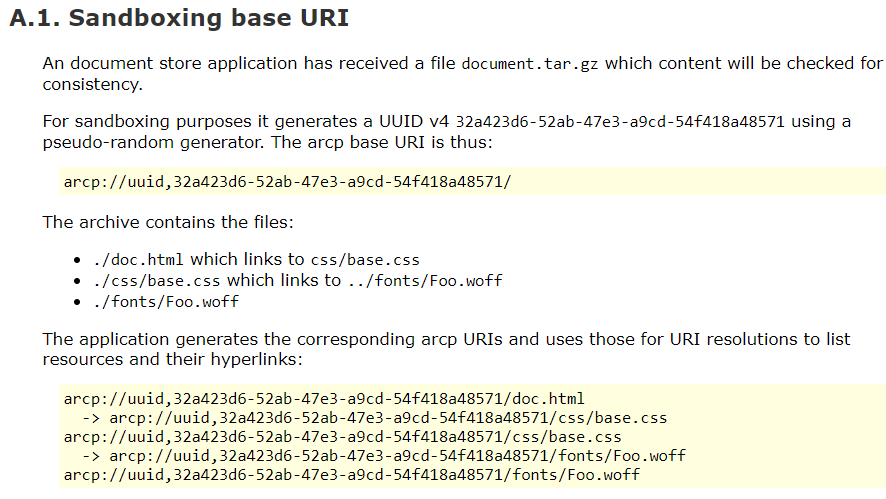
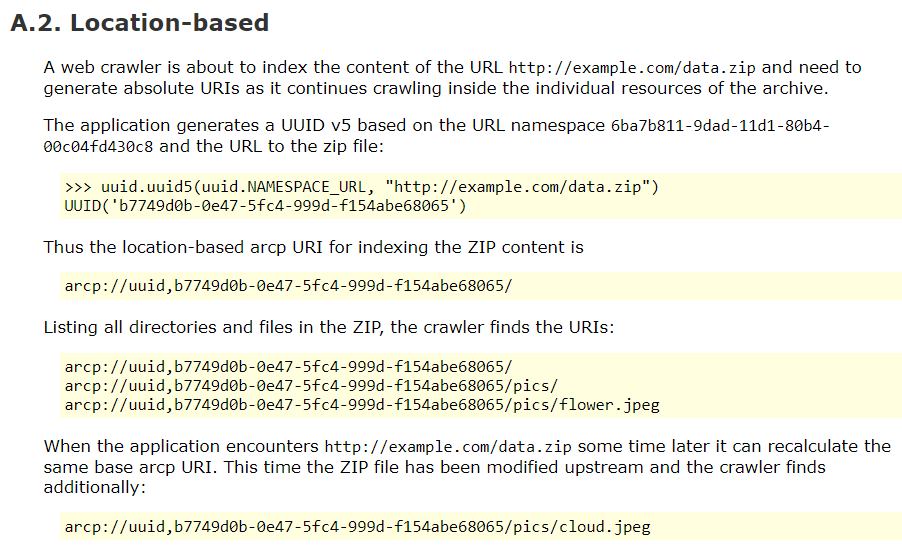
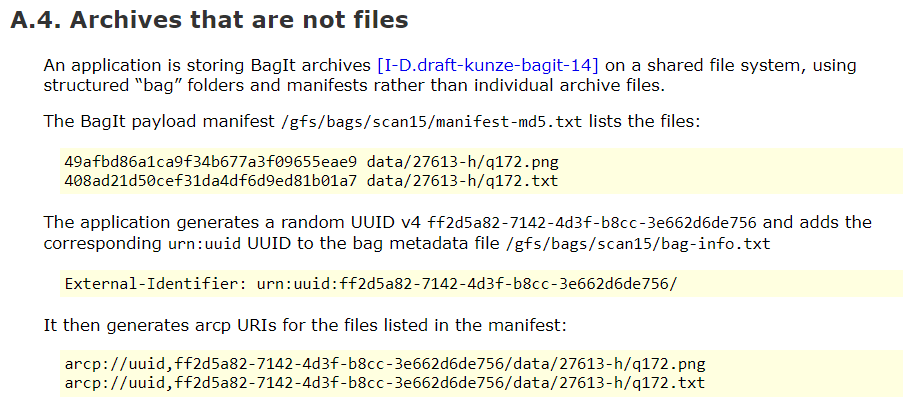
Research Object
Stian Soiland-Reyes, Carole Goble
eScience lab, The University of Manchester
Research Data Packaging workshop
Open Repositories (OR2019), Hamburg, 2019-06-10
This work is licensed under a
Creative Commons Attribution 4.0 International License.






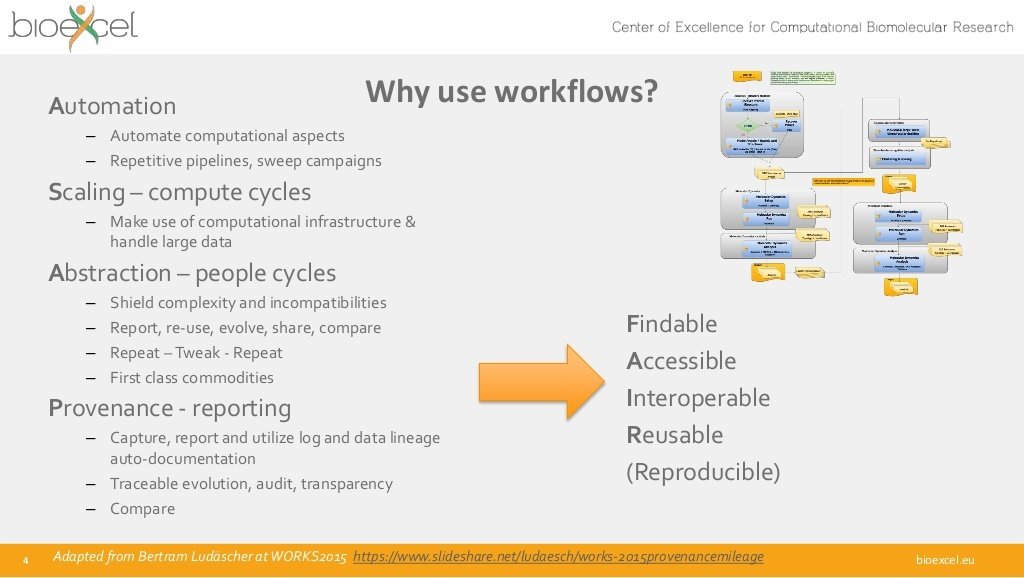
Findable
Accessible
Interoperable
Reusable

F1. (meta)data are assigned a globally unique and persistent identifier
F2. data are described with rich metadata (defined by R1 below)
F3. metadata clearly and explicitly include the identifier of the data it describes
F4. (meta)data are registered or indexed in a searchable resource
To be Findable:
A1. (meta)data are retrievable by their identifier using a
standardized communications protocol
A1.1 the protocol is open, free, and universally implementable
A1.2 the protocol allows for an authentication and authorization
procedure, where necessary
A2. metadata are accessible, even when the data are no longer available
To be Accessible:
I1. (meta)data use a formal, accessible, shared, and
broadly applicable language for knowledge representation.
I2. (meta)data use vocabularies that follow FAIR principles
I3. (meta)data include qualified references to other (meta)data
To be Interoperable:
R1. meta(data) are richly described with a plurality of
accurate and relevant attributes
R1.1. (meta)data are released with a clear and accessible data usage license
R1.2. (meta)data are associated with detailed provenance
R1.3. (meta)data meet domain-relevant community standards
To be Reusable:


They ride with what I refer to as the four horsemen of the reproducibility apocalypse:
- Publication bias
- Low statistical power
- P-value hacking
- HARKing (hypothesizing after results are known)


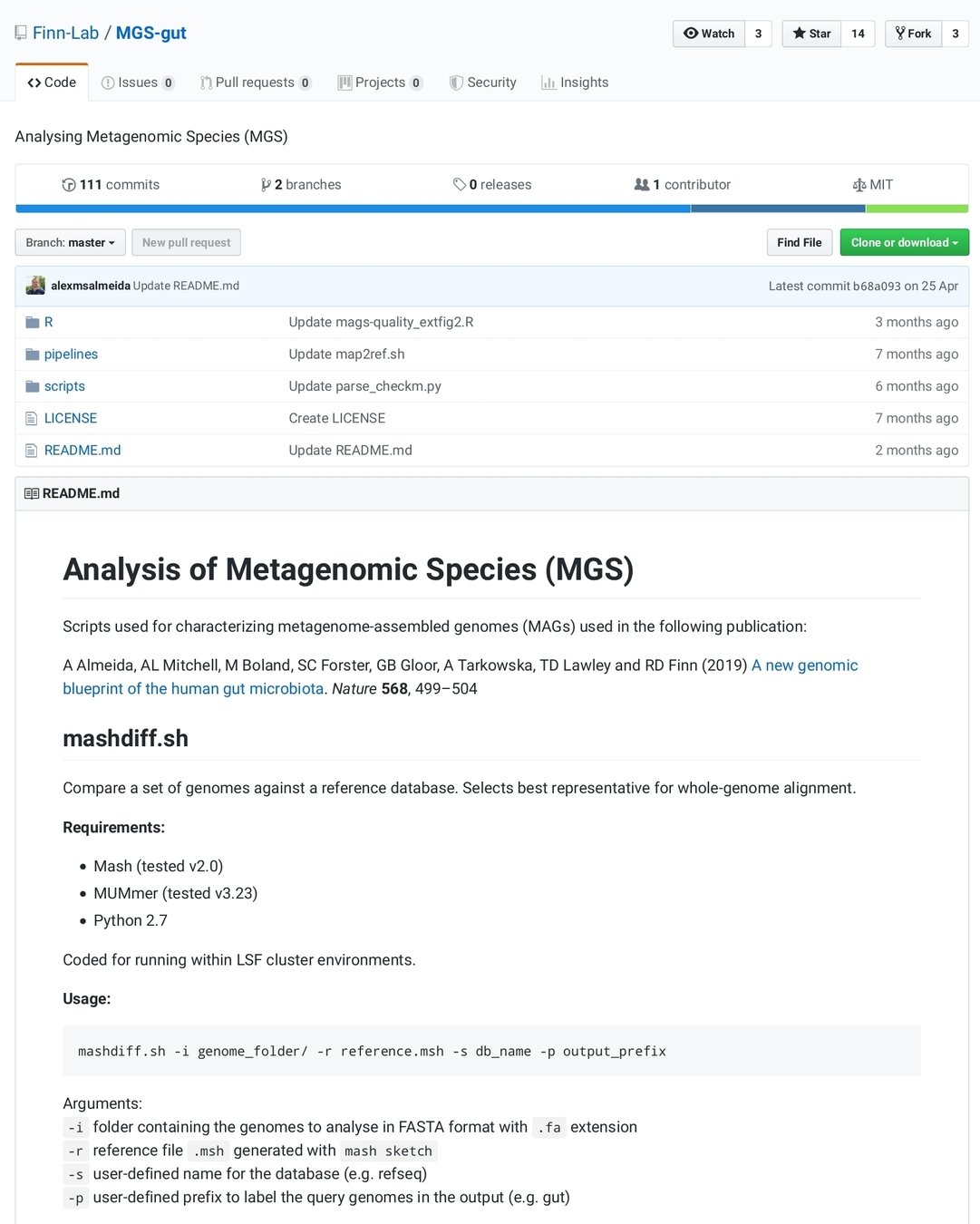

State of art in reproducibility



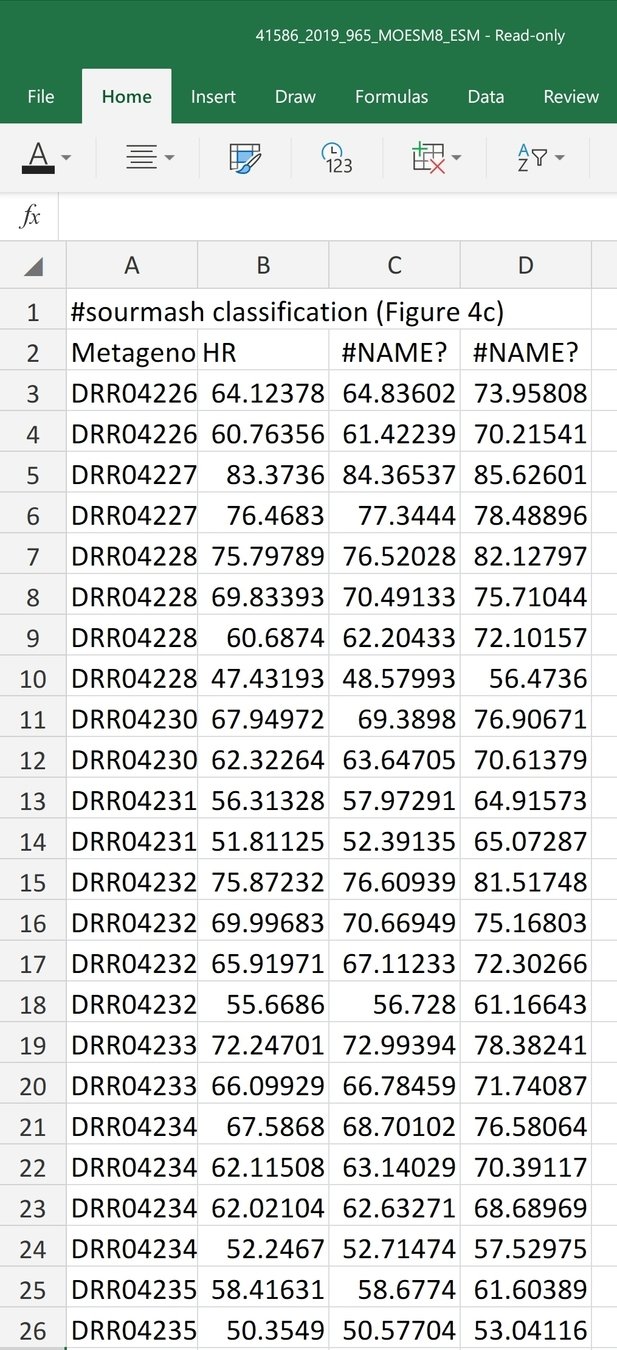

Data availability:
Domain-specific databases
CSV
FTP









Reproducibility?




Attribution


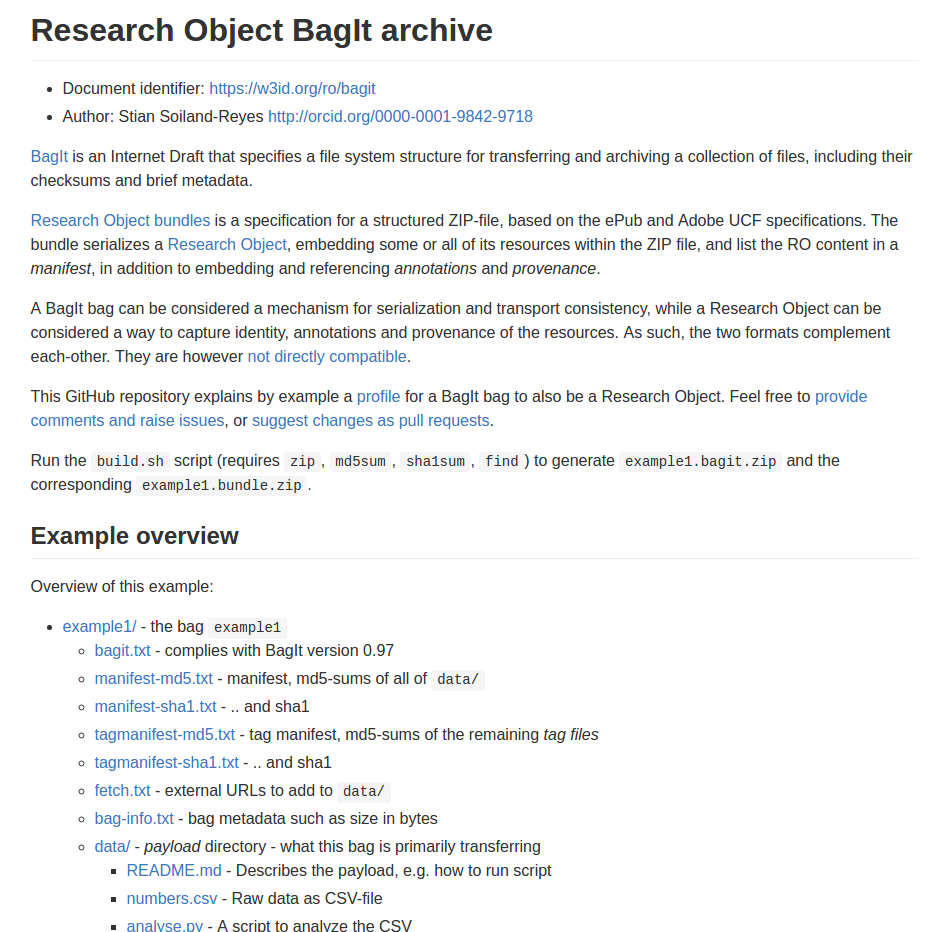
A Research Object bundles and relates digital resources of a scientific experiment or investigation:
Data used and results produced in experimental study
Methods employed to produce and analyse that data
Provenance and settings for the experiments
People involved in the investigation
Annotations about these resources, to improve understanding and interpretation
Research Object
id: doi:10.15490/seek.1.investigation.56
createdOn: 2015-07-10T16:46:00Z
createdBy: http://orcid.org/0000-0001-9842-9718
aggregates:
- id: data/sequence/specimen5.bam
conformsTo: http://gemrb.org/iesdp/file_formats/ie_formats/bam_v1.htm
- id: http://example.com/blog/about-specimen5
authoredBy: http://orcid.org/0000-0001-7066-3350
- id: http://www.myexperiment.org/workflows/3355
history: provenance/workflow-evolution.ttl
annotations:
- about: data/sequence/specimen5.bam
content: annotations/specimen5-properties.jsonld
createdBy: http://orcid.org/0000-0001-7066-3350
- about: data/sequence/specimen5.bam
content: http://example.com/blog/about-specimen5
motivatedBy: oa:questioningResearch Object manifest
(simplified)
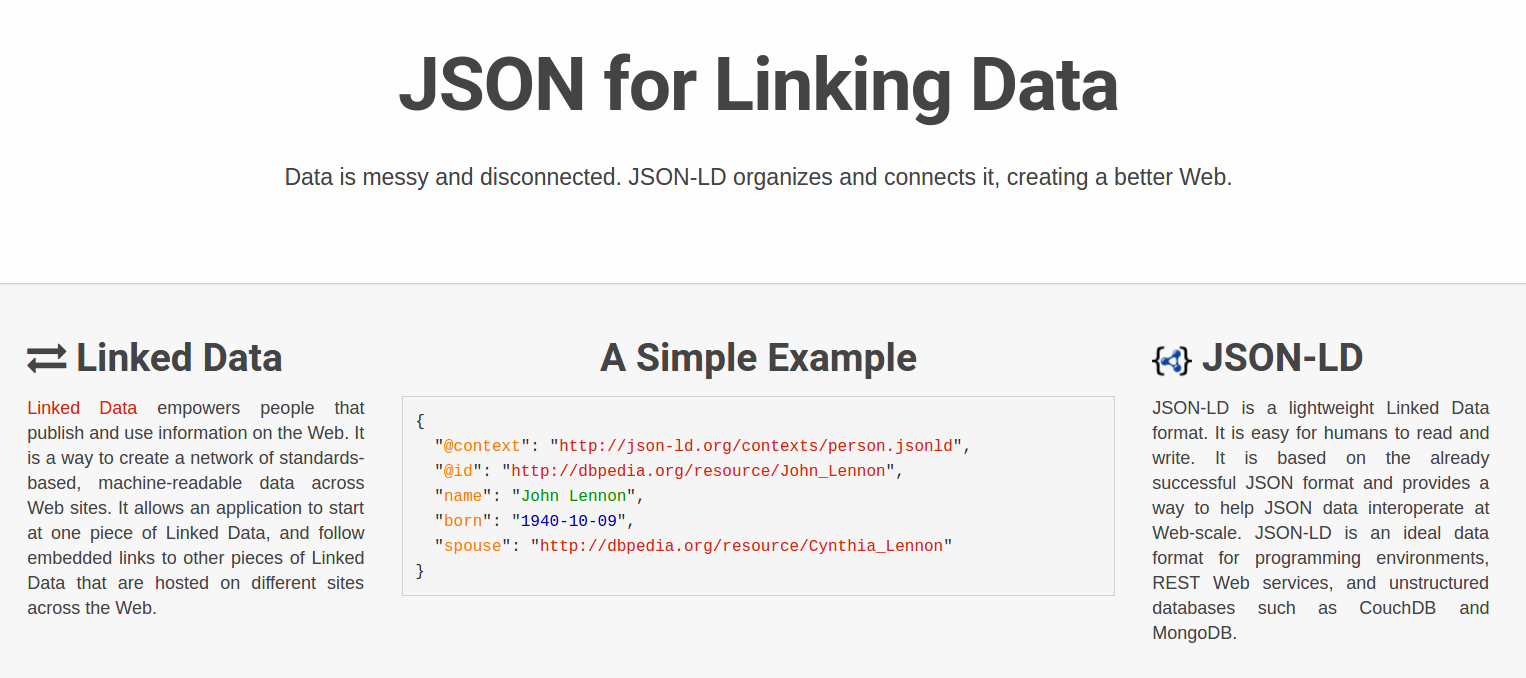
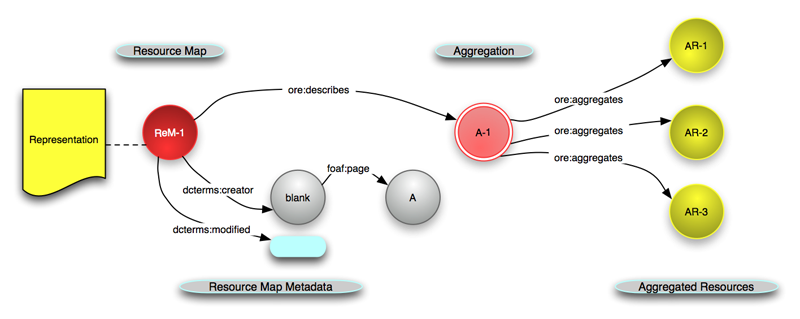
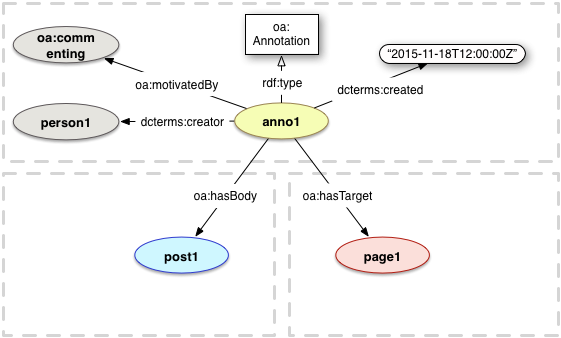
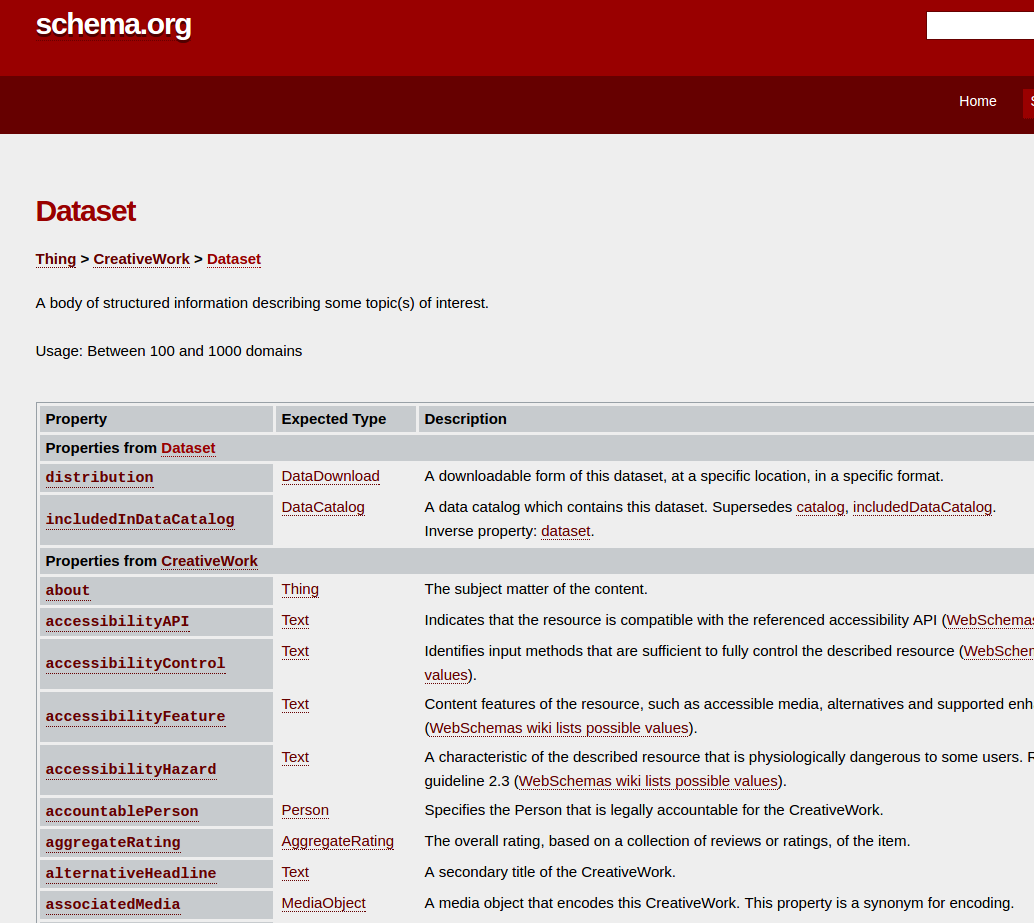
Reuse standards:
OAI-ORE, BagIt, W3C JSON-LD, PROV, Web Annotation Model

metadata/manifest.json
data/sequence/specimen5.bam
provenance/workflow-evolution.ttl
http://example.com/blog/about-specimen5
http://www.myexperiment.org/workflows/335
http://orcid.org/0000-0001-7066-3350
http://gemrb.org/iesdb/
file_formats_ie_formats_bam_v1.html


pip install bdbag





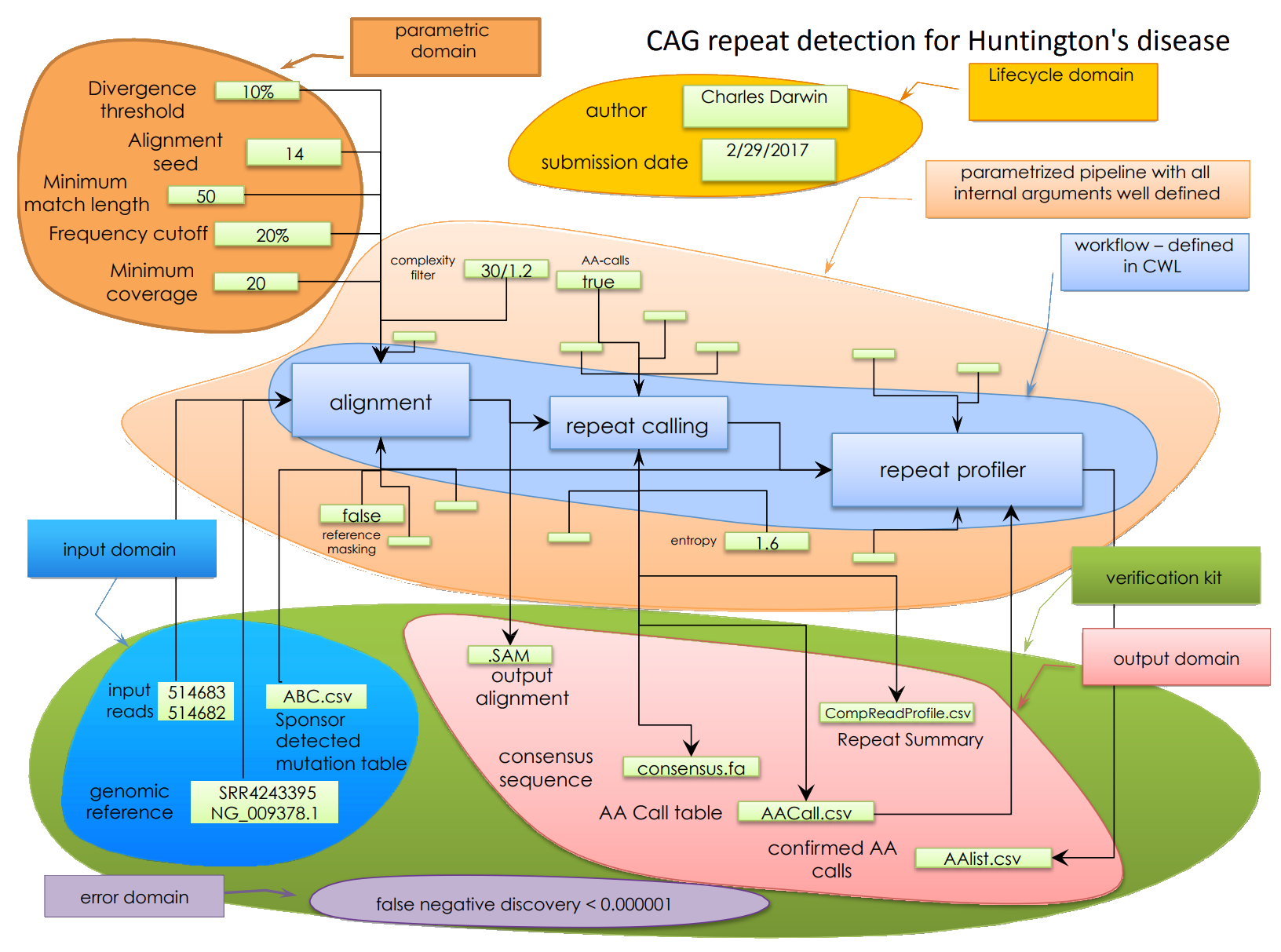

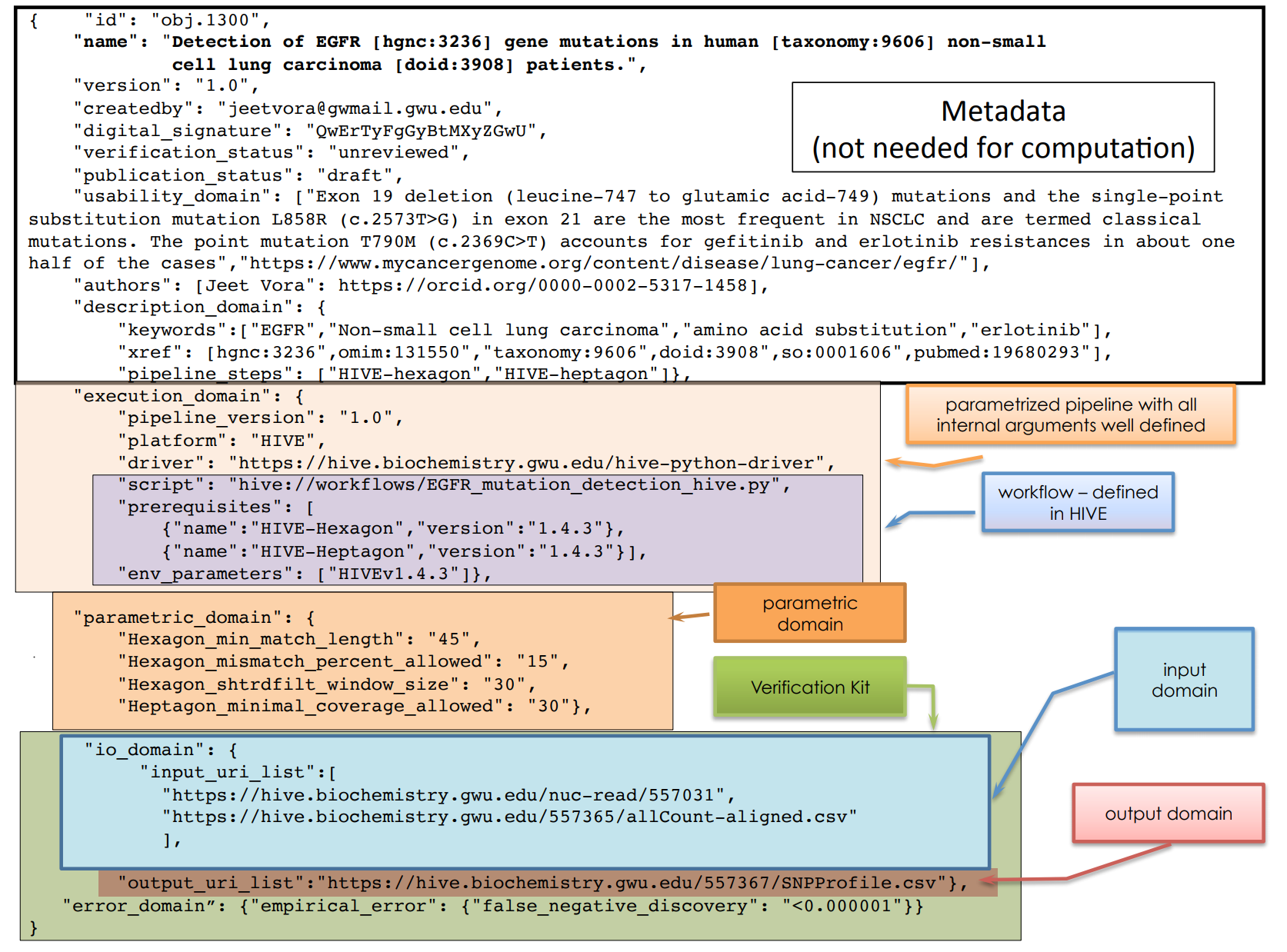


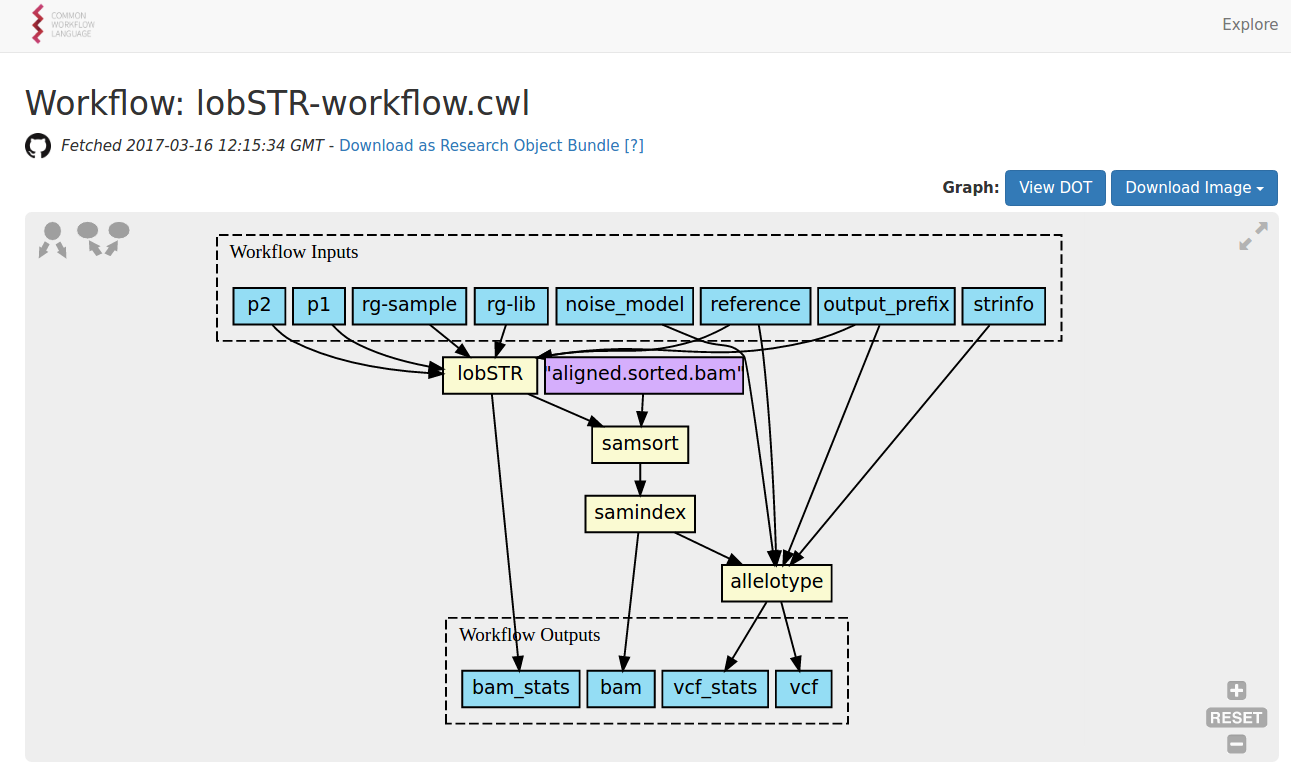
How can we capture the methods?

cwlVersion: v1.0
class: Workflow
inputs:
inp: File
ex: string
outputs:
classout:
type: File
outputSource: compile/classfile
steps:
untar:
run: tar-param.cwl
in:
tarfile: inp
extractfile: ex
out: [example_out]
compile:
run: arguments.cwl
in:
src: untar/example_out
out: [classfile]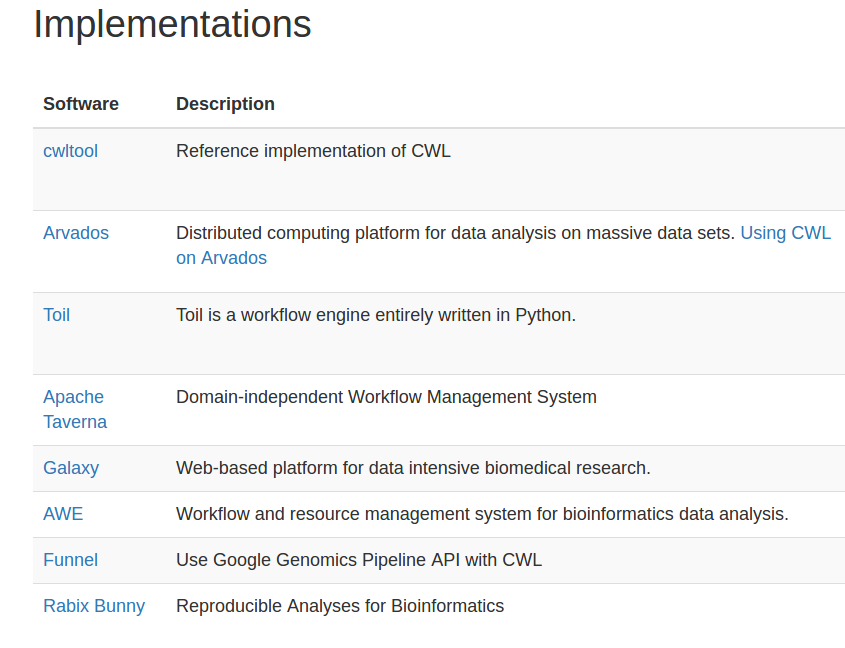



{
"@context" : [ "https://w3id.org/bundle/context" ],
"id" : "/",
"manifest" : [ "manifest.json" ],
"createdOn" : "2017-08-24T10:57:46.325Z",
"createdBy" : {
"uri" : "https://view.commonwl.org",
"name" : "Common Workflow Language Viewer"
},
"authoredBy" : [ {
"uri" : "mailto:peter.amstutz@curoverse.com",
"name" : "Peter Amstutz"
}, {
"uri" : "mailto:luka.stojanovic@sbgenomics.com",
"name" : "Luka Stojanovic"
}, {
"uri" : "mailto:crusoe@ucdavis.edu",
"name" : "Michael R. Crusoe"
}, {
"uri" : "mailto:porter@porter.st",
"name" : "Andrey Kartashov"
}, {
"uri" : "mailto:janko.simonovic@sbgenomics.com",
"name" : "Janko Simonovic"
} ],
"retrievedFrom" : "https://github.com/common-workflow-language/workflows/blob/lobstr-v1/workflows/lobSTR/",
"retrievedOn" : "2017-08-24T10:57:46.325Z",
"retrievedBy" : {
"uri" : "https://view.commonwl.org",
"name" : "Common Workflow Language Viewer"
},
"history" : [ "http:/git2prov.org/git2prov?giturl=https:/github.com/common-workflow-language/workflows.git&serialization=PROV-JSON" ],
"aggregates" : [ {
"uri" : "/workflow/tmp_2.fq",
"mediatype" : "application/octet-stream",
"createdOn" : "2017-08-24T10:57:46.923Z",
"authoredBy" : [ {
"uri" : "mailto:peter.amstutz@curoverse.com",
"name" : "Peter Amstutz"
} ],
"retrievedFrom" : "https://raw.githubusercontent.com/common-workflow-language/workflows/lobstr-v1/workflows/lobSTR/tmp_2.fq",
"retrievedBy" : {
"uri" : "https://view.commonwl.org",
"name" : "Common Workflow Language Viewer"
},
"bundledAs" : {
"uri" : "urn:uuid:61579f3e-63e6-49c2-b780-f67b2df461b7",
"folder" : "/workflow/"
}
}, {
"uri" : "/workflow/lobSTR-demo.json",
"mediatype" : "application/json",
"createdOn" : "2017-08-24T10:57:47.216Z",
"authoredBy" : [ {
"uri" : "mailto:peter.amstutz@curoverse.com",
"name" : "Peter Amstutz"
} ],
"retrievedFrom" : "https://raw.githubusercontent.com/common-workflow-language/workflows/lobstr-v1/workflows/lobSTR/lobSTR-demo.json",
"retrievedBy" : {
"uri" : "https://view.commonwl.org",
"name" : "Common Workflow Language Viewer"
},
"bundledAs" : {
"uri" : "urn:uuid:973caa0e-f3bd-45e8-8d29-70123bc8715a",
"folder" : "/workflow/"
}
}, {
"uri" : "/workflow/models/illumina_v3.pcrfree.stuttermodel",
"mediatype" : "application/octet-stream",
"createdOn" : "2017-08-24T10:57:47.239Z",
"authoredBy" : [ {
"uri" : "mailto:peter.amstutz@curoverse.com",
"name" : "Peter Amstutz"
} ],
"retrievedFrom" : "https://raw.githubusercontent.com/common-workflow-language/workflows/lobstr-v1/workflows/lobSTR/models/illumina_v3.pcrfree.stuttermodel",
"retrievedBy" : {
"uri" : "https://view.commonwl.org",
"name" : "Common Workflow Language Viewer"
},
"bundledAs" : {
"uri" : "urn:uuid:62bbcbea-f34f-463f-990d-6148f8ed5e5c",
"folder" : "/workflow/models/"
}
}, {
"uri" : "/workflow/models/illumina_v3.pcrfree.stepmodel",
"mediatype" : "application/octet-stream",
"createdOn" : "2017-08-24T10:57:47.266Z",
"authoredBy" : [ {
"uri" : "mailto:peter.amstutz@curoverse.com",
"name" : "Peter Amstutz"
} ],
"retrievedFrom" : "https://raw.githubusercontent.com/common-workflow-language/workflows/lobstr-v1/workflows/lobSTR/models/illumina_v3.pcrfree.stepmodel",
"retrievedBy" : {
"uri" : "https://view.commonwl.org",
"name" : "Common Workflow Language Viewer"
},
"bundledAs" : {
"uri" : "urn:uuid:03439ae7-cd94-42a3-b5fe-40bfff6882d8",
"folder" : "/workflow/models/"
}
}, {
"uri" : "/workflow/samtools-sort.cwl",
"mediatype" : "text/x-yaml",
"createdOn" : "2017-08-24T10:57:47.269Z",
"authoredBy" : [ {
"uri" : "mailto:luka.stojanovic@sbgenomics.com",
"name" : "Luka Stojanovic"
}, {
"uri" : "mailto:crusoe@ucdavis.edu",
"name" : "Michael R. Crusoe"
}, {
"uri" : "mailto:porter@porter.st",
"name" : "Andrey Kartashov"
}, {
"uri" : "mailto:peter.amstutz@curoverse.com",
"name" : "Peter Amstutz"
} ],
"retrievedFrom" : "https://raw.githubusercontent.com/common-workflow-language/workflows/lobstr-v1/workflows/lobSTR/samtools-sort.cwl",
"retrievedBy" : {
"uri" : "https://view.commonwl.org",
"name" : "Common Workflow Language Viewer"
},
"conformsTo" : "https://w3id.org/cwl/v1.0",
"bundledAs" : {
"uri" : "urn:uuid:2dc07859-efc2-4945-a95f-ba7815b68d07",
"folder" : "/workflow/"
}
}, {
"uri" : "/workflow/lobSTR-workflow.cwl",
"mediatype" : "text/x-yaml",
"createdOn" : "2017-08-24T10:57:47.42Z",
"authoredBy" : [ {
"uri" : "mailto:luka.stojanovic@sbgenomics.com",
"name" : "Luka Stojanovic"
}, {
"uri" : "mailto:crusoe@ucdavis.edu",
"name" : "Michael R. Crusoe"
}, {
"uri" : "mailto:peter.amstutz@curoverse.com",
"name" : "Peter Amstutz"
} ],
"retrievedFrom" : "https://raw.githubusercontent.com/common-workflow-language/workflows/lobstr-v1/workflows/lobSTR/lobSTR-workflow.cwl",
"retrievedBy" : {
"uri" : "https://view.commonwl.org",
"name" : "Common Workflow Language Viewer"
},
"conformsTo" : "https://w3id.org/cwl/v1.0",
"bundledAs" : {
"uri" : "urn:uuid:58bc1895-3460-46d6-91d7-fa1718d09631",
"folder" : "/workflow/"
}
}, {
"uri" : "/workflow/lobSTR-arvados-demo.json",
"mediatype" : "application/json",
"createdOn" : "2017-08-24T10:57:47.453Z",
"authoredBy" : [ {
"uri" : "mailto:peter.amstutz@curoverse.com",
"name" : "Peter Amstutz"
} ],
"retrievedFrom" : "https://raw.githubusercontent.com/common-workflow-language/workflows/lobstr-v1/workflows/lobSTR/lobSTR-arvados-demo.json",
"retrievedBy" : {
"uri" : "https://view.commonwl.org",
"name" : "Common Workflow Language Viewer"
},
"bundledAs" : {
"uri" : "urn:uuid:30c683bc-69fb-4d93-8dad-65b663783af5",
"folder" : "/workflow/"
}
}, {
"uri" : "/workflow/samtools-index.cwl",
"mediatype" : "text/x-yaml",
"createdOn" : "2017-08-24T10:57:47.458Z",
"authoredBy" : [ {
"uri" : "mailto:luka.stojanovic@sbgenomics.com",
"name" : "Luka Stojanovic"
}, {
"uri" : "mailto:crusoe@ucdavis.edu",
"name" : "Michael R. Crusoe"
}, {
"uri" : "mailto:porter@porter.st",
"name" : "Andrey Kartashov"
}, {
"uri" : "mailto:peter.amstutz@curoverse.com",
"name" : "Peter Amstutz"
} ],
"retrievedFrom" : "https://raw.githubusercontent.com/common-workflow-language/workflows/lobstr-v1/workflows/lobSTR/samtools-index.cwl",
"retrievedBy" : {
"uri" : "https://view.commonwl.org",
"name" : "Common Workflow Language Viewer"
},
"conformsTo" : "https://w3id.org/cwl/v1.0",
"bundledAs" : {
"uri" : "urn:uuid:8235d3f8-6927-4f73-b160-8521838a1cbb",
"folder" : "/workflow/"
}
}, {
"uri" : "/workflow/lobSTR-tool.cwl",
"mediatype" : "text/x-yaml",
"createdOn" : "2017-08-24T10:57:47.476Z",
"authoredBy" : [ {
"uri" : "mailto:luka.stojanovic@sbgenomics.com",
"name" : "Luka Stojanovic"
}, {
"uri" : "mailto:crusoe@ucdavis.edu",
"name" : "Michael R. Crusoe"
}, {
"uri" : "mailto:peter.amstutz@curoverse.com",
"name" : "Peter Amstutz"
} ],
"retrievedFrom" : "https://raw.githubusercontent.com/common-workflow-language/workflows/lobstr-v1/workflows/lobSTR/lobSTR-tool.cwl",
"retrievedBy" : {
"uri" : "https://view.commonwl.org",
"name" : "Common Workflow Language Viewer"
},
"conformsTo" : "https://w3id.org/cwl/v1.0",
"bundledAs" : {
"uri" : "urn:uuid:7fa6fbe4-1fc5-4cb5-9c1a-56b96c5f7aaf",
"folder" : "/workflow/"
}
}, {
"uri" : "/workflow/allelotype.cwl",
"mediatype" : "text/x-yaml",
"createdOn" : "2017-08-24T10:57:47.537Z",
"authoredBy" : [ {
"uri" : "mailto:luka.stojanovic@sbgenomics.com",
"name" : "Luka Stojanovic"
}, {
"uri" : "mailto:janko.simonovic@sbgenomics.com",
"name" : "Janko Simonovic"
}, {
"uri" : "mailto:crusoe@ucdavis.edu",
"name" : "Michael R. Crusoe"
}, {
"uri" : "mailto:peter.amstutz@curoverse.com",
"name" : "Peter Amstutz"
} ],
"retrievedFrom" : "https://raw.githubusercontent.com/common-workflow-language/workflows/lobstr-v1/workflows/lobSTR/allelotype.cwl",
"retrievedBy" : {
"uri" : "https://view.commonwl.org",
"name" : "Common Workflow Language Viewer"
},
"conformsTo" : "https://w3id.org/cwl/v1.0",
"bundledAs" : {
"uri" : "urn:uuid:3706bd2f-e53f-431d-b32a-deb661d9b292",
"folder" : "/workflow/"
}
}, {
"uri" : "/workflow/README",
"mediatype" : "application/octet-stream",
"createdOn" : "2017-08-24T10:57:47.555Z",
"authoredBy" : [ {
"uri" : "mailto:crusoe@ucdavis.edu",
"name" : "Michael R. Crusoe"
}, {
"uri" : "mailto:peter.amstutz@curoverse.com",
"name" : "Peter Amstutz"
} ],
"retrievedFrom" : "https://raw.githubusercontent.com/common-workflow-language/workflows/lobstr-v1/workflows/lobSTR/README",
"retrievedBy" : {
"uri" : "https://view.commonwl.org",
"name" : "Common Workflow Language Viewer"
},
"bundledAs" : {
"uri" : "urn:uuid:ed54c4d6-c585-4dc9-b7bc-0cf299e20b91",
"folder" : "/workflow/"
}
}, {
"uri" : "/workflow/tmp_1.fq",
"mediatype" : "application/octet-stream",
"createdOn" : "2017-08-24T10:57:47.738Z",
"authoredBy" : [ {
"uri" : "mailto:peter.amstutz@curoverse.com",
"name" : "Peter Amstutz"
} ],
"retrievedFrom" : "https://raw.githubusercontent.com/common-workflow-language/workflows/lobstr-v1/workflows/lobSTR/tmp_1.fq",
"retrievedBy" : {
"uri" : "https://view.commonwl.org",
"name" : "Common Workflow Language Viewer"
},
"bundledAs" : {
"uri" : "urn:uuid:5d431f81-ad0b-4acf-903a-9d5aa03b04df",
"folder" : "/workflow/"
}
}, {
"uri" : "/visualisation.png",
"mediatype" : "image/png",
"createdOn" : "2017-08-24T10:57:47.801Z",
"retrievedFrom" : "https://view.commonwl.org/graph/png/github.com/common-workflow-language/workflows/blob/lobstr-v1/workflows/lobSTR/lobSTR-workflow.cwl",
"bundledAs" : {
"uri" : "urn:uuid:ff9ace37-e76c-49f8-8d36-60f11ff6d257",
"folder" : "/"
}
}, {
"uri" : "/visualisation.svg",
"mediatype" : "image/svg+xml",
"createdOn" : "2017-08-24T10:57:47.821Z",
"retrievedFrom" : "https://view.commonwl.org/graph/svg/github.com/common-workflow-language/workflows/blob/lobstr-v1/workflows/lobSTR/lobSTR-workflow.cwl",
"bundledAs" : {
"uri" : "urn:uuid:a6cfb437-8818-4ab2-9081-efc74c5109e8",
"folder" : "/"
}
} ],
"annotations" : [ {
"uri" : "urn:uuid:9f602fff-b280-41c5-9590-ab95a49c85ad",
"about" : "/",
"content" : "annotations/merged.cwl"
}, {
"uri" : "urn:uuid:0ce4b727-ff61-4534-9afb-e3d676d2782d",
"about" : "/",
"content" : "annotations/workflow.ttl"
} ]
}

#!/usr/bin/env cwl-runner
cwlVersion: v1.0
class: Workflow
label: "Hello World"
doc: "Outputs a message using echo"
inputs: []
outputs:
response:
outputSource: step0/response
type: File
steps:
step0:
run:
class: CommandLineTool
inputs:
message:
type: string
doc: "The message to print"
default: "Hello World"
inputBinding:
position: 1
baseCommand: echo
stdout: response.txt
outputs:
response:
type: stdout
in: []
out: [response]

Who is using Research Objects?












Workshop on
Research Objects (RO2019)
At IEEE eScience 2019,
San Diego, CA
2019-06-24 Abstracts due
2019-09-24 RO2019 workshop at IEEE eScience 2019
Submitted abstracts and articles can be in a range of open formats (e.g. HTML, ePub) and are particularly encouraged to be submitted in a FAIR research data packing format.
..extra slides


Identifying intermediate data
Output 1B file is also Input 2C and Input 3D downstream
Simple filenames -> duplications
./data/step1/outputB.txt
./data/step2/inputC.txt
./data/step3/inputD.txt
Content-adressable
SHA-256 hash of bytes as filename:
./data/51/51fb8af0c4ae0422fbe88340d91880ecb9d7537cf57339c1cf1256b7ca58f32d
RFC6920 URI as global identifier:
nih:sha-256;51fb8af0c4ae0422fbe88340d91880ecb9d7537cf57339c1cf1256b7ca58f32d