Stian Soiland-Reyes
eScience lab, The University of Manchester
http://orcid.org/0000-0001-9842-9718
http://slides.com/soilandreyes/2017-11-16-cwl/
2018-04-25
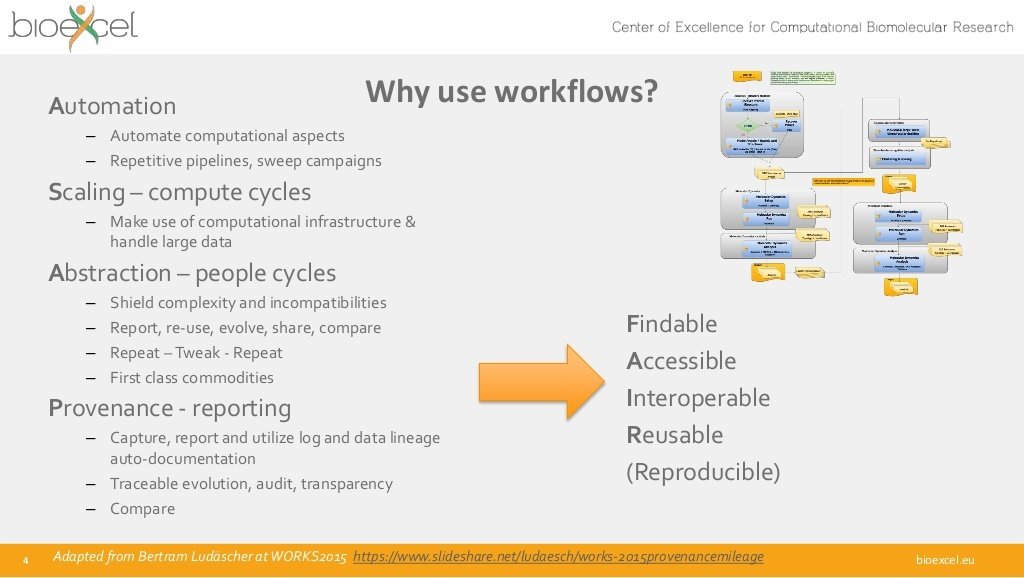
This work is licensed under a
Creative Commons Attribution 4.0 International License.



This work has been done as part of the BioExcel CoE (www.bioexcel.eu), a project funded by the European Union contract H2020-EINFRA-2015-1-675728.


cwlVersion: v1.0
class: Workflow
inputs:
inp: File
ex: string
outputs:
classout:
type: File
outputSource: compile/classfile
steps:
untar:
run: tar-param.cwl
in:
tarfile: inp
extractfile: ex
out: [example_out]
compile:
run: arguments.cwl
in:
src: untar/example_out
out: [classfile]








#!/usr/bin/env cwl-runner
cwlVersion: v1.0
class: CommandLineTool
baseCommand: [tar, xf]
inputs:
tarfile:
type: File
inputBinding:
position: 1
outputs:
example_out:
type: File
outputBinding:
glob: hello.txtCommand line tool



Where to find command line tools?
cwlVersion: v1.0
class: CommandLineTool
baseCommand: node
hints:
DockerRequirement:
dockerPull: node:slim#!/usr/bin/env cwl-runner
cwlVersion: v1.0
class: Workflow
inputs:
inp: File
ex: string
outputs:
classout:
type: File
outputSource: compile/classfile
steps:
untar:
run: tar-param.cwl
in:
tarfile: inp
extractfile: ex
out: [example_out]
compile:
run: arguments.cwl
in:
src: untar/example_out
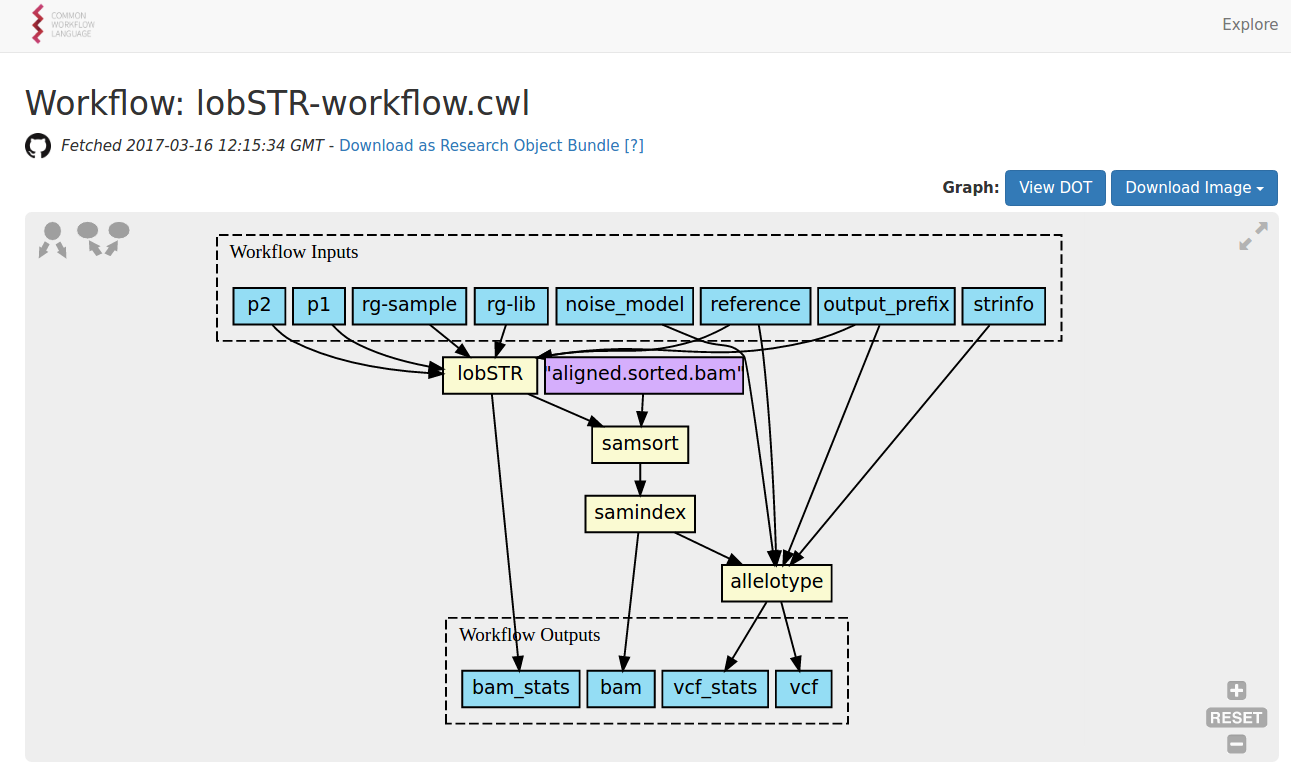
out: [classfile]Composing a workflow
cwlVersion: v1.0
class: Workflow
label: EMG QC workflow, (paired end version). Benchmarking with MG-RAST expt.
requirements:
- class: SubworkflowFeatureRequirement
- class: SchemaDefRequirement
types:
- $import: ../tools/FragGeneScan-model.yaml
- $import: ../tools/trimmomatic-sliding_window.yaml
- $import: ../tools/trimmomatic-end_mode.yaml
- $import: ../tools/trimmomatic-phred.yaml
inputs:
reads:
type: File
format: edam:format_1930 # FASTQ
outputs:
processed_sequences:
type: File
outputSource: clean_fasta_headers/sequences_with_cleaned_headers
steps:
trim_quality_control:
doc: |
Low quality trimming (low quality ends and sequences with < quality scores
less than 15 over a 4 nucleotide wide window are removed)
run: ../tools/trimmomatic.cwl
in:
reads1: reads
phred: { default: '33' }
leading: { default: 3 }
trailing: { default: 3 }
end_mode: { default: SE }
minlen: { default: 100 }
slidingwindow:
default:
windowSize: 4
requiredQuality: 15
out: [reads1_trimmed]
convert_trimmed-reads_to_fasta:
run: ../tools/fastq_to_fasta.cwl
in:
fastq: trim_quality_control/reads1_trimmed
out: [ fasta ]
clean_fasta_headers:
run: ../tools/clean_fasta_headers.cwl
in:
sequences: convert_trimmed-reads_to_fasta/fasta
out: [ sequences_with_cleaned_headers ]
$namespaces:
edam: http://edamontology.org/
s: http://schema.org/
$schemas:
- http://edamontology.org/EDAM_1.16.owl
- https://schema.org/docs/schema_org_rdfa.html
s:license: "https://www.apache.org/licenses/LICENSE-2.0"
s:copyrightHolder: "EMBL - European Bioinformatics Institute"

document
prefix wfprov <http://purl.org/wf4ever/wfprov#>
prefix prov <http://www.w3.org/ns/prov#>
prefix wfdesc <http://purl.org/wf4ever/wfdesc#>
prefix wf <https://w3id.org/cwl/view/git/933bf2a1a1cce32d88f88f136275535da9df0954/workflows/hello/hello.cwl#>
prefix input <app://579c1b74-b328-4da6-80a8-a2ffef2ac9b5/workflow/input.json#>
prefix run <urn:uuid:>
prefix engine <urn:uuid:>
prefix data <urn:hash:sha256:>
default <app://579c1b74-b328-4da6-80a8-a2ffef2ac9b5/>
// Level 1 provenance of workflow run
activity(run:2e1287e0-6dfb-11e7-8acf-0242ac110002, , , [prov:type='wfprov:WorkflowRun', prov:label="Run of workflow/packed.cwl#main"])
wasStartedBy(run:2e1287e0-6dfb-11e7-8acf-0242ac110002, -, -, -, 2017-10-27T14:24:00+01:00)
// The engine is the SoftwareAgent that is executing our Workflow plan
wasAssociatedWith(run:2e1287e0-6dfb-11e7-8acf-0242ac110002, engine:b2210211-8acb-4d58-bd28-2a36b18d3b4f, wf:main)
agent(engine:b2210211-8acb-4d58-bd28-2a36b18d3b4f, prov:type='prov:SoftwareAgent', prov:type='wfprov:WorkflowEngine', prov:label="cwltool v1.2.5")
// prov has no term to relate sub-plans - we'll use wfdesc:hasSubProcess
entity(wf:main,[prov:type='wfdesc:Workflow', prov:type='prov:Plan', wfdesc:hasSubProcess='wf:main/step1', wfdesc:hasSubProcess='wf:main/step2'])
alternateOf(wf:main, workflow/packed.cwl)
entity(wf:main/step1,[prov:type='wfdesc:Process', prov:type='prov:Plan'])
entity(wf:main/step2,[prov:type='wfdesc:Process', prov:type='prov:Plan'])
// First the workflow uses some data; here with a urn:sha:sha256 identifier
used(run:2e1287e0-6dfb-11e7-8acf-0242ac110002, data:5891b5b522d5df086d0ff0b110fbd9d21bb4fc7163af34d08286a2e846f6be03, 2017-10-27T14:29:00+01:00, [prov:role='wf:main/input1']))
entity(data:5891b5b522d5df086d0ff0b110fbd9d21bb4fc7163af34d08286a2e846f6be03, [prov:type='wfprov:Artifact'])
// which we have stored a copy of within the research object
specializationOf(data/58/5891b5b522d5df086d0ff0b110fbd9d21bb4fc7163af34d08286a2e846f6be03, data:5891b5b522d5df086d0ff0b110fbd9d21bb4fc7163af34d08286a2e846f6be03)
// Then there was another activity - wfprov:ProcessRun indicating a command line tool
activity(run:4305467e-6dfb-11e7-885d-0242ac110002, -, -, [prov:type='wfprov:ProcessRun', prov:label="Run of workflow/packed.cwl#main/step1"])
// started by the mother activity
wasStartedBy(run:4305467e-6dfb-11e7-885d-0242ac110002, -, -, run:2e1287e0-6dfb-11e7-8acf-0242ac110002, 2017-10-27T15:00:00+01:00)
// same engine using step1 as plan. In a distributed scenario there might be a different engine
wasAssociatedWith(run:4305467e-6dfb-11e7-885d-0242ac110002, engine:b2210211-8acb-4d58-bd28-2a36b18d3b4f, wf:main/step1)
// This activity also use the same data, but in a different role (e.g. input parameter)
used(run:4305467e-6dfb-11e7-885d-0242ac110002, data:5891b5b522d5df086d0ff0b110fbd9d21bb4fc7163af34d08286a2e846f6be03, 2017-10-27T14:00:00+01:00, [prov:role='wf:main/step1/in1'])
// And we generate some new data
wasGeneratedBy(data:00688350913f2f292943a274b57019d58889eda272370af261c84e78e204743c, run:4305467e-6dfb-11e7-885d-0242ac110002, 2017-10-27T16:00:00+01:00, [prov:role='wf:main/step1/out1']))
entity(data:00688350913f2f292943a274b57019d58889eda272370af261c84e78e204743c, [prov:type='wfprov:Artifact'])
// again stored in the RO
specializationOf(data/00/00688350913f2f292943a274b57019d58889eda272370af261c84e78e204743c, data:00688350913f2f292943a274b57019d58889eda272370af261c84e78e204743c)
// step1 finished
wasEndedBy(run:4305467e-6dfb-11e7-885d-0242ac110002, -, -, run:2e1287e0-6dfb-11e7-8acf-0242ac110002, 2017-10-27T15:30:00+01:00)
// the master workflow then "generate" that same value, but now at a different time and role (the resultA master workflow output)
wasGeneratedBy(data:00688350913f2f292943a274b57019d58889eda272370af261c84e78e204743c, run:2e1287e0-6dfb-11e7-8acf-0242ac110002, 2017-10-27T15:00:00+01:00, [prov:role='wf:main/resultA'])
// next step activity
activity(run:c42dc36e-6dfd-11e7-bc24-0242ac110002, -, - [prov:type='wfprov:ProcessRun', prov:label="Run of workflow/packed.cwl#main/step2"])
wasStartedBy(run:c42dc36e-6dfd-11e7-bc24-0242ac110002, -, -, run:2e1287e0-6dfb-11e7-8acf-0242ac110002, 2017-10-27T16:00:00+01:00)
// associated with step2
wasAssociatedWith(run:c42dc36e-6dfd-11e7-bc24-0242ac110002, engine:b2210211-8acb-4d58-bd28-2a36b18d3b4f, wf:main/step2)
// Uses two data artifacts; one which came from previous step, other as workflow input
used(run:4305467e-6dfb-11e7-885d-0242ac110002, data:5891b5b522d5df086d0ff0b110fbd9d21bb4fc7163af34d08286a2e846f6be03, 2017-10-27T15:00:00+01:00, [prov:role='wf:main/step2/valueA'])
used(run:4305467e-6dfb-11e7-885d-0242ac110002, data:00688350913f2f292943a274b57019d58889eda272370af261c84e78e204743c, 2017-10-27T15:00:00+01:00, [prov:role='wf:main/step2/valueB'])
// and generate two new data artifacts
wasGeneratedBy(data:952f537d1f3116db56703787ace248fe00ae46fa77ea3803aa3d8dc01d221a9d, run:c42dc36e-6dfd-11e7-bc24-0242ac110002, 2017-10-27T16:34:20+01:00, [prov:role='wf:main/step2/out1'])))
entity(data:952f537d1f3116db56703787ace248fe00ae46fa77ea3803aa3d8dc01d221a9d, [prov:type='wfprov:Artifact'])
specializationOf(data/95/2f537d1f3116db56703787ace248fe00ae46fa77ea3803aa3d8dc01d221a9d, data:952f537d1f3116db56703787ace248fe00ae46fa77ea3803aa3d8dc01d221a9d)
wasGeneratedBy(data:3deb00bd0decd1f21d015a178c4f23a5eb537588c08eeee9d55059ec29637be0, run:c42dc36e-6dfd-11e7-bc24-0242ac110002, 2017-10-27T16:34:20+01:00, [prov:role='wf:main/step2/out2'])))
entity(data:3deb00bd0decd1f21d015a178c4f23a5eb537588c08eeee9d55059ec29637be0, [prov:type='wfprov:Artifact'])
specializationOf(data/3d/eb00bd0decd1f21d015a178c4f23a5eb537588c08eeee9d55059ec29637be0, data:3deb00bd0decd1f21d015a178c4f23a5eb537588c08eeee9d55059ec29637be0)
// step2 ends
wasEndedBy(run:c42dc36e-6dfd-11e7-bc24-0242ac110002, -, -, run:2e1287e0-6dfb-11e7-8acf-0242ac110002, 2017-10-27T16:30:00+01:00)
// only step output out1 captured by mother workflow, sent to resultB workflow output
wasGeneratedBy(data:952f537d1f3116db56703787ace248fe00ae46fa77ea3803aa3d8dc01d221a9d, run:2e1287e0-6dfb-11e7-8acf-0242ac110002, 2017-10-27T15:00:00+01:00, [prov:role='wf:main/resultB'])
// mother workflow ends
wasEndedBy(run:2e1287e0-6dfb-11e7-8acf-0242ac110002, -, -, run:2e1287e0-6dfb-11e7-8acf-0242ac110002, 2017-10-27T16:34:40+01:00)
endDocument
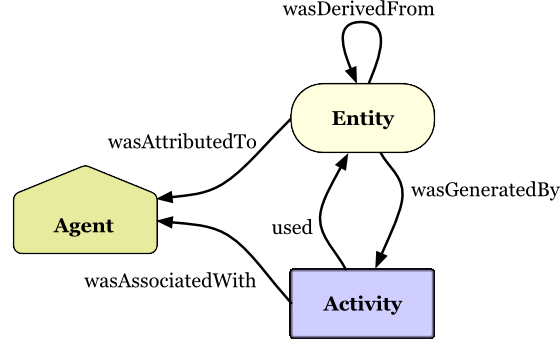
CWLProv

