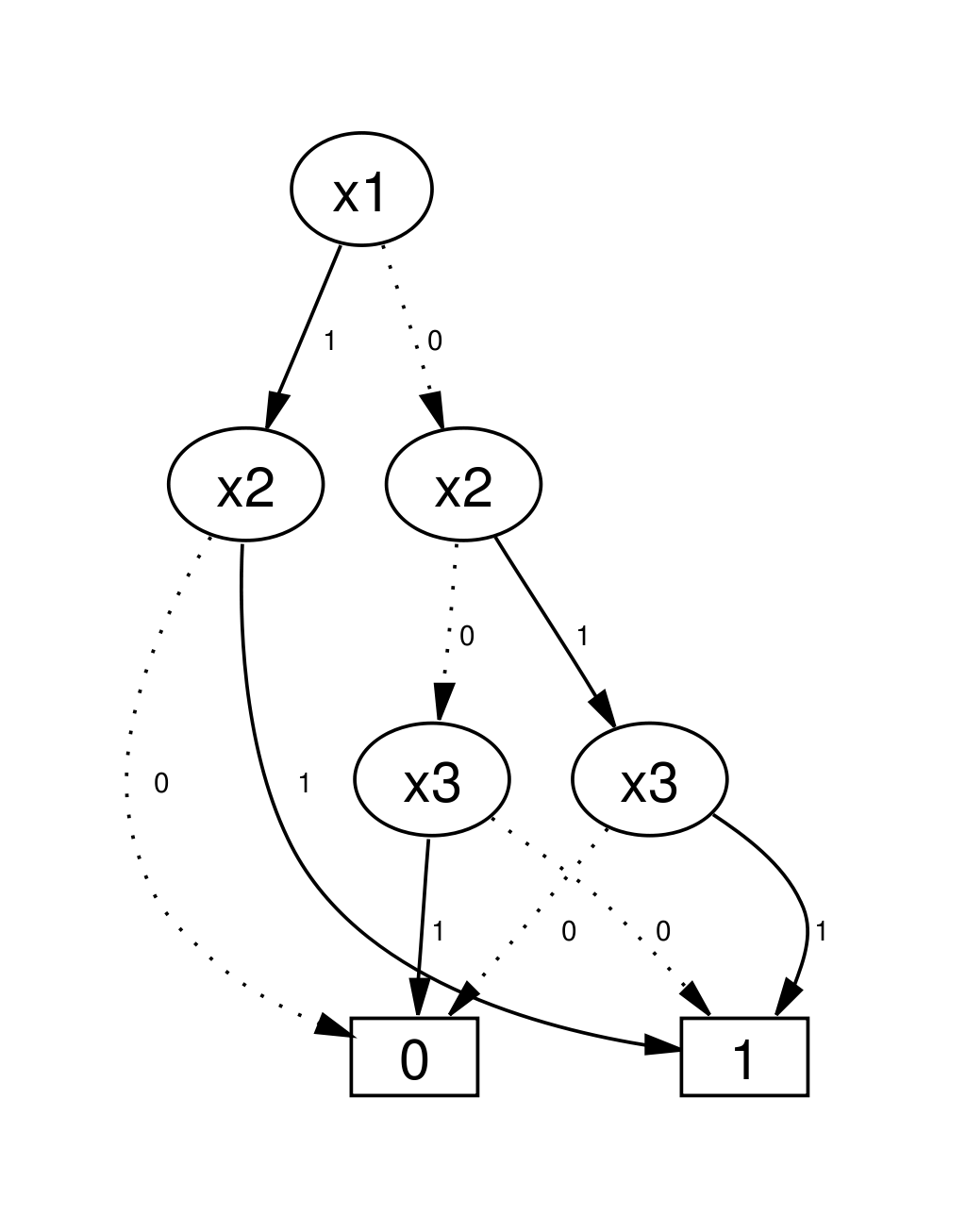
Parametrisation+State of a BN is a bit-vector. Sets/relations of these can be encoded via Boolean functions.
Set operations correspond to logical operations on functions (and graph operations on BDDs)
Since Boolean network is described by Boolean functions, computing successors/predecessors (including parameters) is relatively straightforward.
Benes, N., Brim, L., Pastva, S., & Šafránek, D. (2022). BDD-Based Algorithm for SCC Decomposition of Edge-Coloured Graphs. Log. Methods Comput. Sci., 18.
The first case is in pracitce much better!
Nobody can usually give you such guarantees...
Benes, N., Brim, L., Pastva, S., & Šafránek, D. (2021). Computing Bottom SCCs Symbolically Using Transition Guided Reduction. CAV.
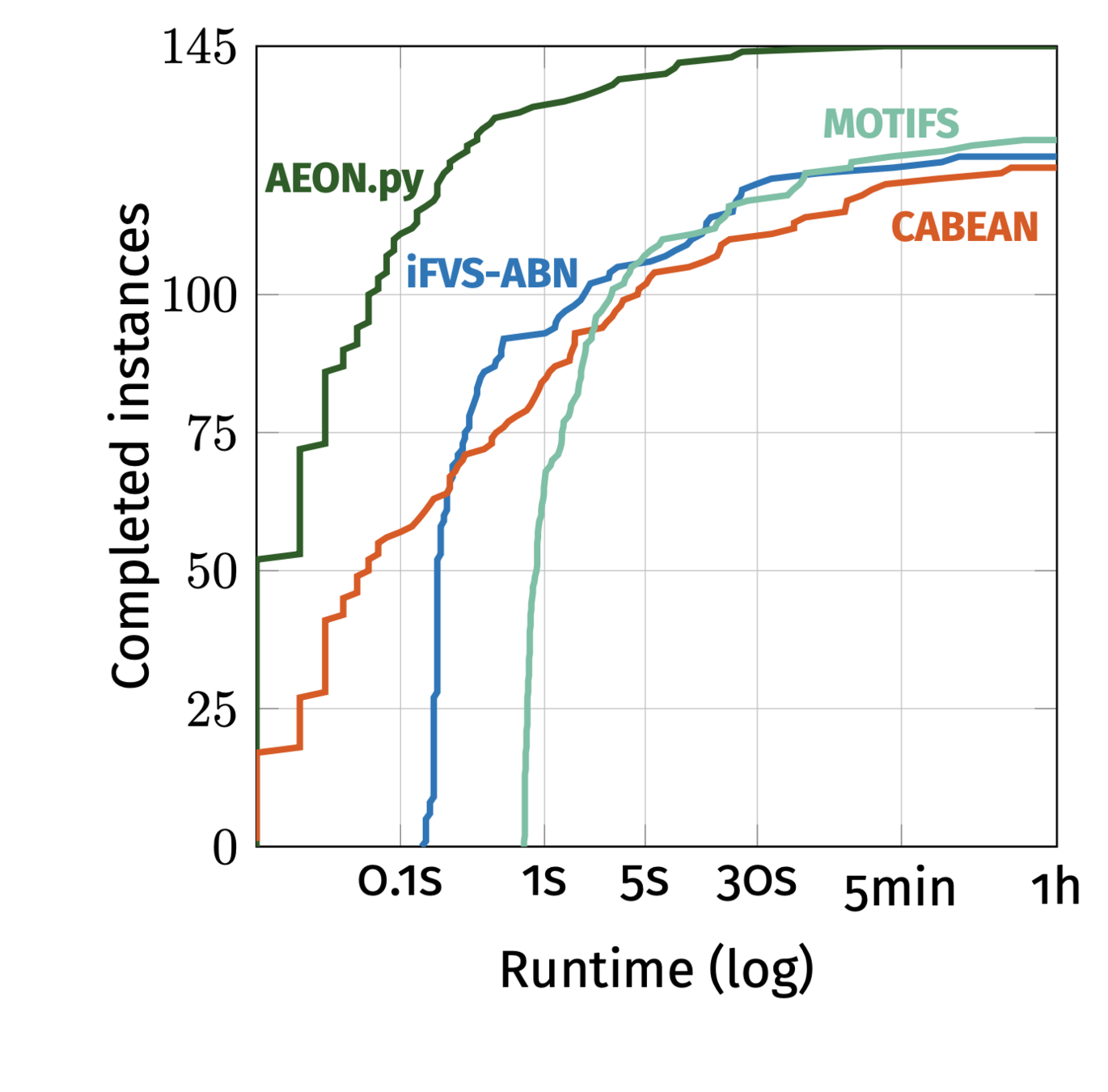
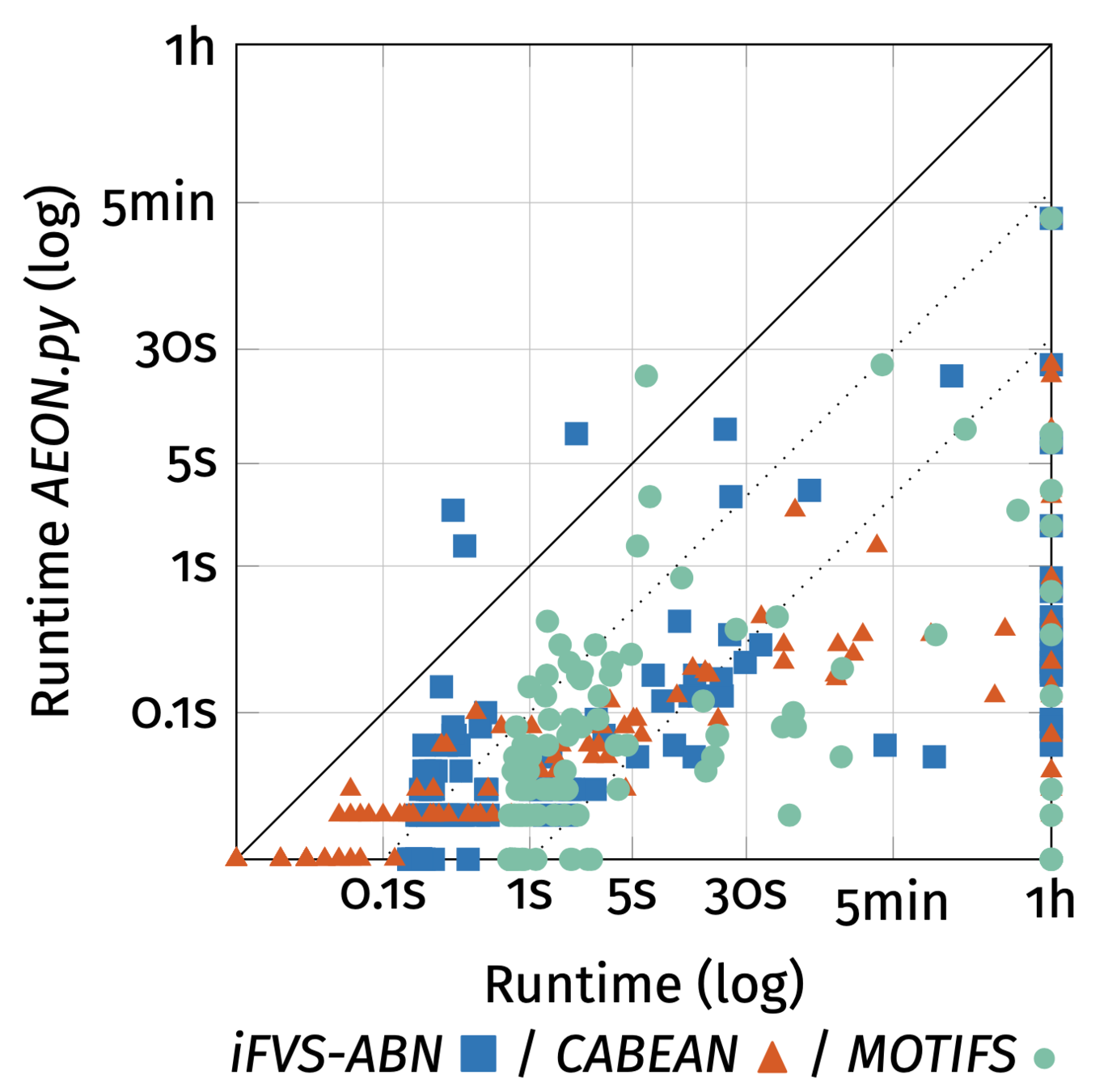
BDDs generally don't work, except when they do.
- We have a parameter space partitioning.
- We know how to compute it. (e.g. for 50+ Boolean parameters)
- How can we show it to you?
Bifurcation decision tree
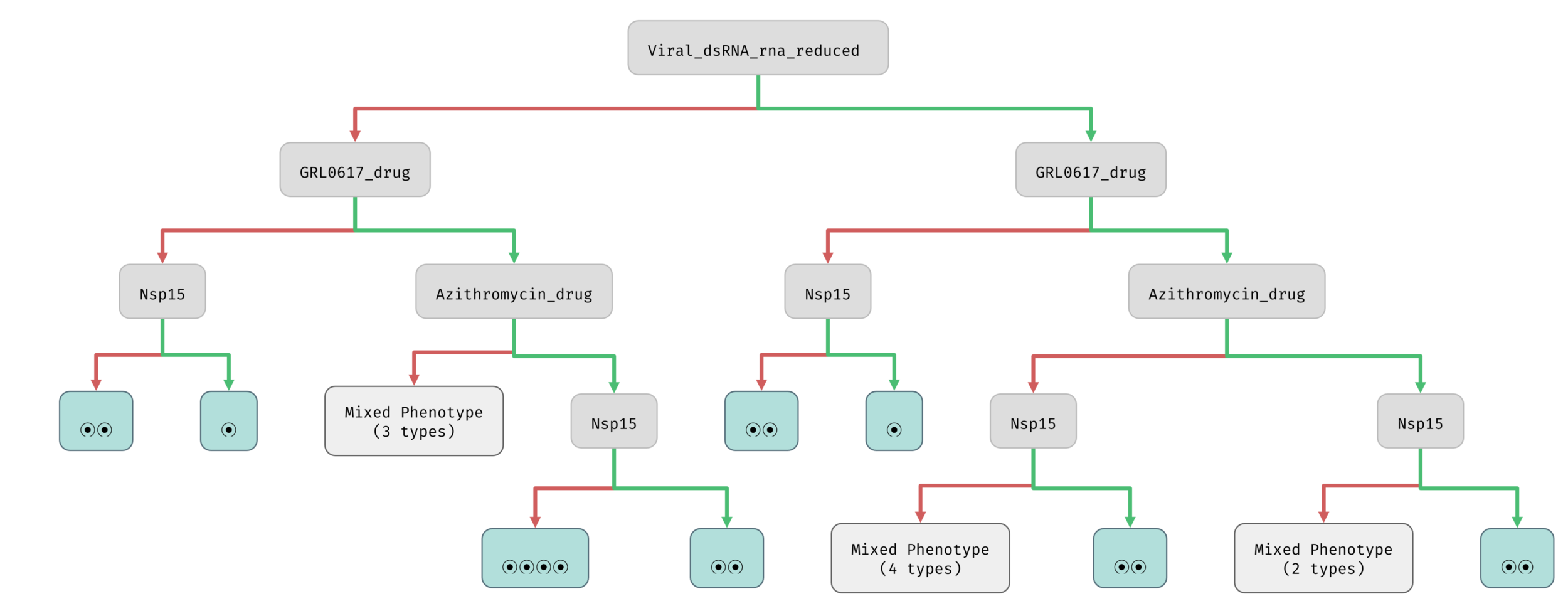
Different outcomes (classes of attractors)
Network parameters
Interactive, on-the-fly construction
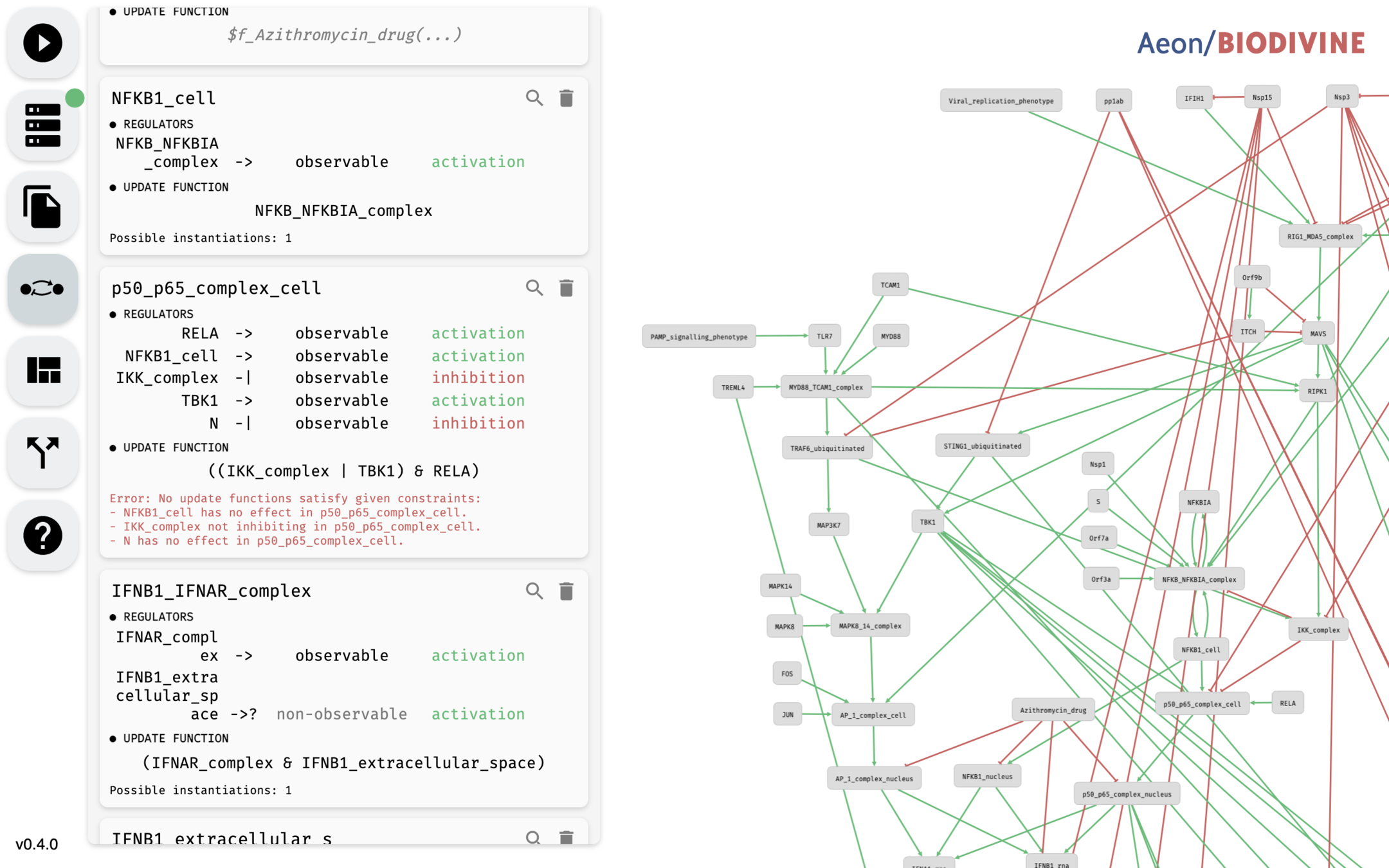
AEON
- Interactive web-based network and decision tree editor.
- Rust backend running symbolic BDD algorithms.
- Python API wrapper for use in Jupyter notebooks.
Benes, N., Brim, L., Kadlecaj, J., Pastva, S., & Šafránek, D. (2020). AEON: Attractor Bifurcation Analysis of Parametrised Boolean Networks. CAV.
Benes, N., Brim, L., Huvar, O., Pastva, S., Šafránek, D., & Smijáková, E. (2022). AEON.py: Python Library for Attractor Analysis in Asynchronous Boolean Networks. Bioinformatics.
T-cell survival network
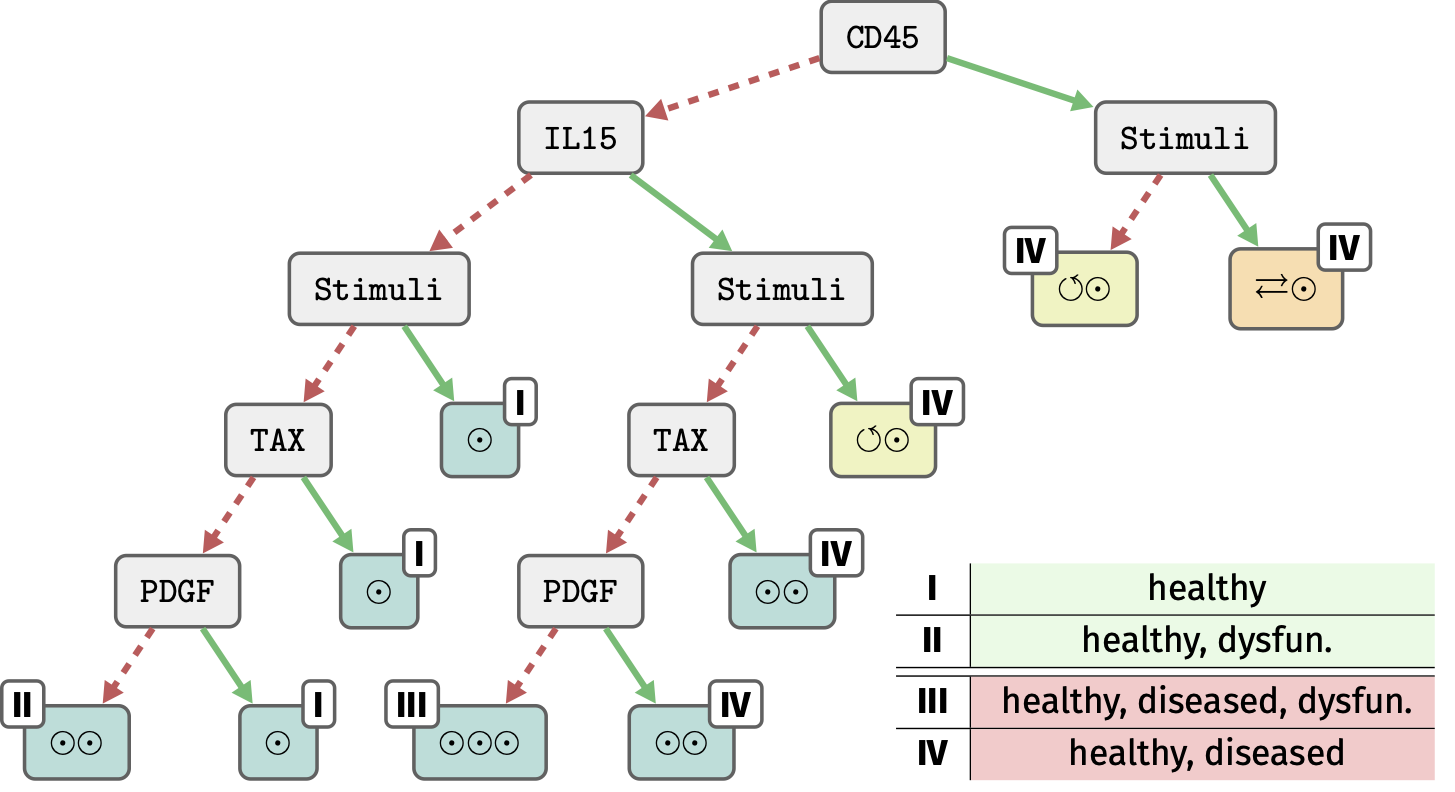
Zhang, R., et al. (2008). Network model of survival signaling in large granular lymphocyte leukemia. Proceedings of the National Academy of Sciences, 105, 16308 - 16313.
Newly characterised "phenotype"
Butanol pathway repair
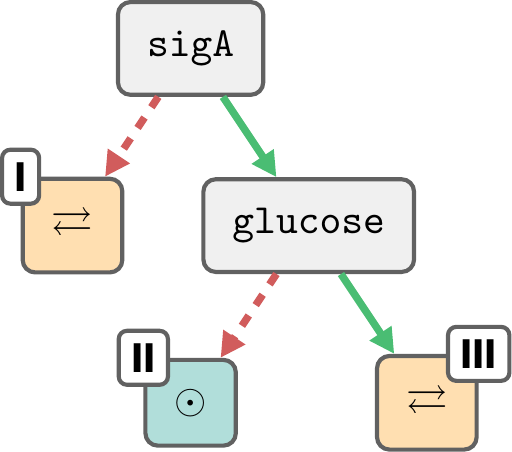

Out of 13 parameters, only 2 play a role in the long-term behaviour.
This relationship is maintained regardless of \(F_1, F_2, F_3\).
But we can further narrow down \(F_1, F_2, F_3\) through other means...
Jana Musilová. “Signaling pathway for butanol produc- tion in solventogenic clostridium bacteria”. MA thesis. Brno University of Technology, 2019.
Summary
- Computer science can stydy gene regulation through the lens of Boolean networks.
- Logical and functional parameters play a critical role in the long-term behaviour of Boolean networks.
- Symbolic methods can often work quite well as long as you are aware of the tradeoffs.
- Useful results are explainable: embrace interactivity and flexibility in communication.
For details, meet me in room 01.02.073 :)








Luboš Brim
David Šafranek
Nikola Beneš
Eva Šmijáková
Jakub Kadlecaj
Ondřej Huvar
Jakub Poláček