
Introduction to
Felipe Delestro
delestro@biologie.ens.fr

several programming languages and kernels
markdown
text for annotation and explanation
inline output with interactivity
EXAMPLE
Notebook running localy
Language Kernel
Jupyter notebook
user
| python | julia | ghc | nodejs | coffeescript |
| Brainfuck | R | F# | Go | Scala |
| Erlang | Torch | Elixir | Aldor | OCaml |
| Forth | Perl | PHP | Octave | Scilab |
| C | Matlab | Clojure | Hy | redis |
| io.js | Babel | Mathics | Wolfram | Lua |
| Scheme | Processing.js | IDL | Mochi | VPython |
| C# | Q | Cryptol | C++ | Xonsh |
| Prolog | Lisp | Maxima | Yacas | Jython |
| Gnuplot | Tcl | bash | TaQL | Coconut |
| NodeJS | Pike | Typescript | Kotlin | Babel |
installing
jupyter notebook
pip install jupyterin case of running a notebook server on bioclust, we need to use a virtual environment for the installation
pip install numpy scipy tornado
pyzmq pandas ipython pygments
matplotlib jinja2 jupyterif you find dependency problems....
Config file
to generate the config file for the first time, run:
jupyter notebook --generate-configthe file will be generated on:
~/.jupyter/jupyter_notebook_config.py
Config file
c = get_config()
c.NotebookApp.ip = '*'
c.NotebookApp.open_browser = False
c.NotebookApp.password = u'sha1:f6f35cb74557:ddaba774b3ef705b89b7639fdaee05eed2e4e0db'
c.NotebookApp.port = 4444
c.NotebookApp.base_url = '/local_notebook/'
c.NotebookApp.base_kernel_url = '/local_notebook/'
c.NotebookApp.tornado_settings = {'static_url_prefix':'/local_notebook/static/'}
c.NotebookApp.notebook_dir = u'/users/biocomp/delestro/memotrack/Python'Example
Interactive notebook
Example
running on bioclust using Tmux
With the same notebook server, you can run several kernels

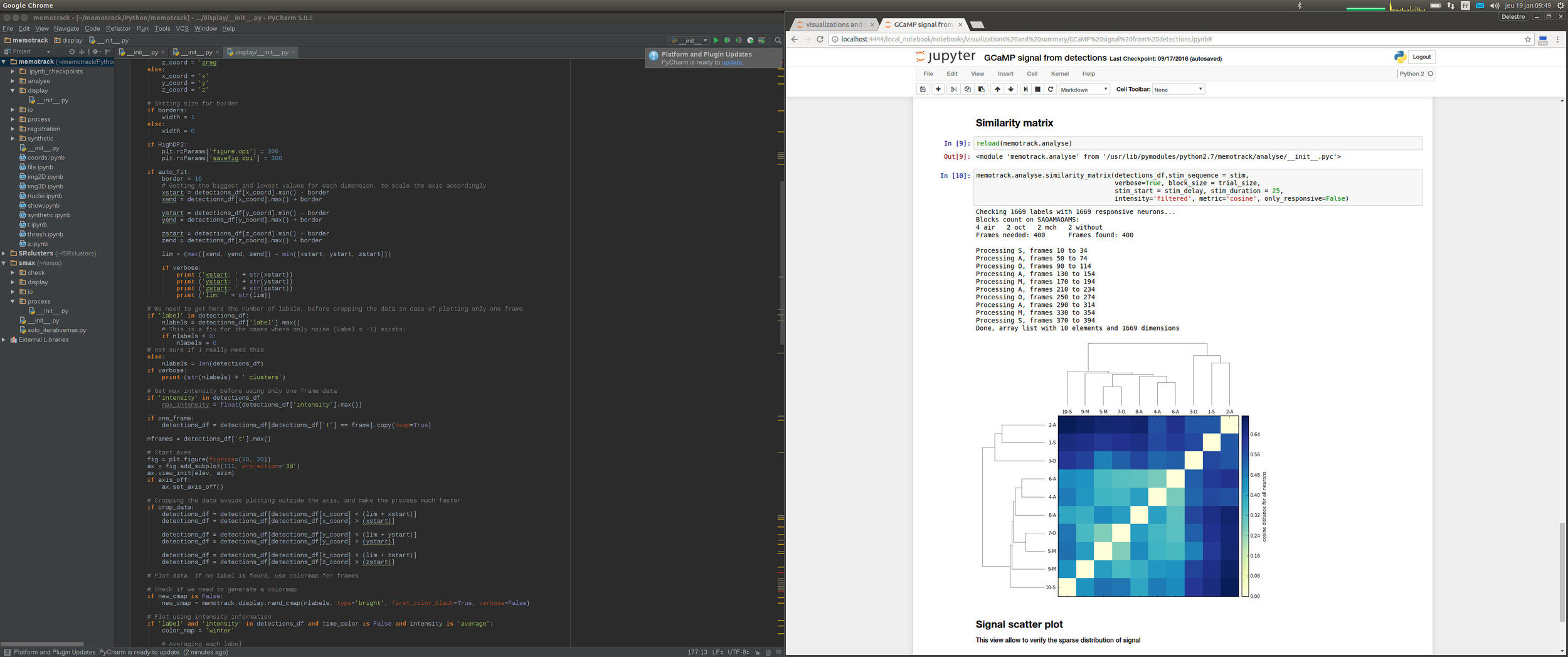
Pycharm for coding, Jupyter for running