Uncovering A Digital Twin of the Maturing Infant Microbiome

Sizemore, Nicholas, Kaitlyn Oliphant, Ruolin Zheng, Camilia R. Martin, Erika C. Claud, and Ishanu Chattopadhyay. "A digital twin of the infant microbiome to predict neurodevelopmental deficits." Science Advances 10, no. 15 (2024): eadj0400.

ishanu chattopadhyay
Nicholas Sizemore
Kaitlyn Oliphant
Erika Claud
Camilia Martin
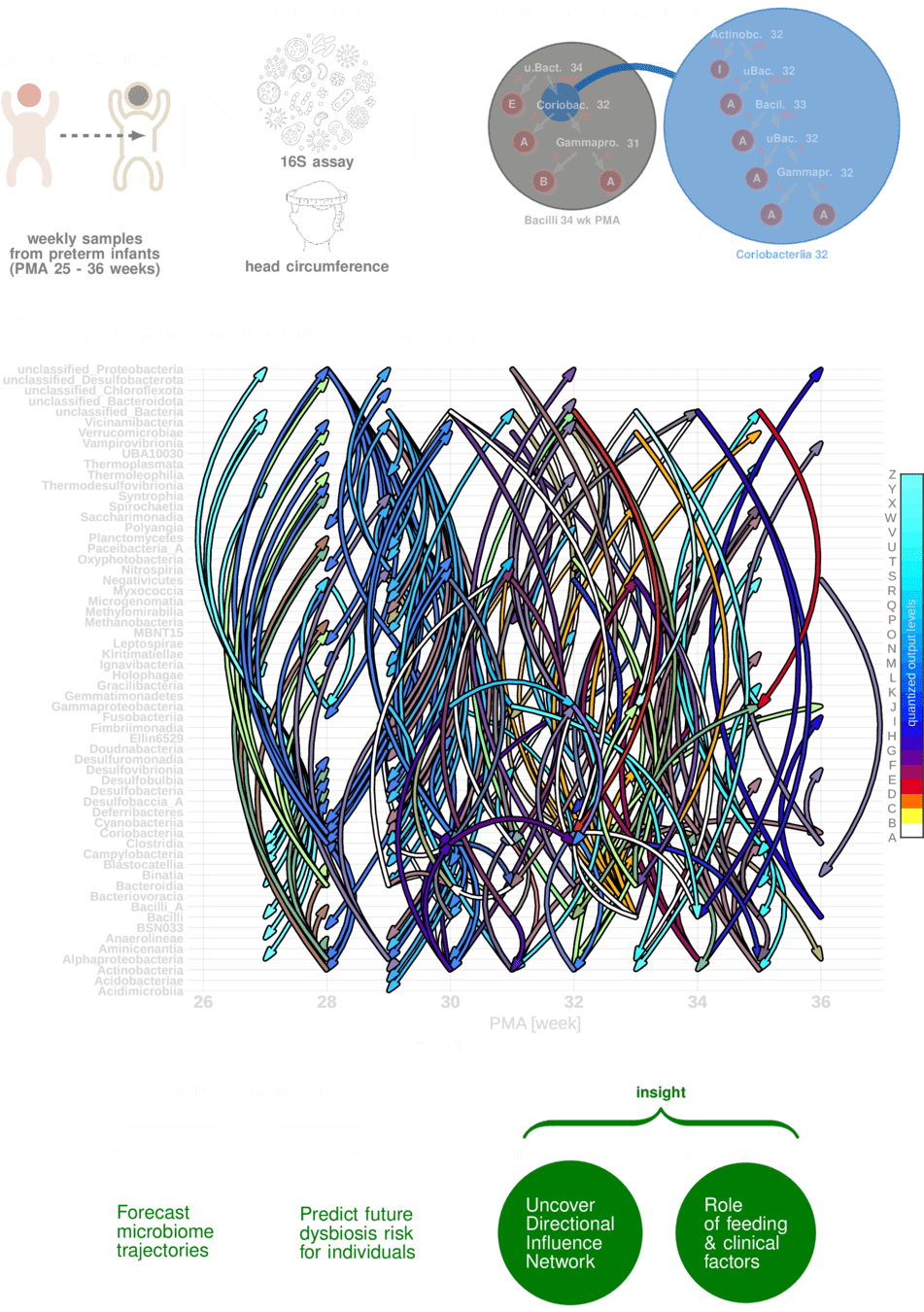
THE PROBLEM
Can microbial assay from gut actionably
pre-empt developmental deficit?
Assuming a 1000 species ecosystem, and 1 successful experiment every day to discern a single two-way relationship, we would need 1,368 years to go through all possibilities. If we look for 3 way interactions, we would need 454,844 years

The Need for AI-accelerated Scale-up
Digital Twin Inference
- Consider ecosystem of many inbitants
- The abundunce of each entity is a variable
- The variables interact in complex ways
- We are going to model this cross-talk
The QNet Framework
Lets see an example of this approach

Chattopadhyay, Ishanu, Kevin Wu, Jin Li, and Aaron Esser-Kahn. "Emergenet: Fast Scalable Pandemic Risk Assessment of Influenza A Strains Circulating In Non-human Hosts." (2023). Under Review in Nature
PREEMPT
Consider how mutations in a viral genome "cross-talk"
We can infer these co-depndecies using the same approach
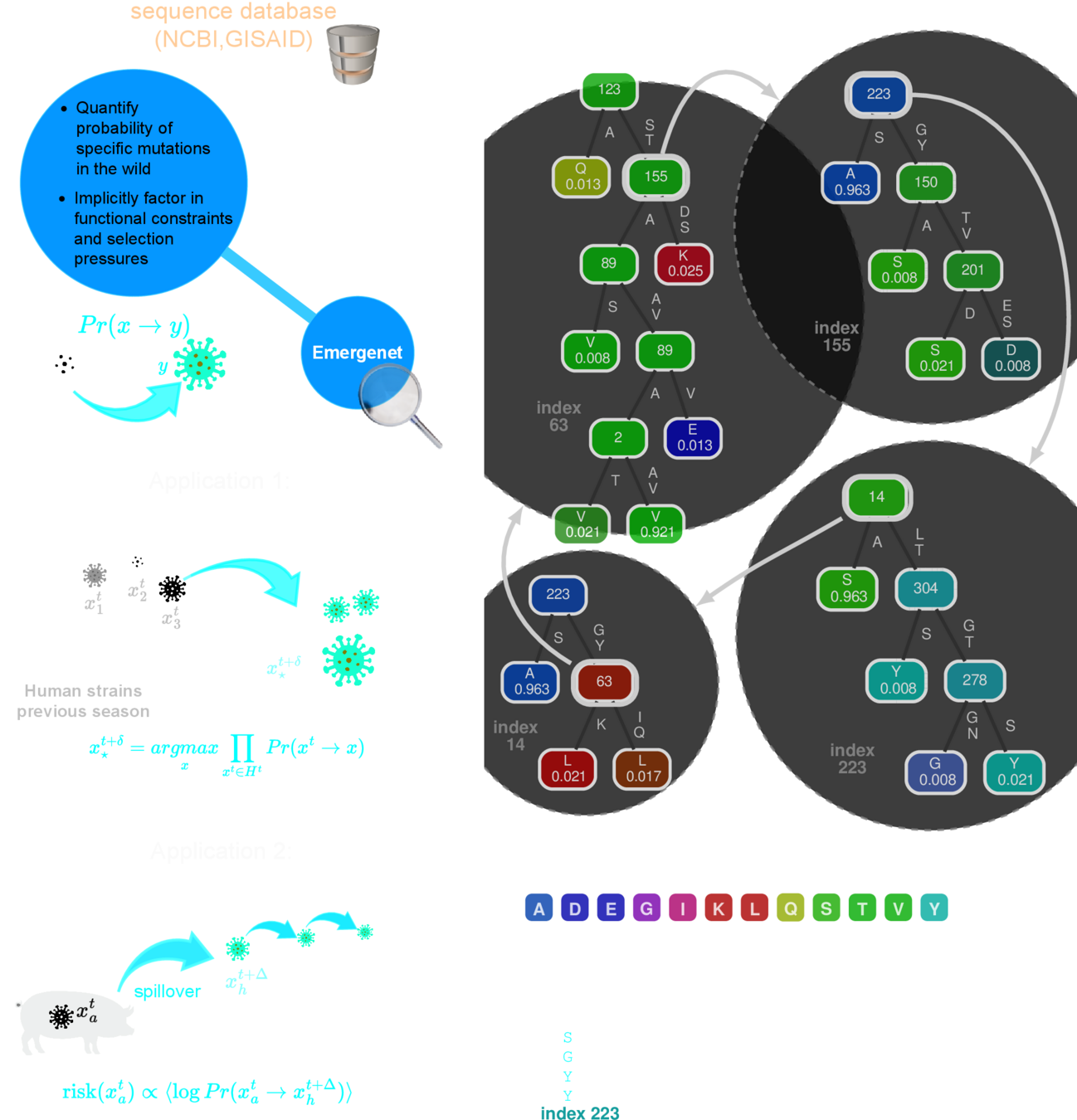


Q-Net
recursive forest
Each tree (model for variable) uses other variables as features
An Emergent Recursive Model
Lets go back to the Microbiome Problem
<class>_<observation_time>
<actinobacteria>_<30wk>
<clostridia>_<28wk>
construct qnet


Q-net inferred with typical patients

Q-net inferred with patients with neurodevelopmental deficit
We infer two digital twins: One for typical development, one for sub-optimal development
completely uninformative state
observed
state
Q-net inferred with typical patients
Q-net inferred with patients with neurodevelopmental deficit
Then we need to decide if an observed profile was generated by the typical model or the sub-optimal model
Risk of Time-stamped Microbial Profile to lead to Developmental Deficit
How different are the individual estimators for typical and deficit models?
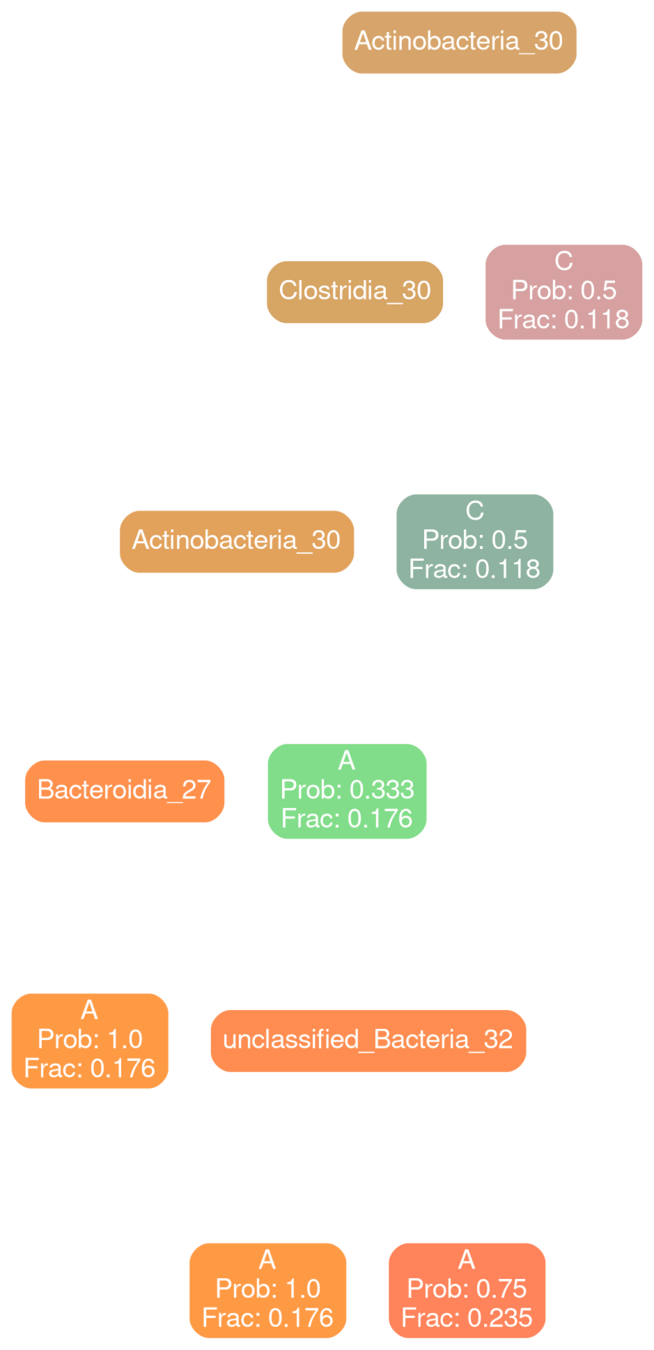
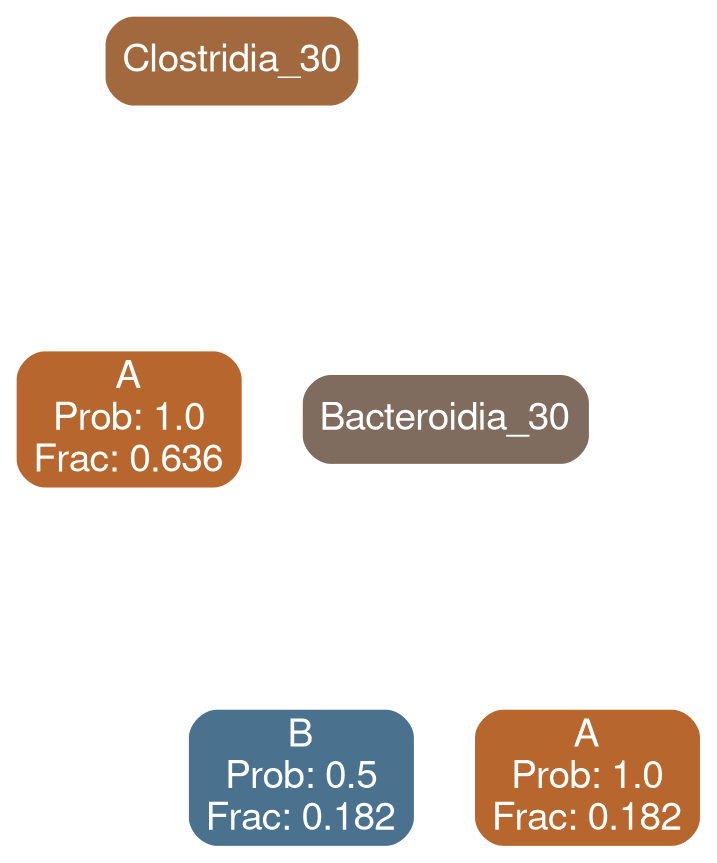
Bacilli 30
typical
deficit
Coriobacteria 32
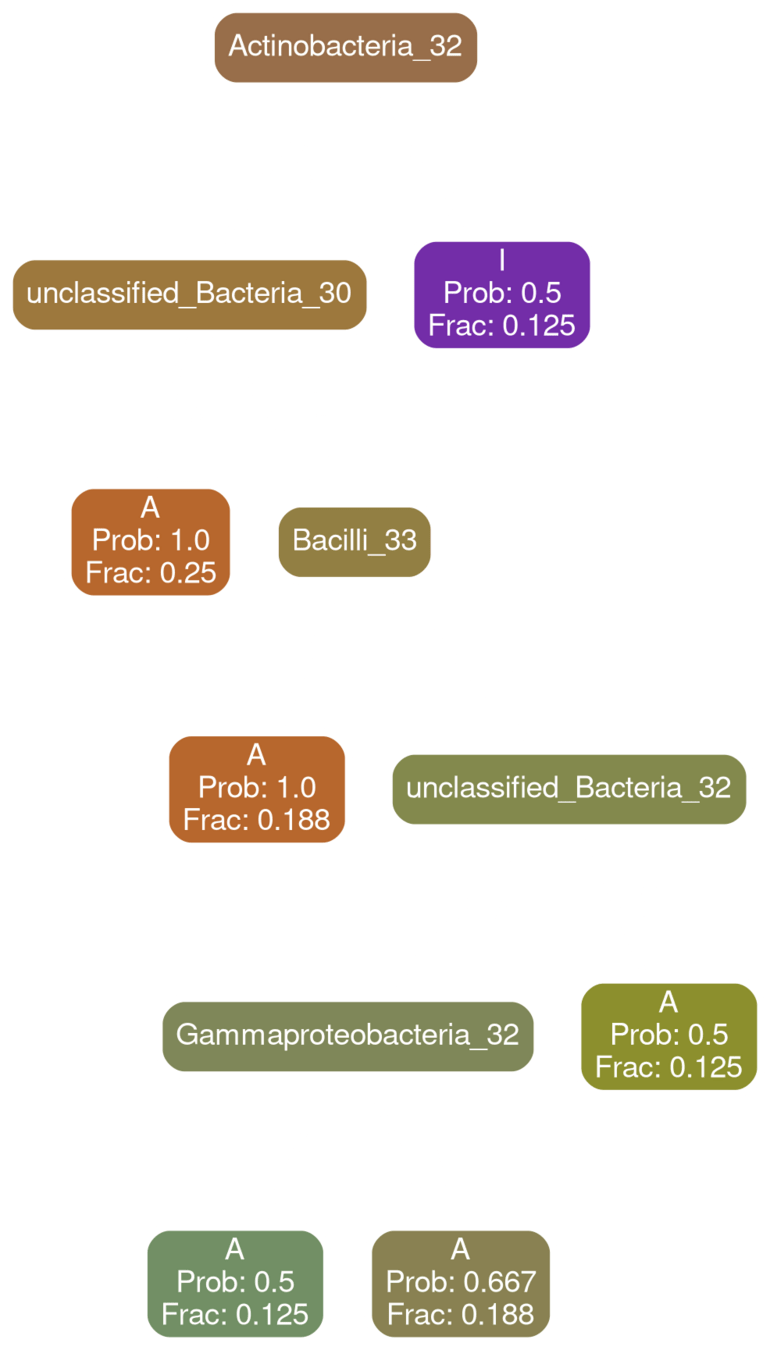
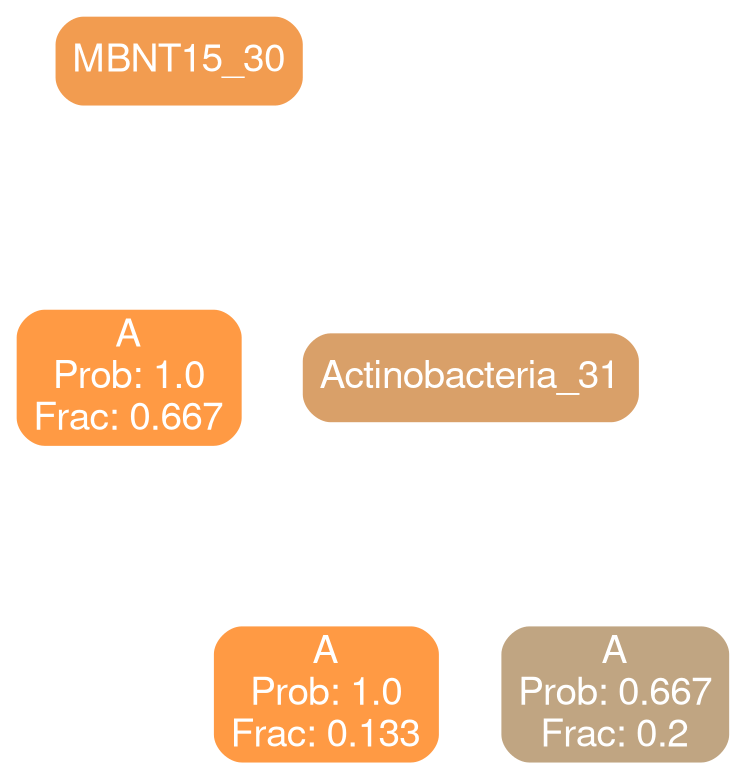
typical
deficit
Gammaproteobacteria 32
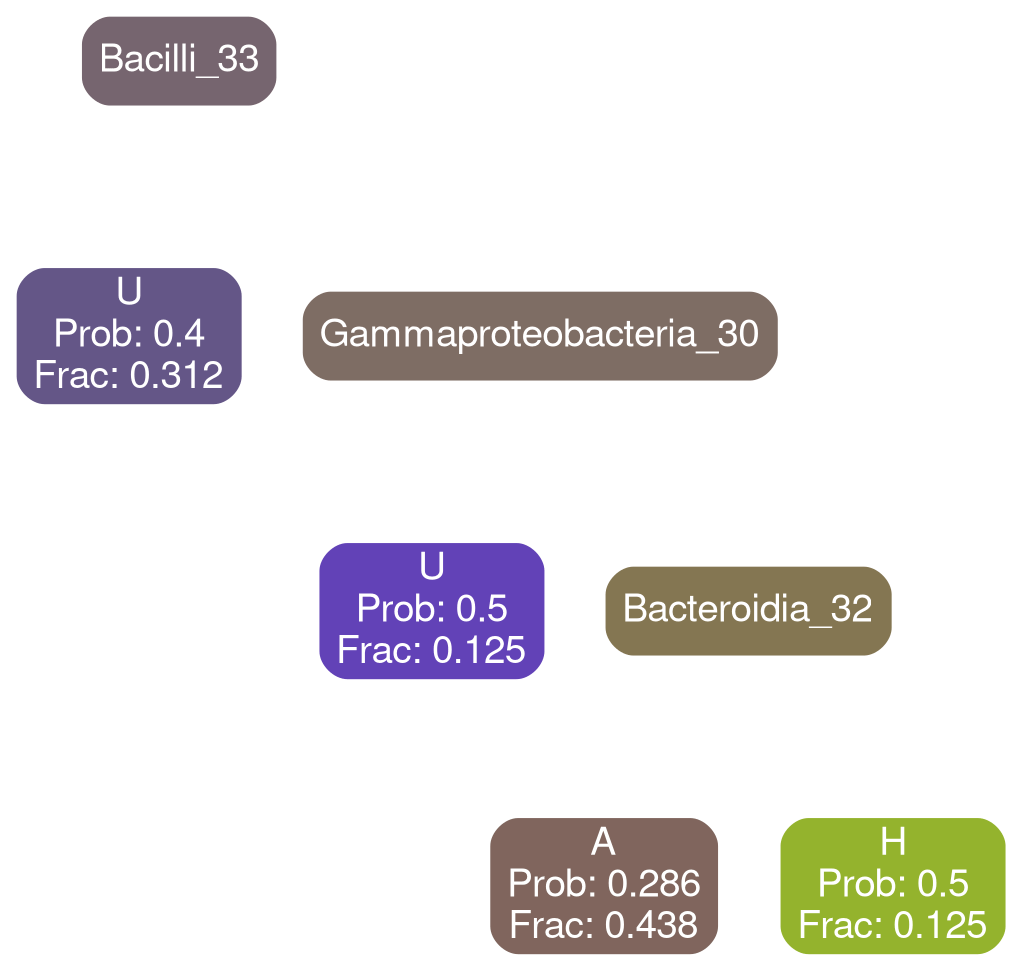
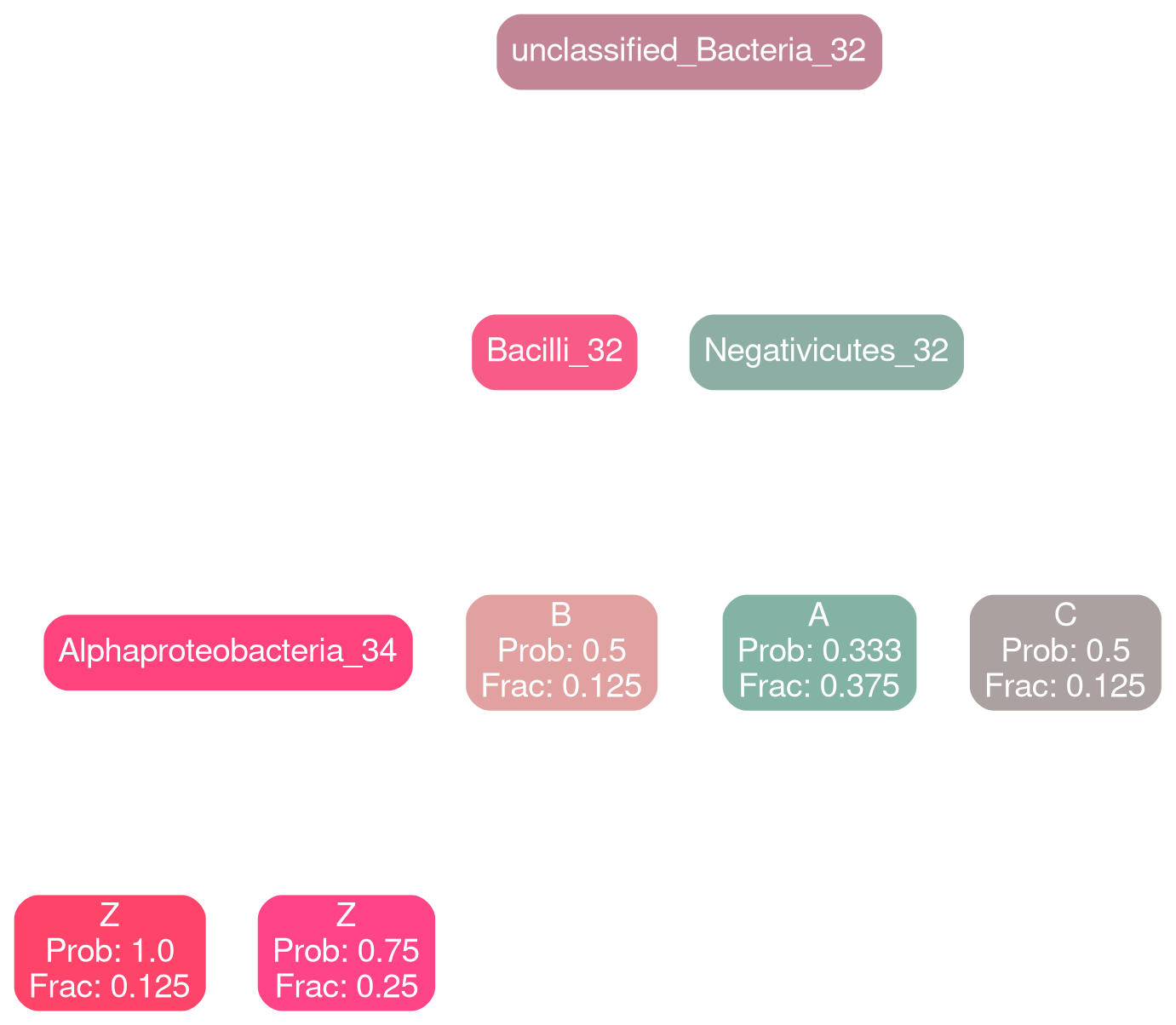
typical
deficit
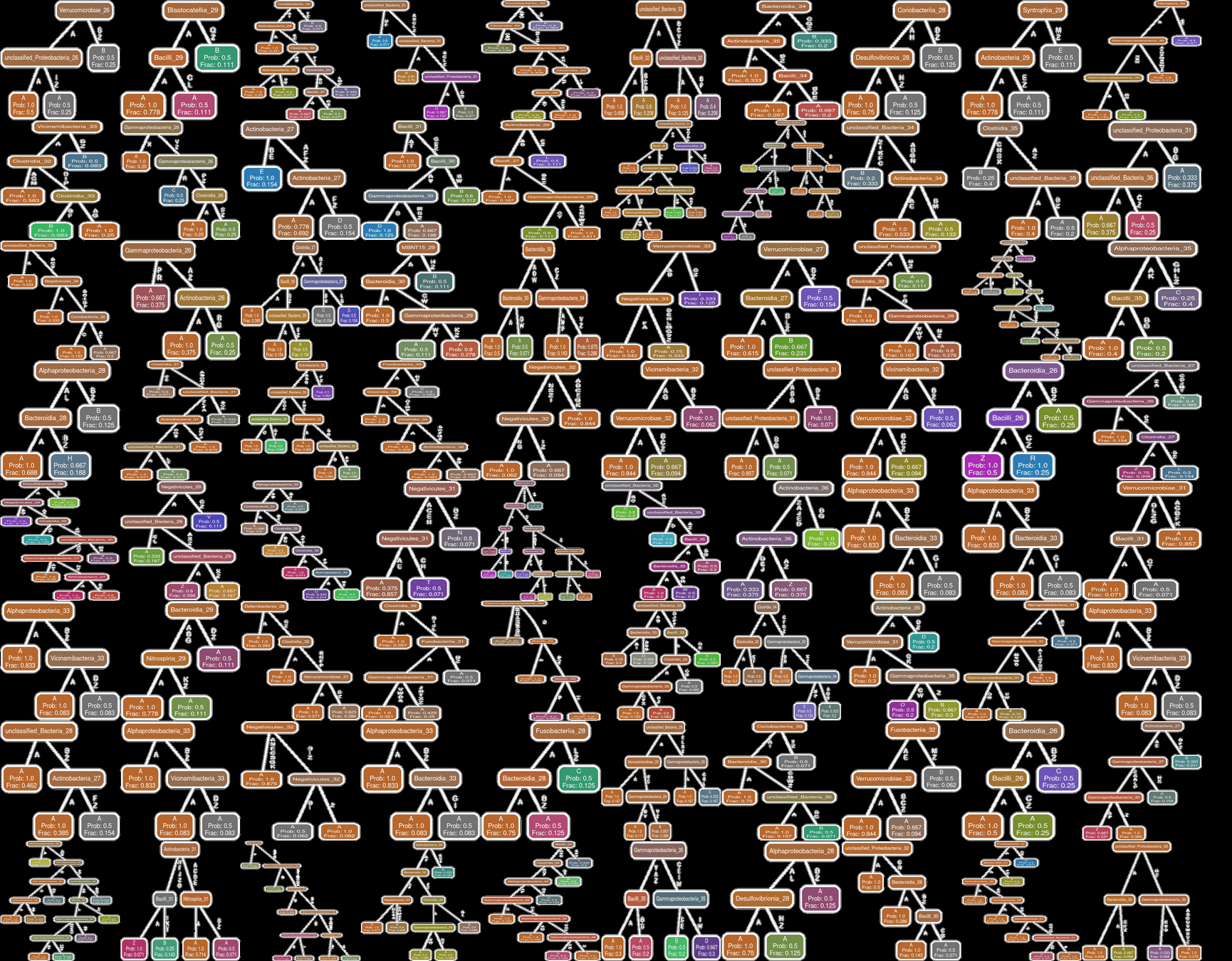
All Patients
Feeding Variables added
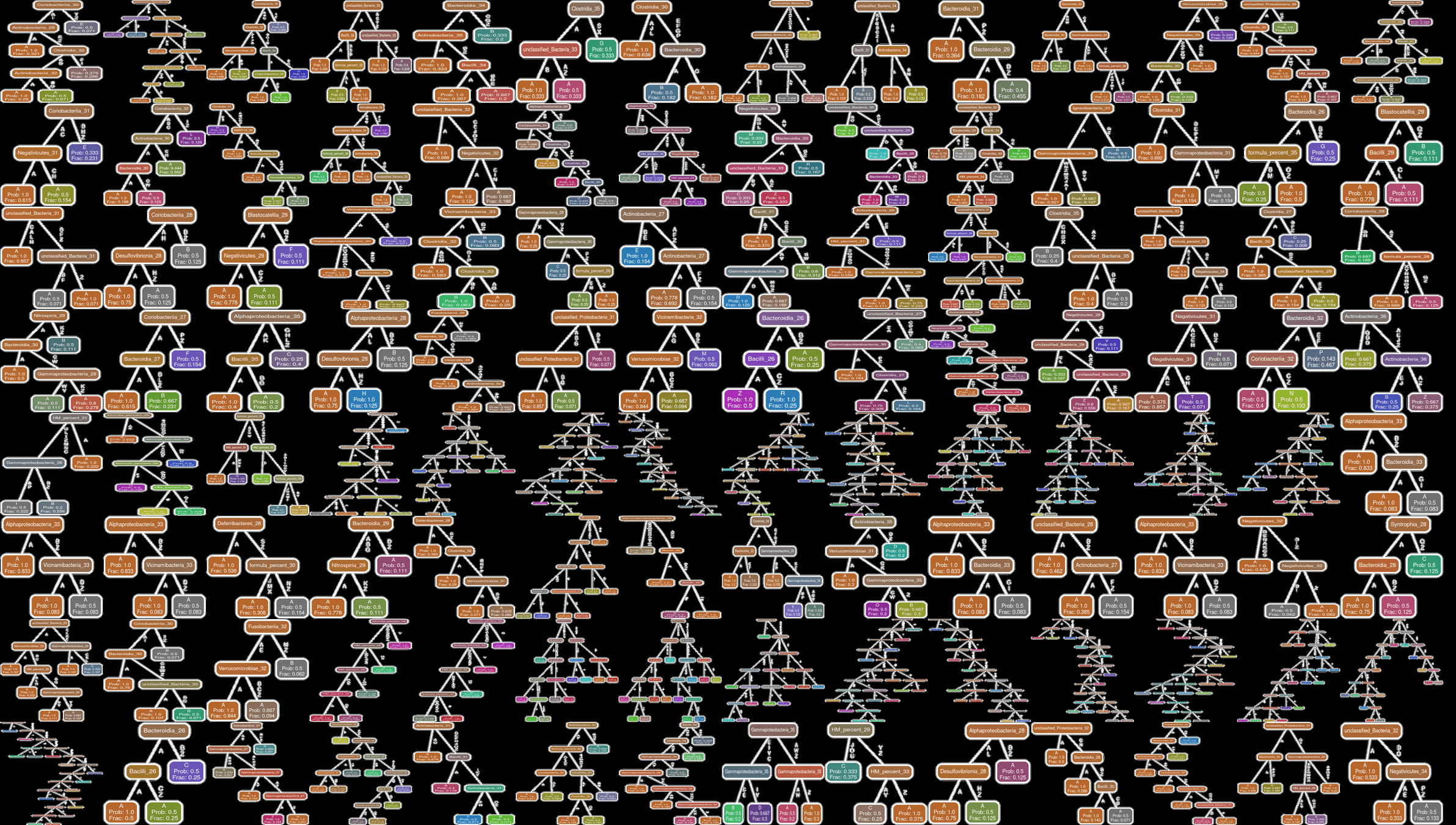
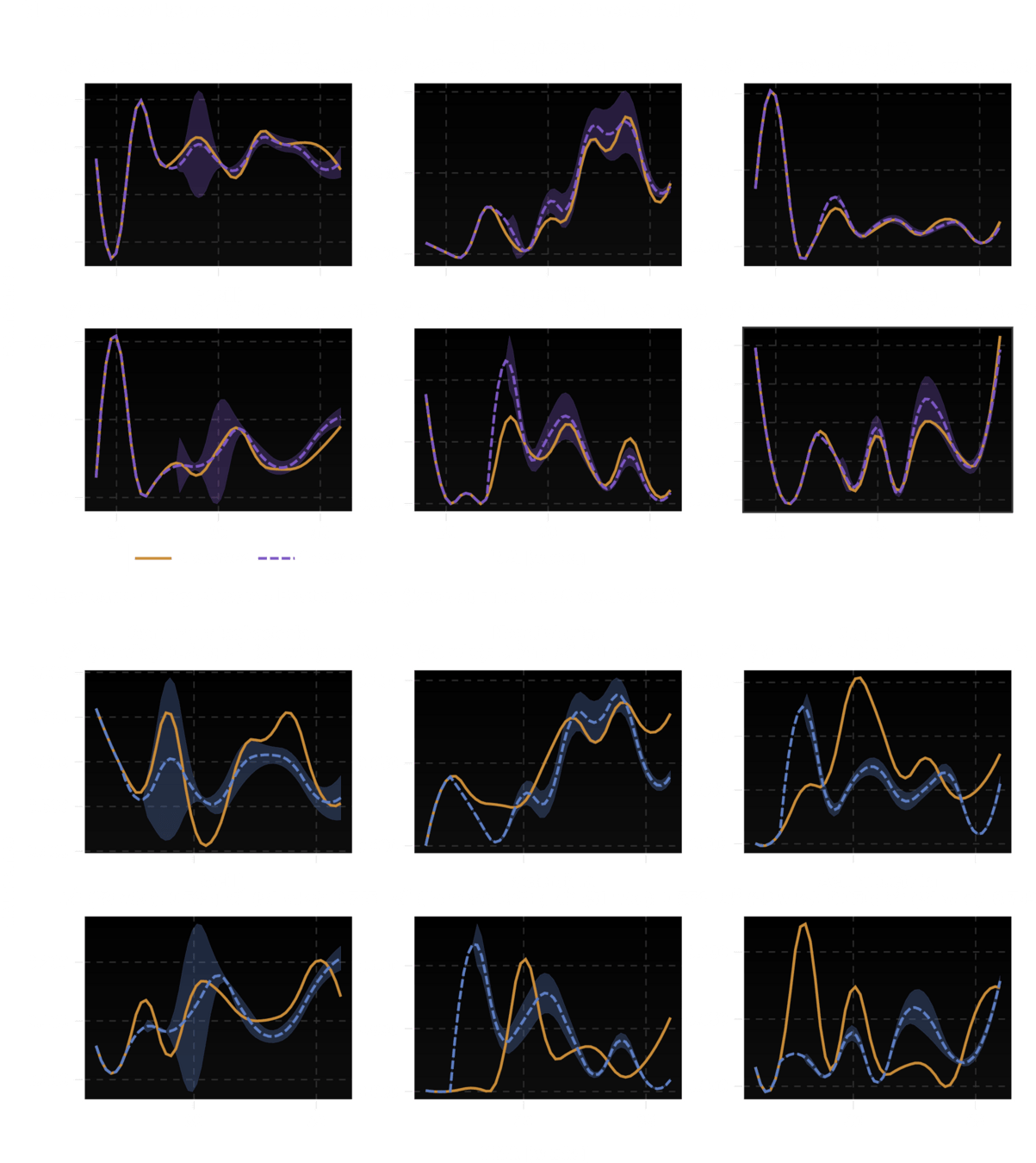
Ability to "fill in" missing data is equivalent to making trajectory forecasts
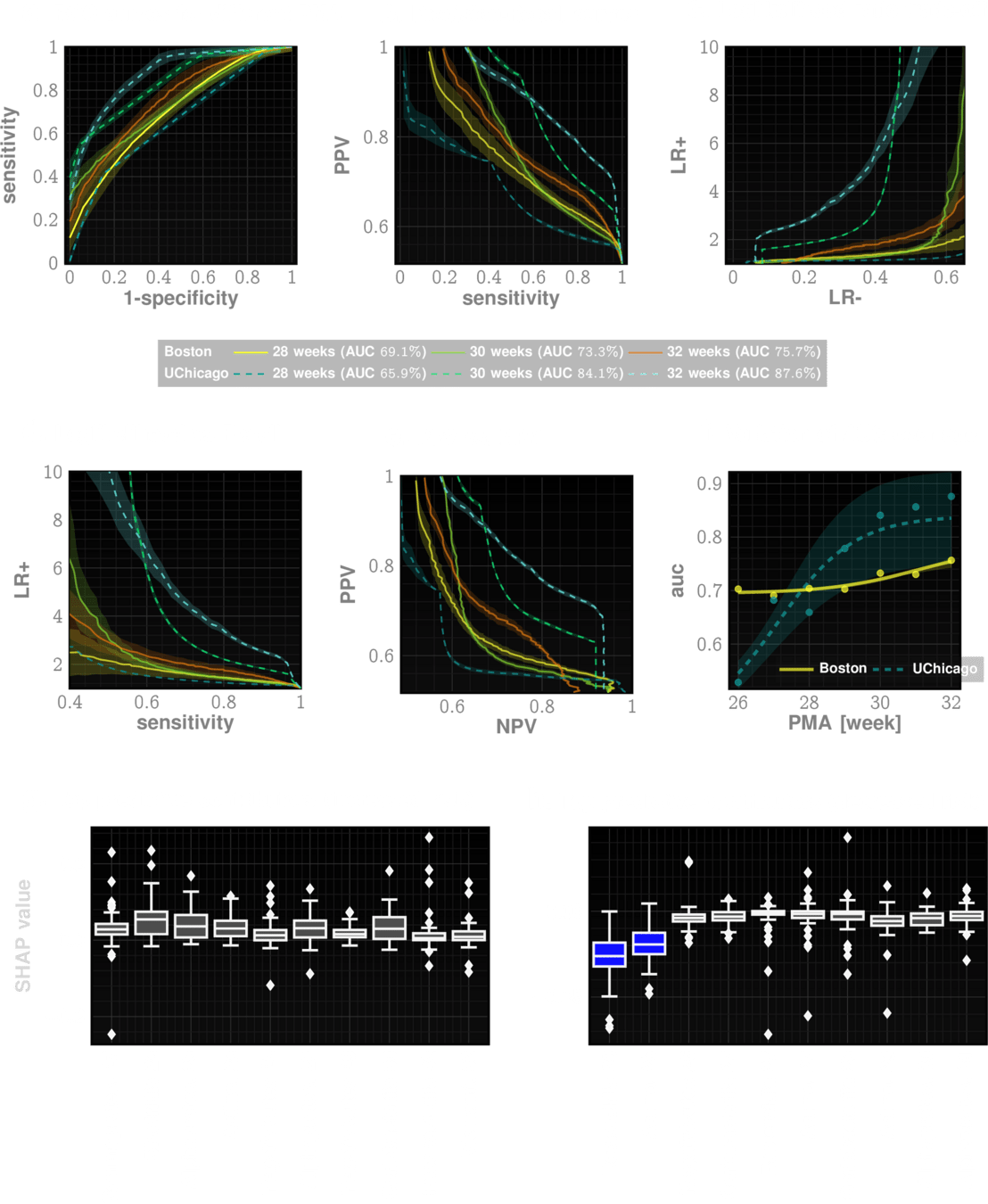



Our risk measure is highly predictive and actionable

Which entities are most predictive?
Just add those microbes back?
No! Our results indicate that supplantations need to be patient specific
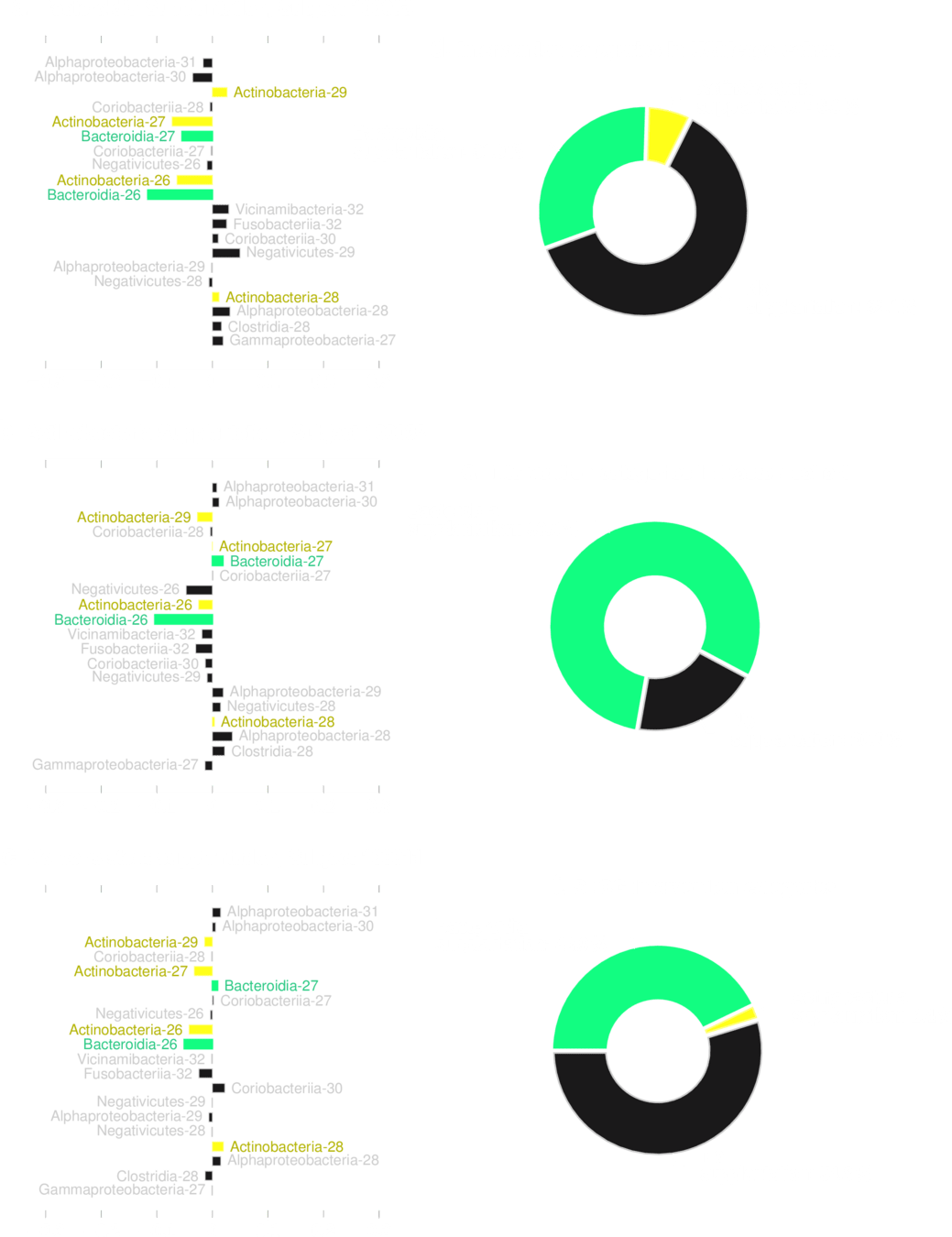
No transplantation is guaranteed to work reliably
Predicted to reduce
risk reliably
Predicted to reduce
risk reliably

Supplantation MUST be bacteroidia

Supplantation MUST be Actinobacteria

No risk-decreasing supplantation
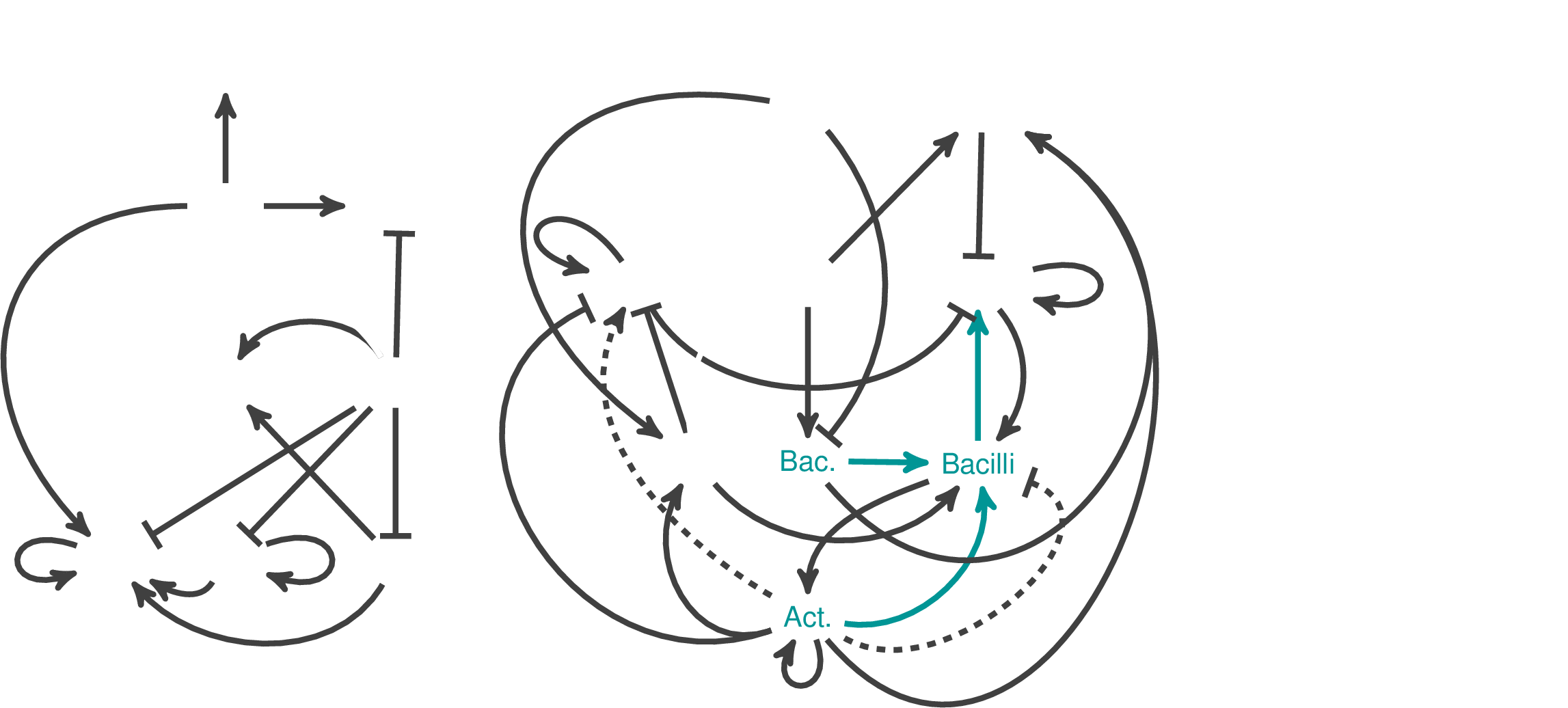
Network Interpretations? We see clear differences between two cases
Typical
Deficit
Future
Answer the question: "what is a healthy microbiome?"
Explicit supplantation profiles that are tuned to individual ecosystems
Bioreactor experiments