Boolean Network Synthesis From Experimental Data and Prior Knowledge With Formal Guarantees
Samuel Pastva
MOTIVATION
Project
IST
Single Cell RNA Sequencing
Single Cell RNA
Sequencing
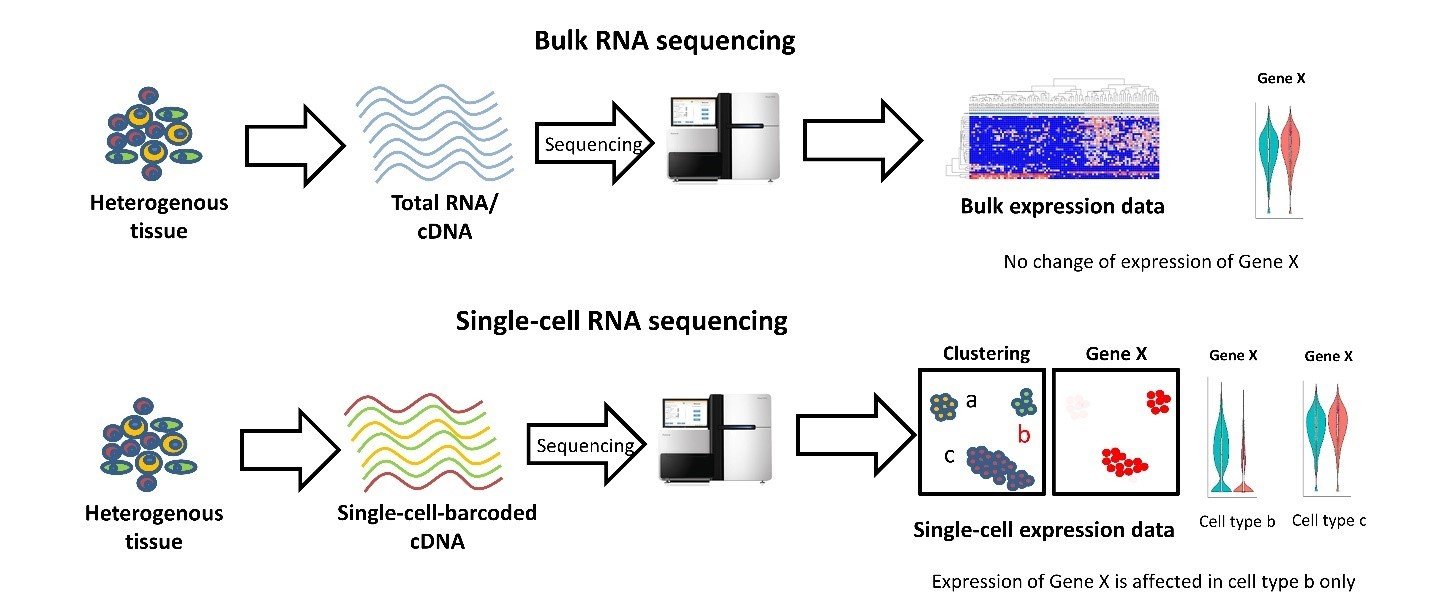
Complete snapshot of gene expression (active cell "logic")
Boolean Networks
Boolean Networks
Logical Modelling
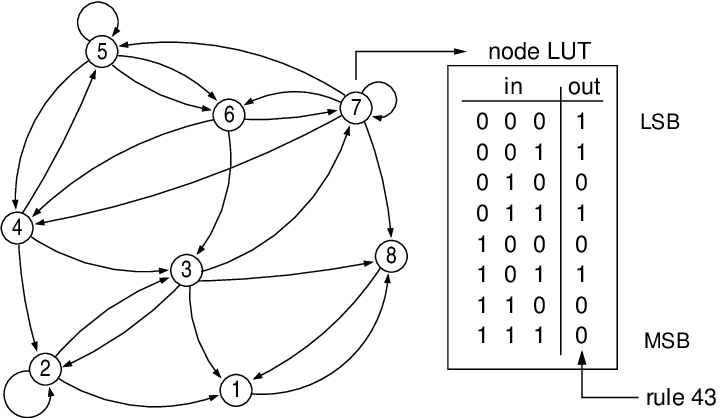
Boolean Networks
Logical Modelling
-
Qualitative approach is computationally less demanding than e.g. ODEs, but still powerful
-
White-box models: explainable, composable, generalisable to different organisms
Boolean Networks
Logical Modelling
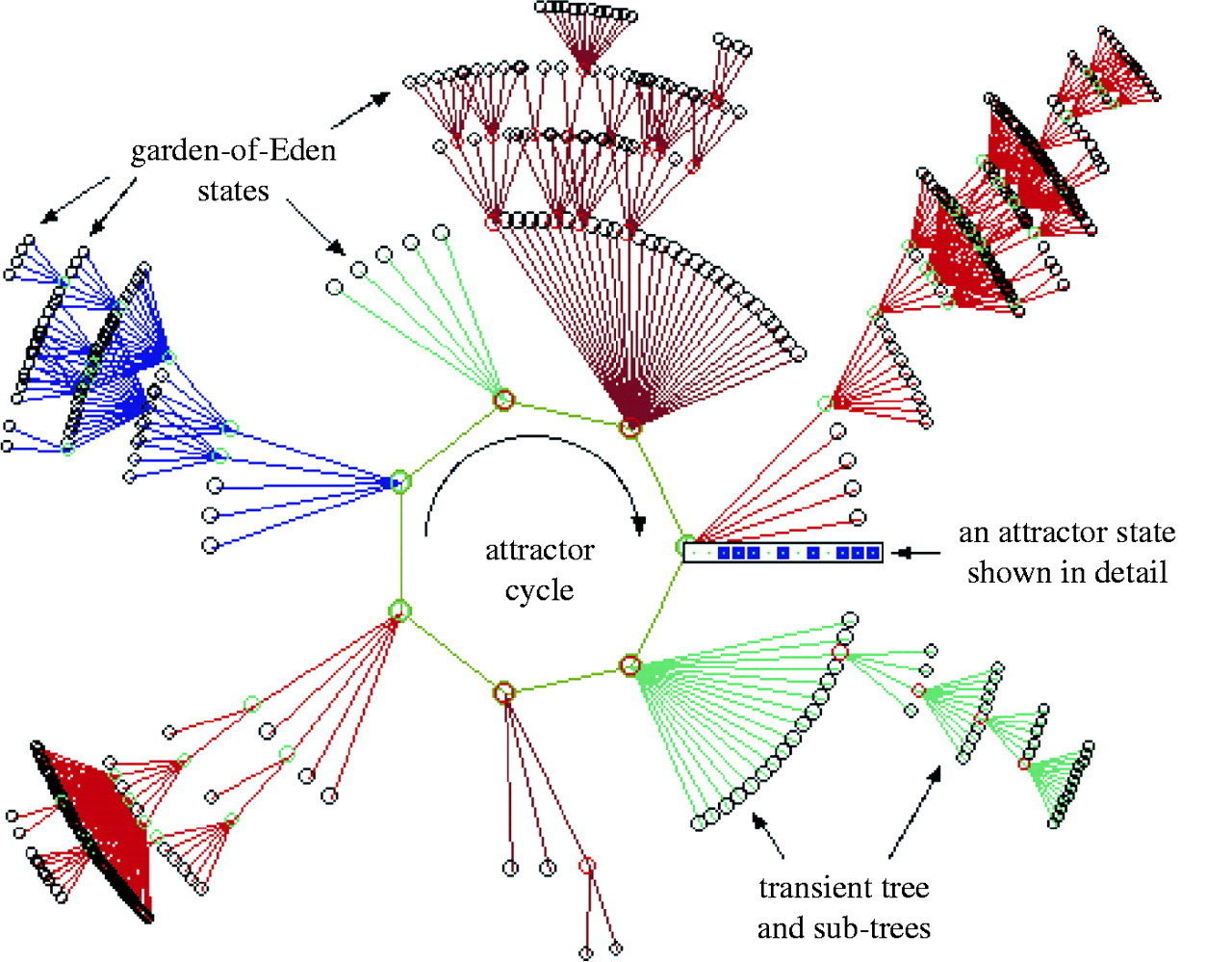
State-space of a synchronous Boolean network
Boolean Networks
Logical Modelling
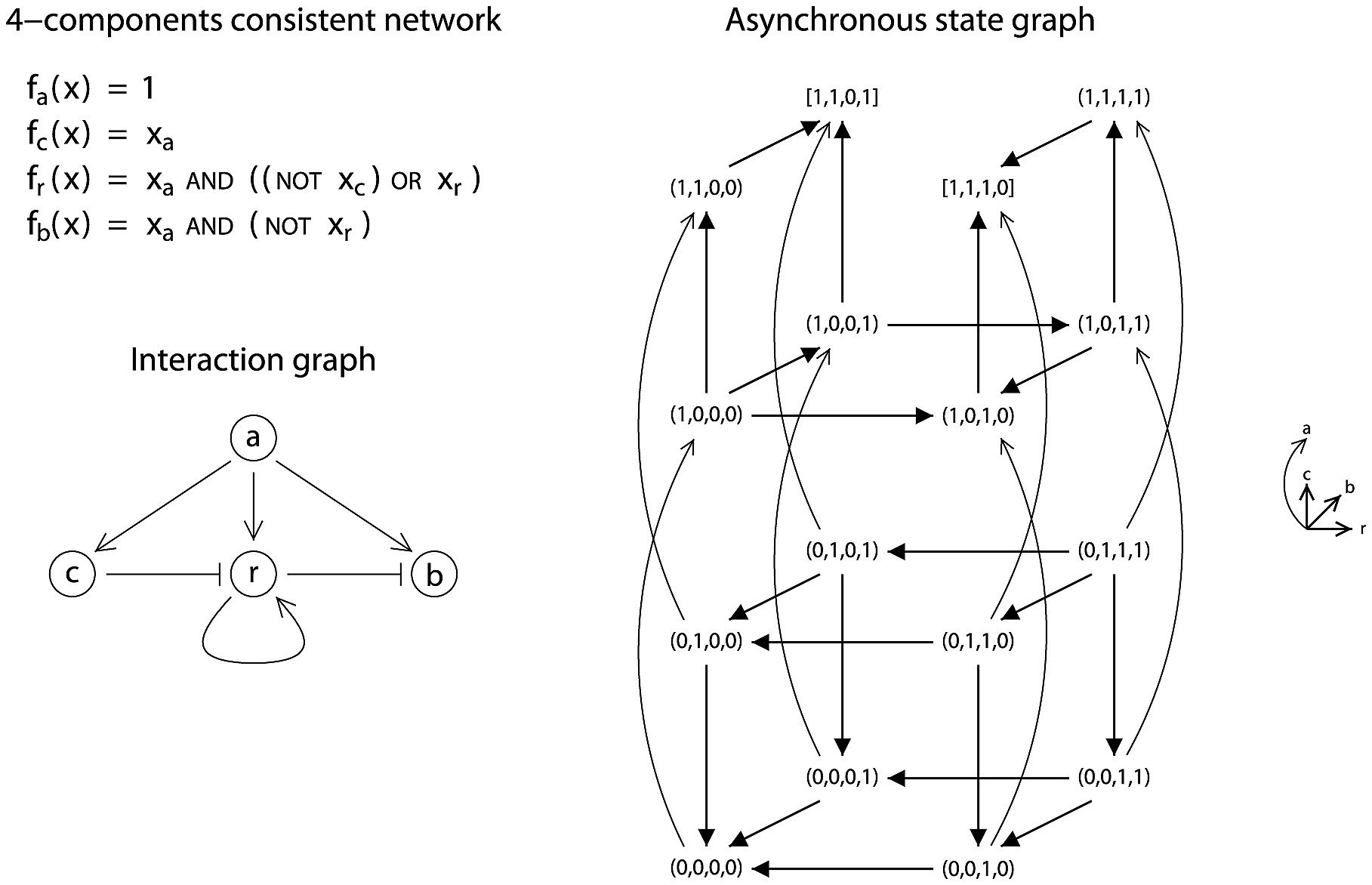
State-space of an asynchronous Boolean network
"Model gap"
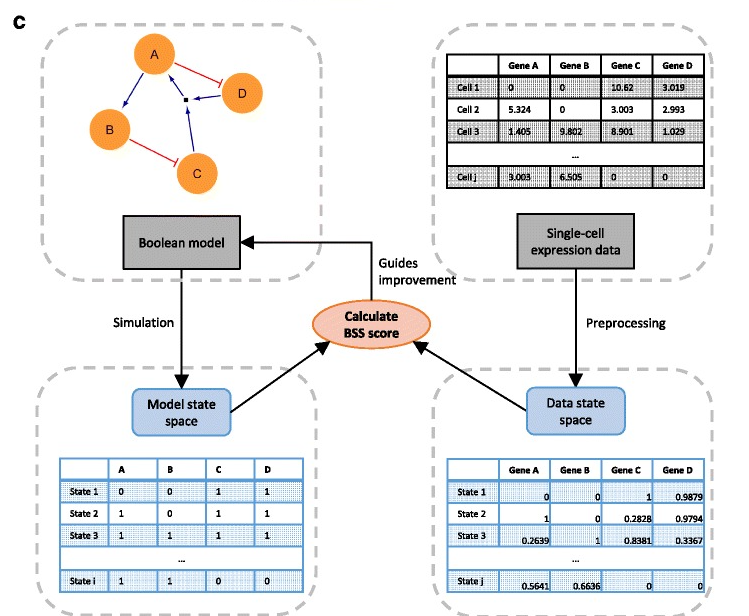
Boolean network
synthesis
Boolean network
synthesis
-
Cannot use single cell data (pre-dates single cell genomics)
-
Fits only synchronous Boolean networks
-
Only considers static properties (update function tables and their fixed-points)
-
Generates a single "correct" network instead of conservatively communicating uncertainty: How specific and sensitive is the result?
Project
Workflow which accounts for different types of partial information (literature, data, assumptions):
- Network structure (prior regulatory knowledge)
- Static attributes (local properties of update functions)
- Dynamic attributes (global properties of network dynamics)
Project
Proper communication of uncertainty through gradual refinement of "partial" Boolean networks.
Partial Boolean Network
Refinement
Rejected Networks
Necessary and sufficient conditions for valid networks
IST
prof. Thomas Henzinger
- Expert both in formal methods and systems biology
- Logical modelling not explored at the moment
- Inherently interdisciplinary project with application in biological research
IST has multiple groups investigating gene regulation
- Source of data and knowledge for the project
- Benefit to these groups in the form of new/improved models and tooling