
Future Algorithms and Future Leaders in AI for Medicine:
From Test-free Screening, to
Digital twins in Medicine to Training
The Next Generation of BioAI-Experts
Ishanu Chattopadhyay, PhD
Assistant Professor of Medicine
University of Chicago
ishanu@uchicago.edu
first wave
rule-based systems
second wave
Big Data / ML / Deep Learning
recognize patterns, make predictions, struggle on tasks not trained for
third wave
contextual reasoning, generlizable, towards true intelligence



Artificial Intelligence
1940 - 2024
Medicine is poised to enter a transformative era, ushered by the emergence of sophisticated Artificial Intelligence (AI) models.
Enable more holistic approaches to medicine, where predictive patterns can be rapidly recognized and exploited

Future

mathematics

computer science

social science

medicine

AI/ML learning theory and applications
Complex systems
Implication of AI in Future of Societay


The Laboratory for Zero Knowledge Discovery

collaborators
Alex Leow
Psychiatry UIC
Anna Podolanczuk, Pulmonary Care, Weill Cornell
Gary Hunninghake, Pulmonary C, Harvard




Robert Gibbons, Bio-statistics

Daniel Rubins, Anesthesia and Critical Care

Peter Smith, Pediatrics

Michael Msall Pediatrics

Fernando Martinez, Pulmonary Critical Care, Weill Cornell

James Mastrianni, Neurology




James Evans, sociology

Erika Claud, Pediatrics
Aaron Esser-Kahn Molecular Engineering

David Llewellyn
University of Exeter
Kenneth Rockwood
Dalhousie University
Andrew Limper Mayo Clinic

David Scwartz
University of Colorado, Pulmonary Genetics

zed.uchicago.edu
Department of Pediatrics
UChicago
Department of Neurology & The Memory Center
UChicago
Department of Psychiatry
UChicago
Pulmonary Critical Care, Weill Cornell
Department of Anesthesia and Critical Care
UChicago
Center for Health Statistics
UChicago
Pulmonary Critical Care, Harvard Medical School
Department of Psychiatry
UIC
Demon Network, Exeter, Alan Turing Institute, UK
Dalhousie University, Canada
Pritzker School of Molecular ENgineering
Social Science
UChicago
Pulmomary and Genomics University of Colorado Anschutz
Los Alamos National Laboratory
collaboratorions







zed.uchicago.edu
D3M (I2O)
PAI (DSO)
PREEMPT (BTO)
YFA (DSO)
NIA
Nature Medicine
Nature Human Behavior
Nature Commun-ication
Science Advances
(3)
PNAS
JAMA
JAHA
JACC
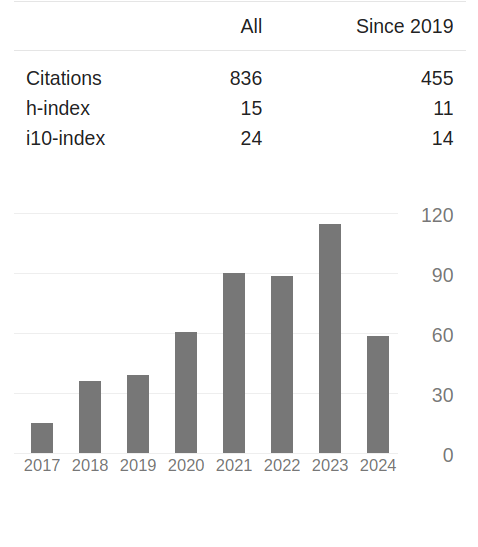
Publications
ALTMETRIC
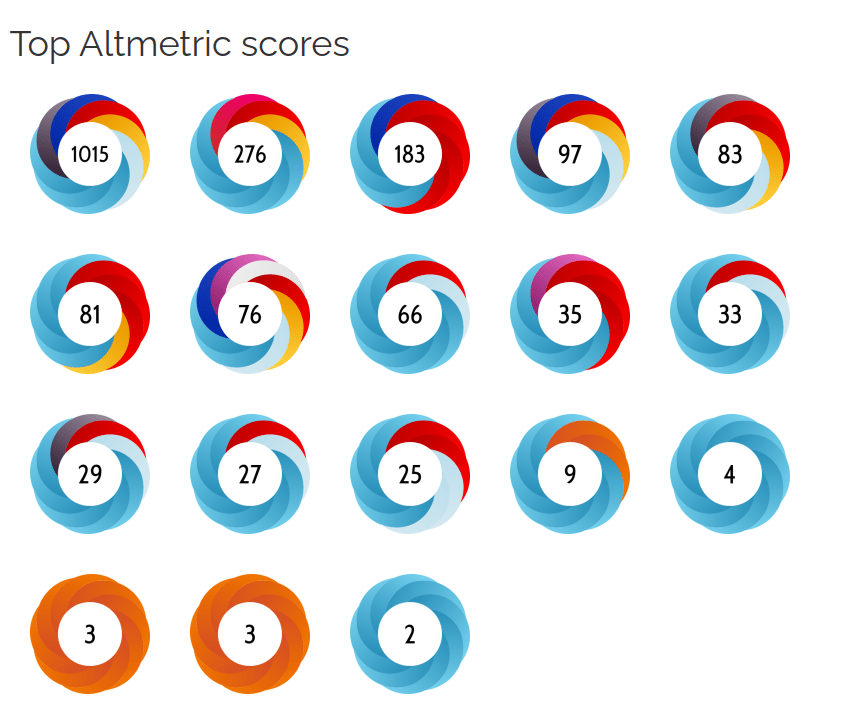

Scores
Impact on Popular Discourse on AI

In
National Pop-culture Discourse
Interviews, Op-eds, and Forum Appearences
- Joe Rogan Podcast
- Walter Isaacson Interview
- Speaker on Pritzker Forum on Global Cities
- >150 News articles written on published papers
Media Coverage
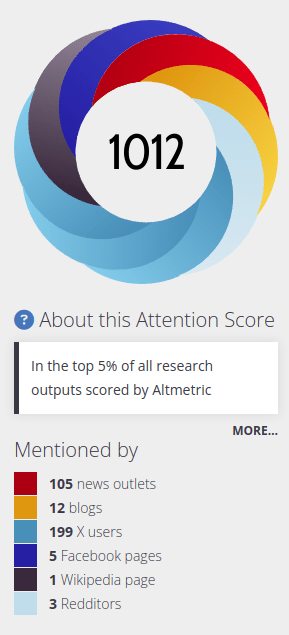
Rotaru, Victor, Yi Huang, Timmy Li, James Evans, and Ishanu Chattopadhyay. "Event-level prediction of urban crime reveals a signature of enforcement bias in US cities." Nature human behaviour 6, no. 8 (2022): 1056-1068.






MEDIA
Research Direction 1
point-of-care screening for complex diseases
Can we use existing EHR to reliably screen for complex diseases such as pulmonary fibrosis, dementia and rare cancers?
Ai
Electronic Healthcare Record
IPF
ASD
ADRD



Onishchenko, Dmytro, Robert J. Marlowe, Che G. Ngufor, Louis J. Faust, Andrew H. Limper, Gary M. Hunninghake, Fernando J. Martinez, and Ishanu Chattopadhyay. "Screening for idiopathic pulmonary fibrosis using comorbidity signatures in electronic health records." Nature Medicine 28, no. 10 (2022): 2107-2116.
Universal screening for complex diseases
Research Direction II
Can We Model Ecosystems As They Evolve ?
Can we predict future mutations?
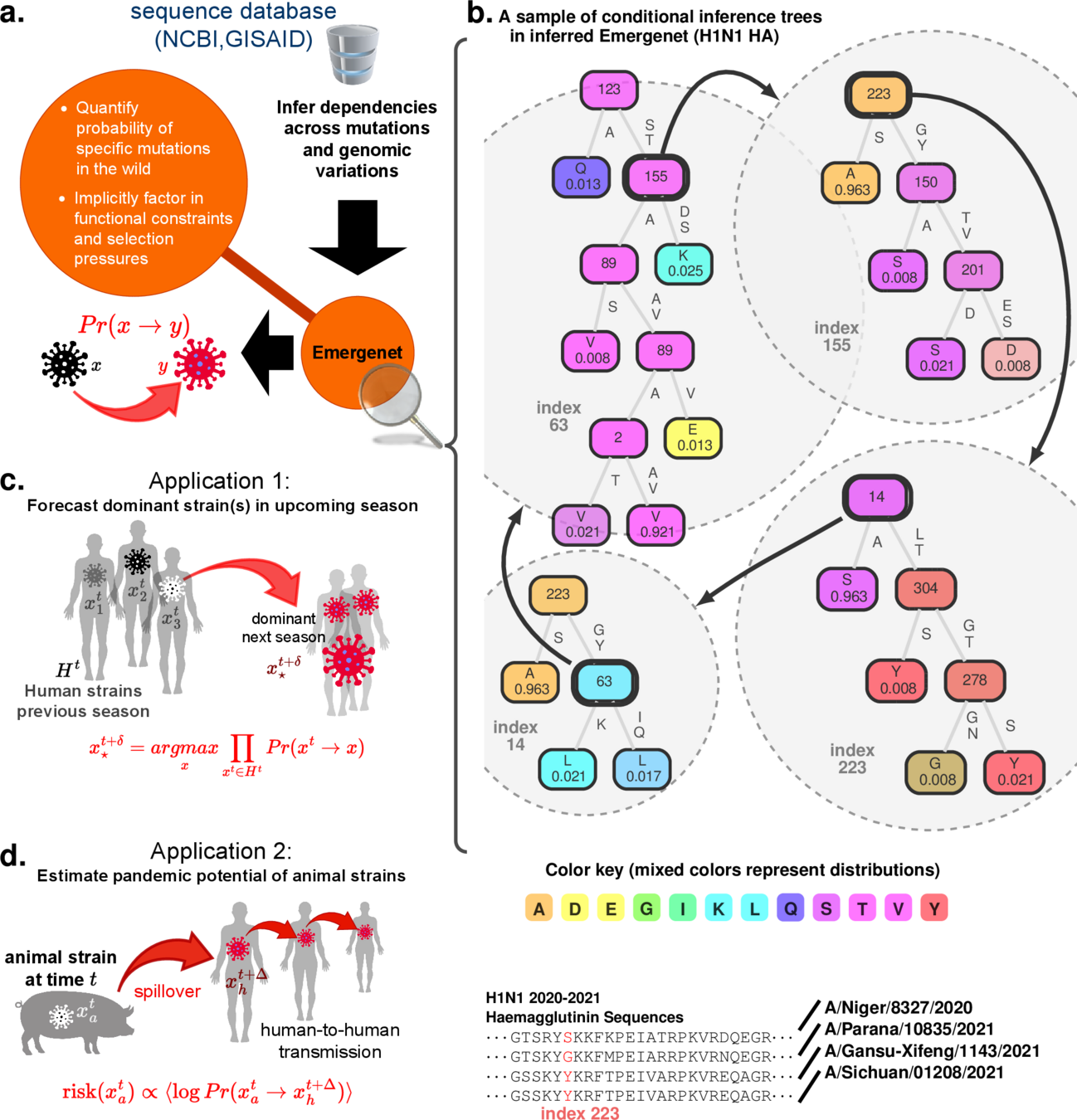
Digital Twins for complex systems
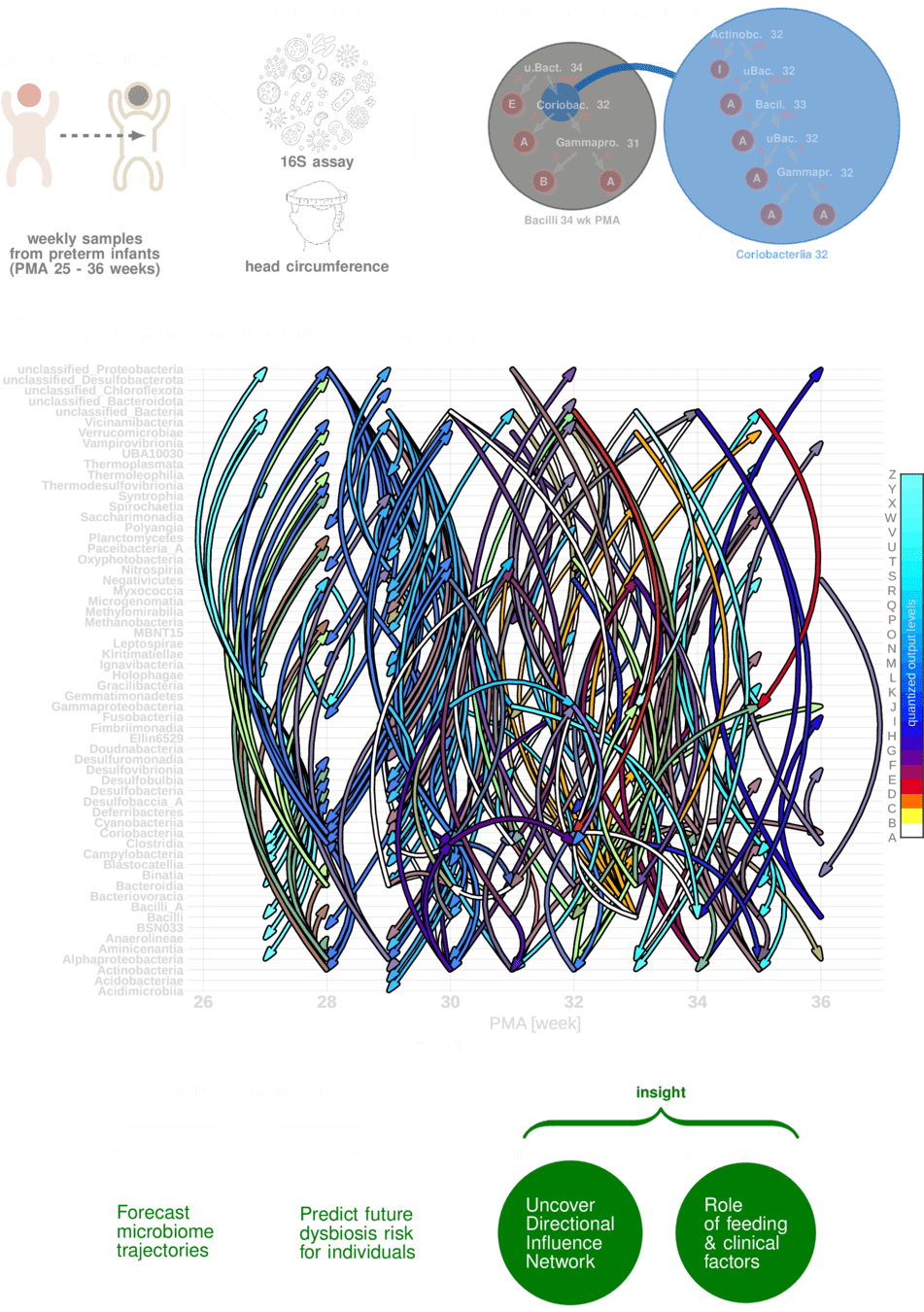
Can we find generative models for microbiome dynamics?
Teaching and Mentoring
- CCTS 40500/ CCTS 20500 / BIOS 29208 MACHINE LEARNING IN BIOMEDICINE (2019-, Winter Quarter, 30 hrs, 100 units)
- Mentored and guided research projects for undergraduate /graduate students/ Post-doctoral Associates (2016-)
- MEDC/ISTP 42000 Topics In Biomedical Data Analysis - Big Data 2017 (6 hrs)
- Mentoring undergraduate student (J. Lee) as a part of the Collegiate Translational Medicine Program (CTMP) (2018-2019)

-
Dr. Shahab Asoodeh
-
Dr. Yi Huang
-
Dmytro Onishenko
-
Victor Rotaru
-
Jin Li
-
Ruolin Zhang
-
David Yang






-
Dr. Nicholas Sizemore
-
Drew Vlasnik
-
Lucas Mantovani
-
Jaydeep Dhanoa
-
Jasmine Mithani
-
Angela Zhang
-
Warren Mo
-
Kevin Wu







Students, Postdocs and Mentees
Postdoc Placement:
Brookhaven National Laboratory, McGill University
Teaching AI
Math, Software, or Insight?
Developing the Human Capital To Lead The AI-Revolution

How do we teach AI?
Mathematics, Statistics, Data Science?
Start with the theorems?
medical education
Bio-AI
A Future AI-Expert Must Have A Lay of The Land
Ultimately the actual coding is increasingly simple: However Need To Know What To Use When and Why?
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.5)
clf = RandomForestClassifier(max_depth=10, class_weight='balanced',n_estimators=100).fit(X_train, y_train)
y_pred = clf.predict(X_test)
from sklearn.ensemble import RandomForestRegressor
from sklearn.ensemble import GradientBoostingRegressor
from sklearn.ensemble import ExtraTreesRegressor
from sklearn.ensemble import BaggingRegressor
from sklearn.svm import SVR
regr = BaggingRegressor(n_estimators=10,).fit(X_,y_)
regr = GradientBoostingRegressor(max_depth=None).fit(X_,y_)
regr = ExtraTreesRegressor(max_depth=None,n_estimators=100).fit(X_,y_)
regr = RandomForestRegressor(max_depth=None,n_estimators=100).fit(X_,y_)
Ignite Interest
Show how You can Change The World
Example: Make problem-solving fun and adventurous

Midterm

1854 London
Imagine you have been transported to 1854 London, during a cholera outbreak.
People are dying
The current scientific knowledge on germ theory is rudimentary, and mostly incorrect. There is no notion of epidemiology.
Your Goal:
Use data science
to correctly infer that:
1) cholera is most likely water-borne,
2) isolate source of the infection
Midterm

1854 London
The data is as follows:
- Confirmed cholera deaths
- Water pump location (black square)
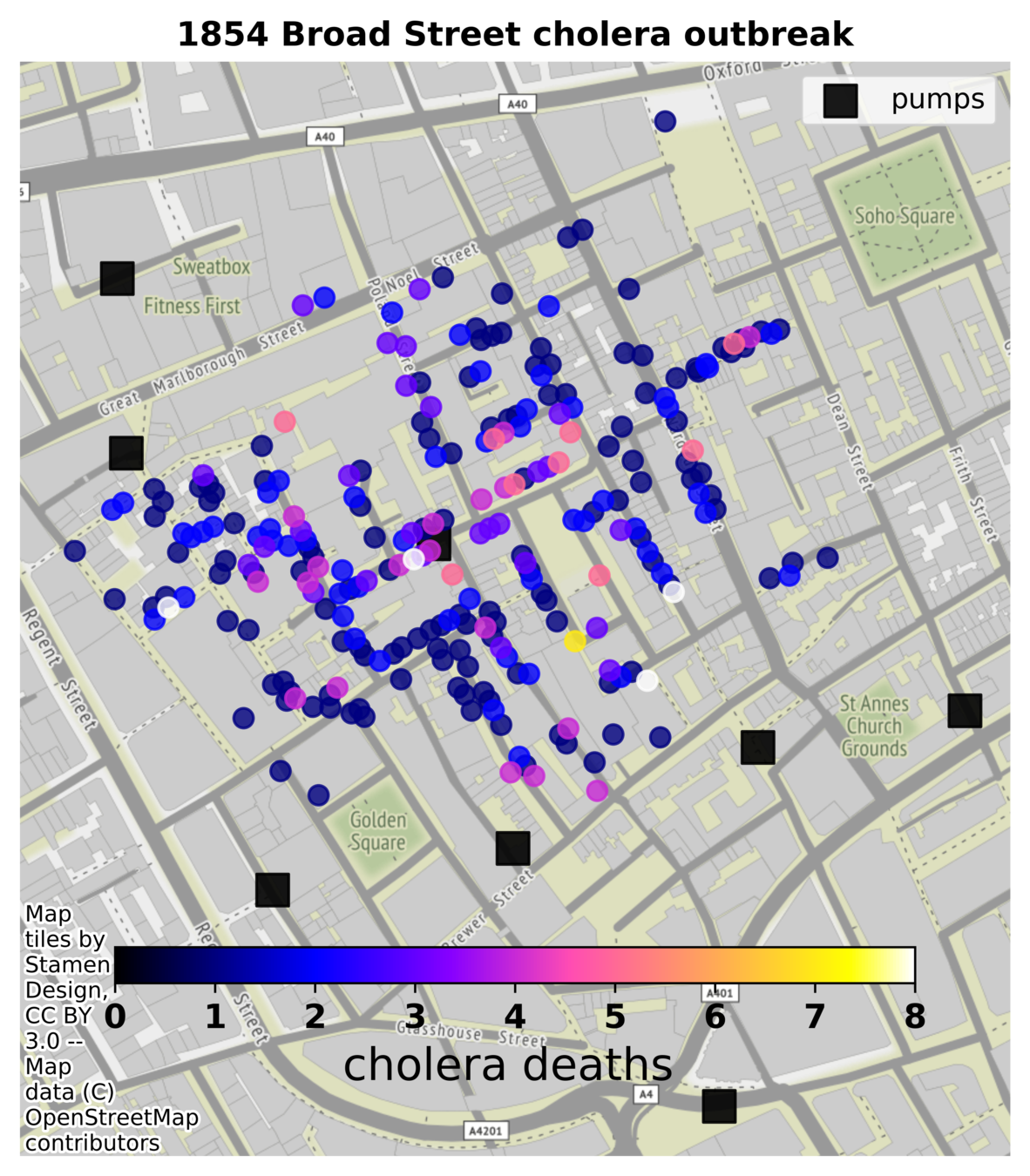
Midterm

1854 London
John Snow (15 March 1813 – 16 June 1858) was an English physician and a leader in the development of anaesthesia and medical hygiene. He is considered one of the founders of modern epidemiology, in part because of his work in tracing the source of a cholera outbreak in Soho, London, in 1854, which he curtailed by removing the handle of a water pump.
Snow used a dot map to illustrate the cluster of cholera cases around the pump. He also used statistics to illustrate the connection between the quality of the water source and cholera cases. Snow's study is regarded as the founding event of the science of epidemiology.


Q1. Is the disease waterborne?
- Show that the probability of cholera deaths are strongly spatially associated with location of water pumps
Q2. Locate the pump that is most likely the source of the disease, with estimated probability
- Show that an unique pump is central to the estimated distribution of deaths
Hint: Use different ensemble regressors to estimate the distrbution of deaths on a fine grid, trained with the data that is available
- How do you know which regressor is most "believable"?
Can you retrace J. Snow's argument?
Service
- AMGEN Scientific Advisory Board, Future trends in asthma, COPD, and IPF
- AlphaRecon Scientific Advisory Board, AI In Event prediction in Complex systems
- MacLean Center For Bioethics - Summer Lectures
- Harper Lecture, 2023
- Organizer: Workshops on Crimes of Prediction, Neubauer Collegium
- Organizer: Workshop on Data-driven Discovery of Models (D3M)
- Scientific Advisor, Adiona Health, 2022-current (Dementia management)
- Co-Mentoring MD Trainees at Brigham and Women’s Hospital, Harvard Medical School in critical pulmonary care (with Gary Hunninghake)
- Advisory Presentations, Cook County Commission on Social Innovation
Service
- Steering Committee Member proposed AI in Medicine Center IN Biological Sciences Division (2022)
- Faculty Search Committee for proposed AI in Medicine Center IN Biological Sciences Division (2022)
- Faculty Search Committee for Research-Focused Faculty in Medicine/BSD with Social Sciences emphasis (2021)
- Member of the Committee on Committee on Quantitative Methods in Social, Behavioral, and Health Sciences (2019-)
- Member of the Committee on Genetics, Genomics and Systems Biology (GGSB) (2018-)
- Interviewed potential graduate students for GGSB (2019,2020,2021,2022,2023)
- Member of Research Committee in the Section of Hospital Medicine, Department of Medicine, University of Chicago (2016-)
- Interviewed candidates for the Clinical Informatics Fellowship in the Department of Medicine (2018)
- Helped identify candidates for BSD Dean’s faculty search (Spearheaded by T. Conrad Gilliam) in the area of clinical/bio-medical data science (September 2017)
- REVIEWED AND SCREENED APPLICANTS FOR SUMMER QUANTITATIVE BIOLOGY FELLOWSHIP IN THE COLLEGE ALONG WITH Dmitry Kondrashov (2018)
Mission
Vision
Transform bio-surveillance
Transform modeling of complex systems
Transform early diagnosis
Democratize AI unleashing its power for social good
"Empowering Minds, Transforming Futures: AI Innovation with Purpose."
Enabling technology to reduce human suffering
Enabling the Vulnerable
To Have a Voice
Early Life Interventions to maximize human potential
Health Equity
Values
"Fusing Innovation with Compassion:
Advancing AI Research for the Greater Good."
Classroom
Social Justice and Diversity, Equity and Inclusion
Research Group Composition
Highly diverse, more than 40% women, and representative of UoC student population
40% Women in Undergraduate and graduate students
50% women among past postdoctoral associates
Research Direction 1
Universal Screening?
- Autism
- Idiopathic Pulmonary Fibrosis
- Alzheimer's Disease and related dementia
- Suicidality, PTSD
- Perioperative Cardiac Event
- Aggressive Melanoma
- Uterine Cancer
- Pancreatic Cancer
- ...


- non-existent biomarkers
- expensive, time-consuming diagnostic tests
- Lack of Universal Screening at the point of care
- Early diagnosis is difficult, late or missed diagnosis costs lives
Prognosis at Point-of-Diagnosis
- Optimizing Management
Patient Journey
- Continuous Risk Monitoring
Early Diagnosis
- Universal Screening
- Cohort Selection
Reduce screen failure rates
Holistic health surveillance
Predict antifibrotics continuation
improve outcomes
1
2
3
Interstitial Lung Disease / Pulmonary Fibrosis
Rapid Universal Point-of-care Screening for ILD/IPF Using Comorbidity Signatures in Electronic Health Records
Flag patients before they (or doctors) suspect
Primary Care
Pulmonologist
Zero-burden Co-morbid Risk Score (ZCoR)
Referral



shortness of breath
dry cough
doctor can hear velcro crackles
Non-specific Symptoms
>50 years old
more men than women
IPF
Rare disease
~5 in 10,000
Post-Dx
Survival
~4 years
Cannot always be seen on CXR
At least one misdiagnosis
~55%
Two or more misdiagnosis
38%
Initially attributed to age related symptoms:
72%

PCP workflow demands
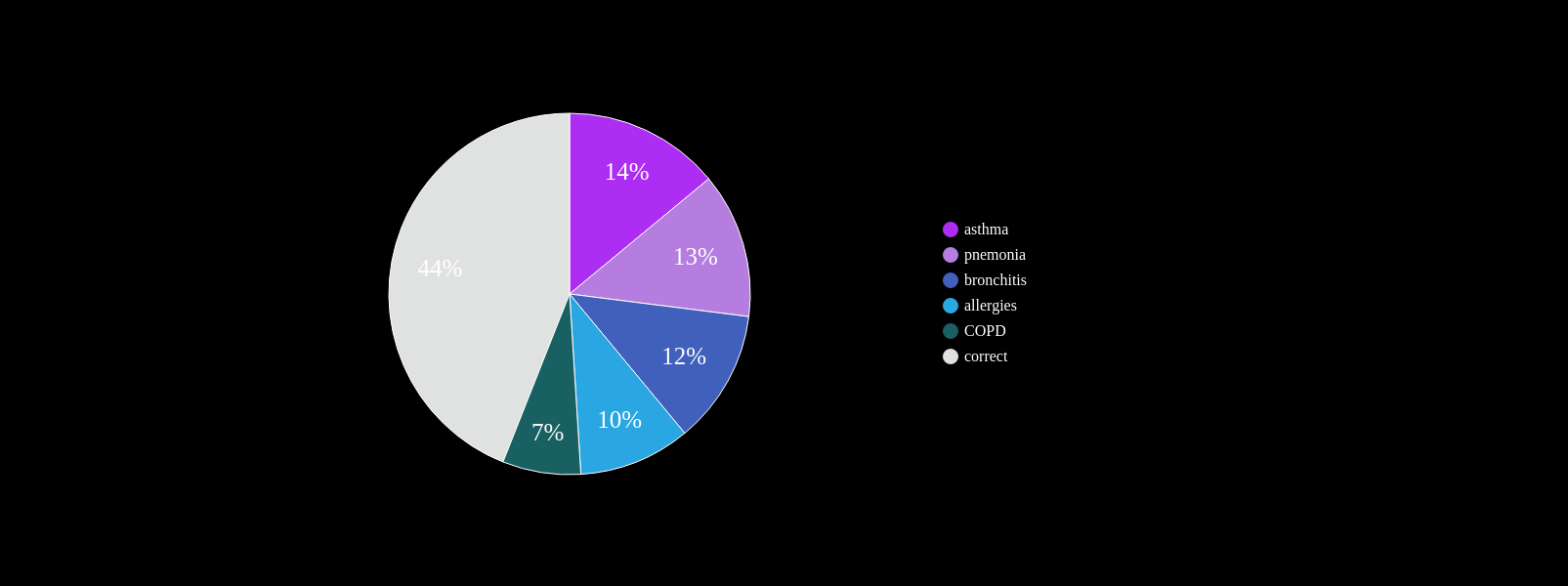
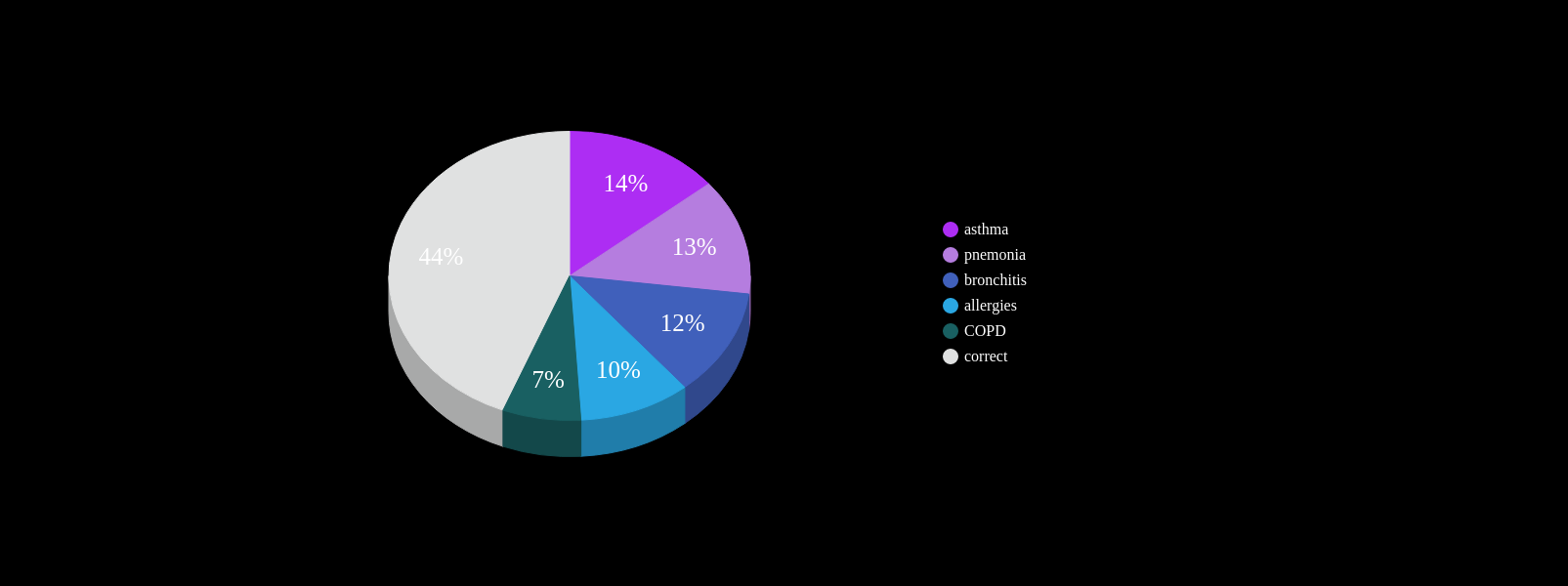
Known Co-morbidities of PF
Are there more? Subtle footprints in the medical history that are more heterogeneous?

~ 4yrs
current survival ~4yrs
~ 4yrs
current clinical DX
ZCoR screening



Onishchenko, D., Marlowe, R.J., Ngufor, C.G. et al. Screening for idiopathic pulmonary fibrosis using comorbidity signatures in electronic health records. Nat Med 28, 2107–2116 (2022). https://doi.org/10.1038/s41591-022-02010-y
n=~3M
AUC~90%
Likelihood ratio ~30

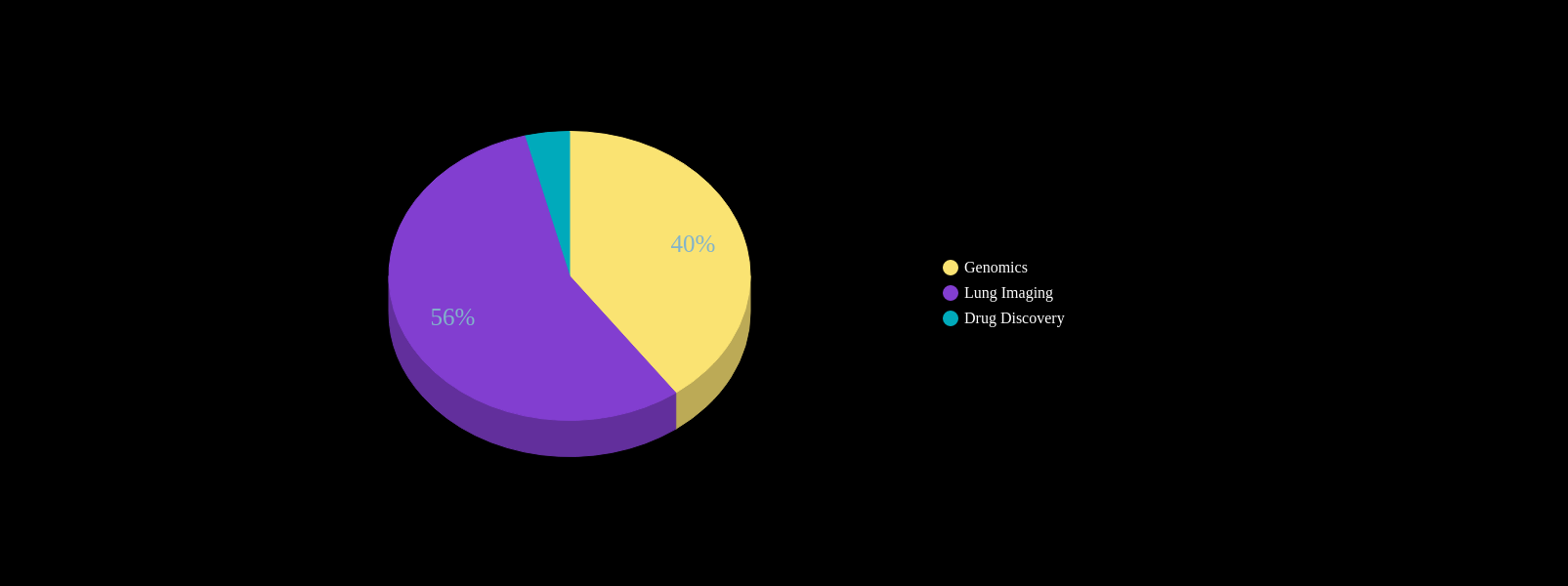


Conventional AI/ML attempts to model the physician
AI in IPF Research
- Co-morbidity patterns
- No data demands
- Use whatever data is already on patient file
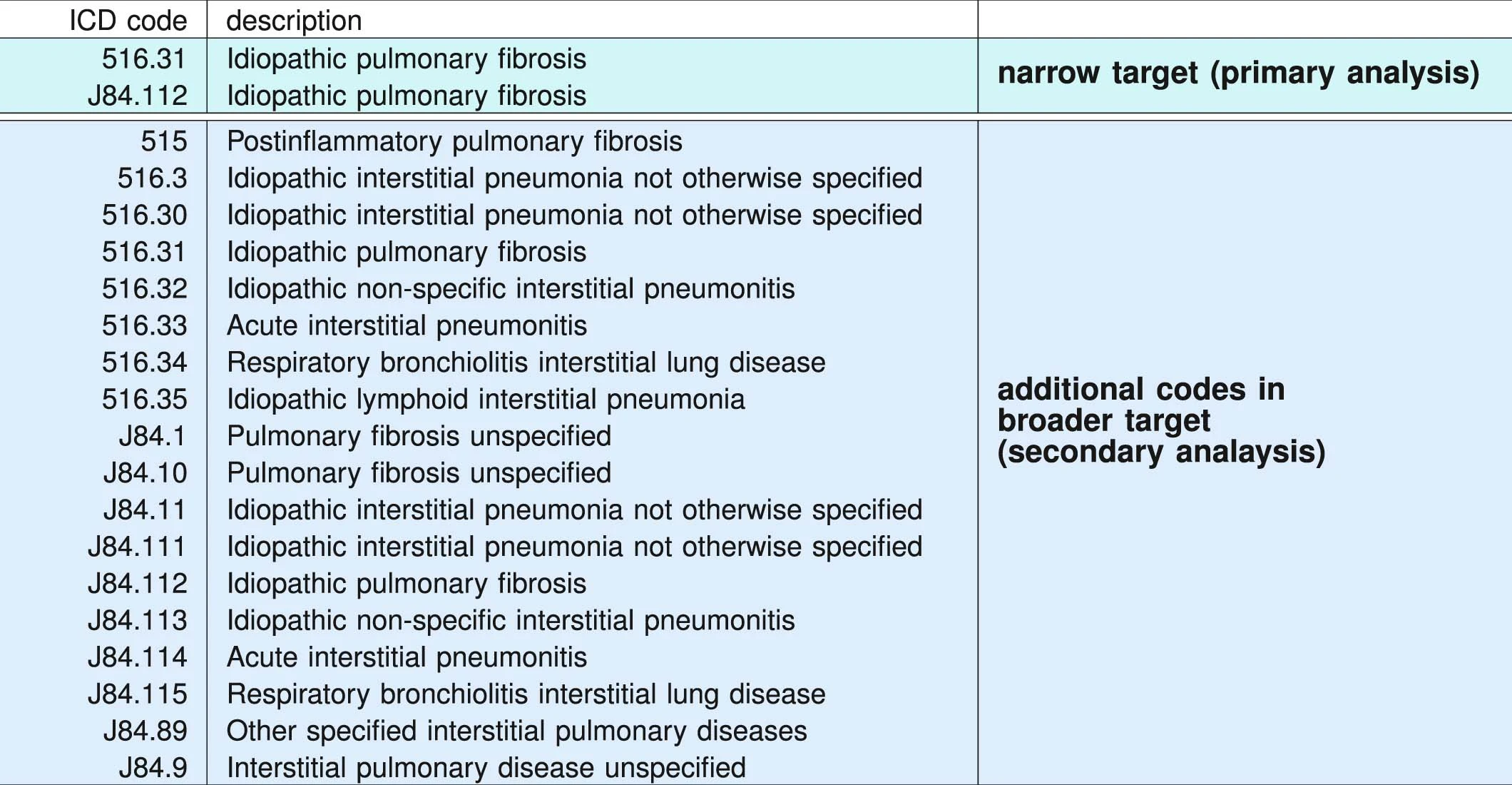
ICD administrative codes

IPF
ILD
target codes appear
Past medical history
No target codes appear
case
control
2yrs
2yrs
prediction
target codes appear
Past medical history
No target codes appear
case
control
2yrs
2yrs
IPF drugs prescribed
Signature of IPF diagnostic sequence
pirfenidone or nintedanib
- age > 50 years
- at least two IPF target codes identified at least 1 month apart
- chest CT procedure (ICD-9-CM 87.41 and Current Procedural Terminology, 4th Edition, codes 71250, 71260 and 71270) before the first diagnostic claim for IPF
- no claims for alternative ILD codes occurring on or after the first IPF claim
ICD Codes can be noisy
"cases" are not always true IPF
Truven MarketScan (IBM) Commerical Claims & Encounters Database 2003-2018
>100M patients visible
>7B individual claims
>87K unique diagnostic codes
>7% Medicare data present
2,053,277 patients included in study

University of Chicago Medical Center 2012-2021
68,658 patients
Random sample from Optumlabs Data Warehouse courtsey Mayo Clinic
861,280 patients
2,983,215 patients
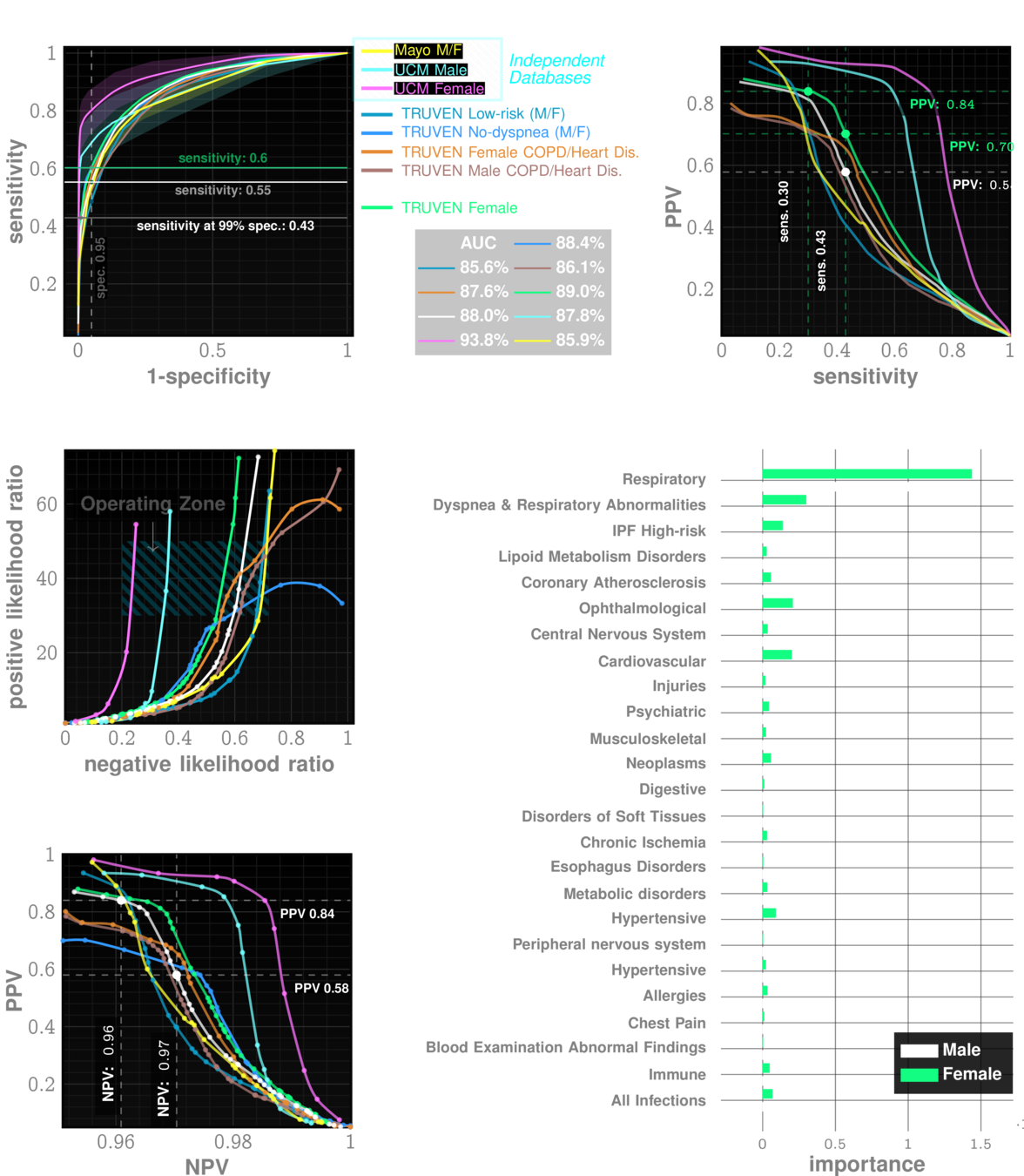

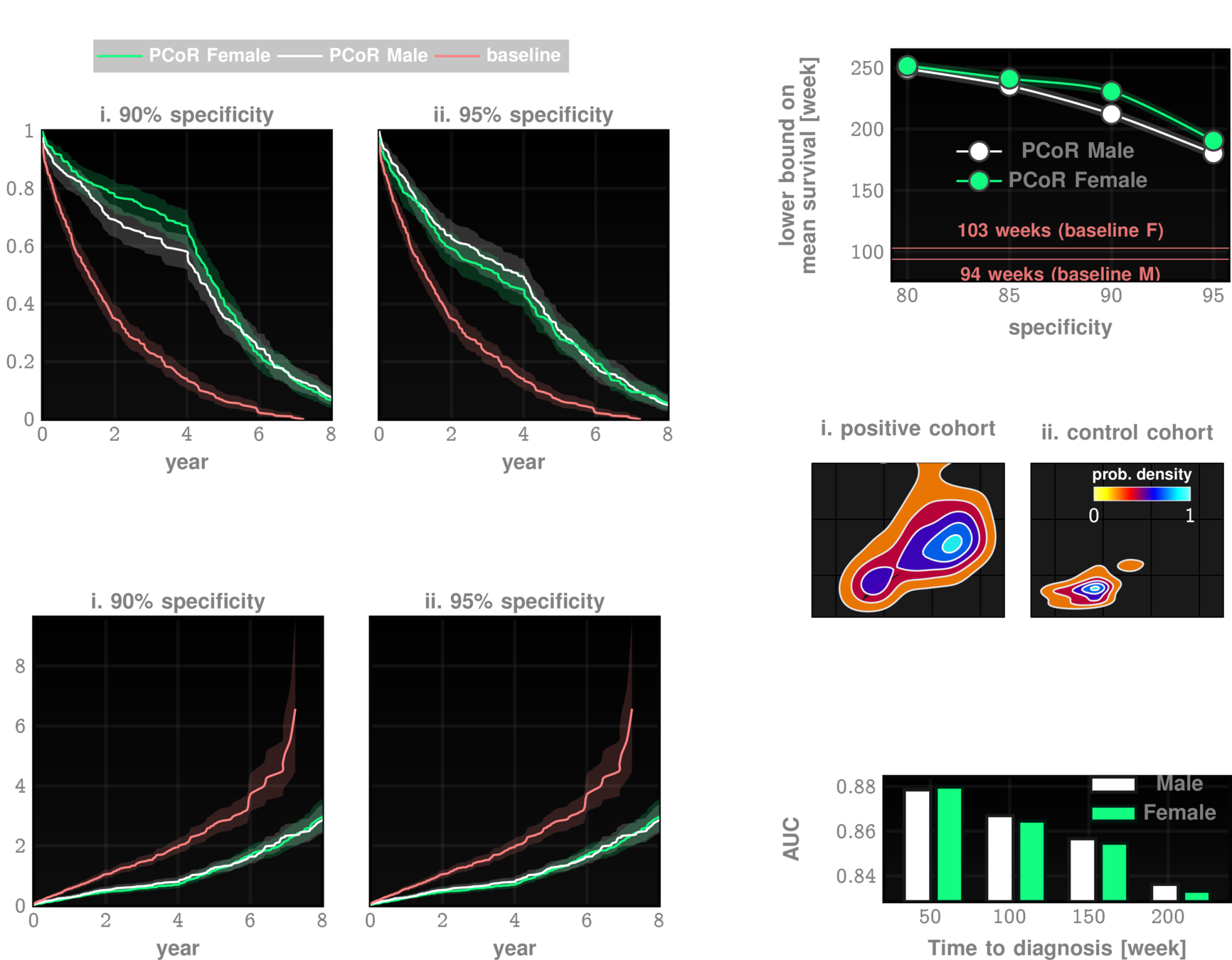
Data: Onishchenko etal. Nat. Medicine 2022
performance tables
Out-of-sample Results
specificity ~99%
NPV >99.9%
IPF
ILD




Comorbidity Spectra



patient A
patient B
patient C
Beyond "risk factors" to personalized risk patterns
False Positives:
- Heathcare Capacity
Ethics:
- Risk from Imaging Tests
For every 20-30 flags,
1 is positive
- General likelihood ratio 60-80
- PPV 3.5-5%
- Notifying patients 4 years early?
- No cure, why screen
minimal
acceptable?
Better outcomes
- early anti-fibrotic therapy seems increasingly promising
- better shot at lung transplant
- early dx reduces hospital-izations by a factor of 1-3
Collard, Harold R., Alex J. Ward, Stephan Lanes, D. Cortney Hayflinger, Daniel M. Rosenberg, and Elke Hunsche. "Burden of illness in idiopathic pulmonary fibrosis." Journal of medical economics 15, no. 5 (2012): 829-835.
Clinical Trial Cohort Selection
Current screen failure rate ~50-60%
ZCoR boosted screen failure rate ~20%
cohort size: 2000
initial cohort size: 5000
initial cohort size with ZCoR: 2500
Cost per patient for confirmatory tests: ~7k USD
Savings: ~20M USD
Cloud Deployment
Theoretical formulation
Multi-cohort validation
Launch User-Accessible Platform
3 years
2 years
[
{
"patient_id": "P000038",
"sex": "F",
"birth_date": "01-01-2006",
"DX_record": [
{"date": "07-31-2006", "code": "Z38.00"},
{"date": "08-07-2006", "code": "P59.9"},
{"date": "08-29-2016", "code": "J01.90"},
{"date": "09-10-2016", "code": "J01.90"},
{"date": "11-14-2016", "code": "J01.91"}
],
"RX_record": [
{"date": "10-29-2011", "code": "rxLDA017"},
{"date": "05-16-2015", "code": "rxIDG004"},
{"date": "08-08-2015", "code": "rxIDG004"},
{"date": "06-04-2016", "code": "rxIDD013"}
],
"PROC_record": [
{"date": "02-05-2007", "code": "90723"},
{"date": "11-05-2007", "code": "J1100"}
]
}
]{
"predictions": [
{
"error_code": "",
"patient_id": "P000012",
"predicted_risk": 0.005794344620009157,
"probability": 0.8253881317184486
}
],
"target": "TARGET"
}Data In
Data Out
The Paraknowledge API
curl -X POST -H "Content-Type: application/json" -d '[{"patient_id": "P28109965201", "sex": "M", "age": 89, "fips": "35644", "DX_record": [{"date": "12-16-2011", "code": "R09.02"}, {"date": "12-30-2011", "code": "H04.129"}, {"date": "12-30-2011", "code": "H02.109"}], "RX_record": [], "PROC_record": [{"date": "09-28-2012", "code": "71100"}]}]' "https://us-central1-pkcsaas-01.cloudfunctions.net/zcor_predict?target=IPF&api_key=7eea9f70d79c408f2b69847d911303c"Current Targets
IPF
ILD
ADRD
CKD
CKD_SEVERE
MELANOMA
CANCER_PANCREAS
CANCER_UTERUS
SISACohort Selection and Risk Analysis Testbed
Misleading Diagnosis of Idiopathic Pulmonary Fibrosis: A Clinical Concern
Javier Ramos-Rossy, MD, Onix Cantres-Fonseca, MD, Ginger Arzon-Nieves, Yomayra Otero-Dominguez, MD, Stella Baez-Corujo, MD, and William Rodríguez-Cintrón, MD

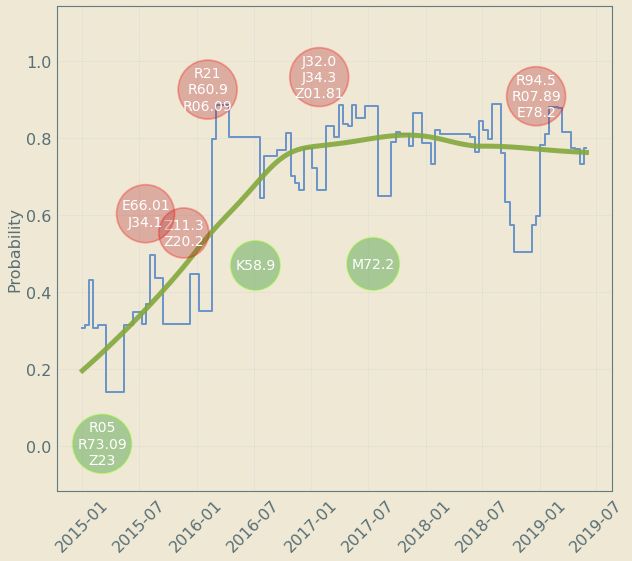
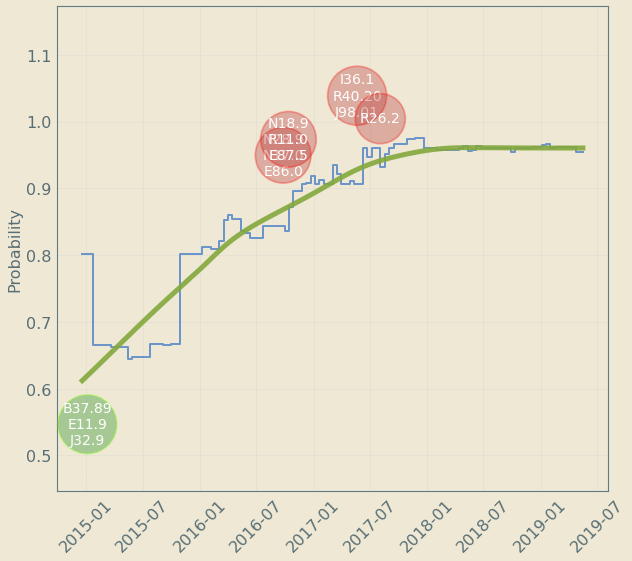
Upto 4 year "signal" resolution
decreases risk
increases risk
Patient Journey: Tracking Risk over time
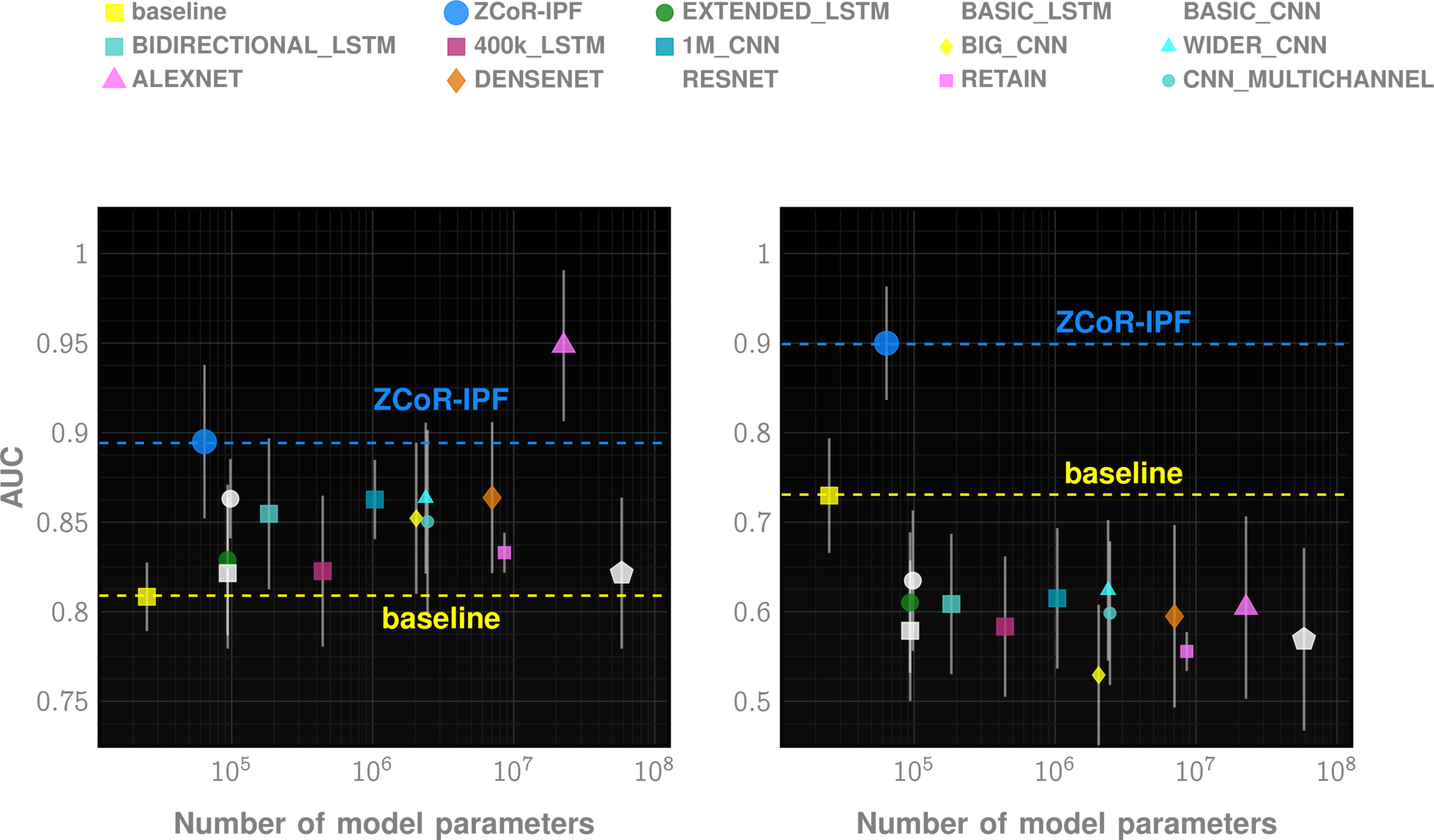
Off-the-shelf AI does not suffice
Modeling Longitudinal Patterns
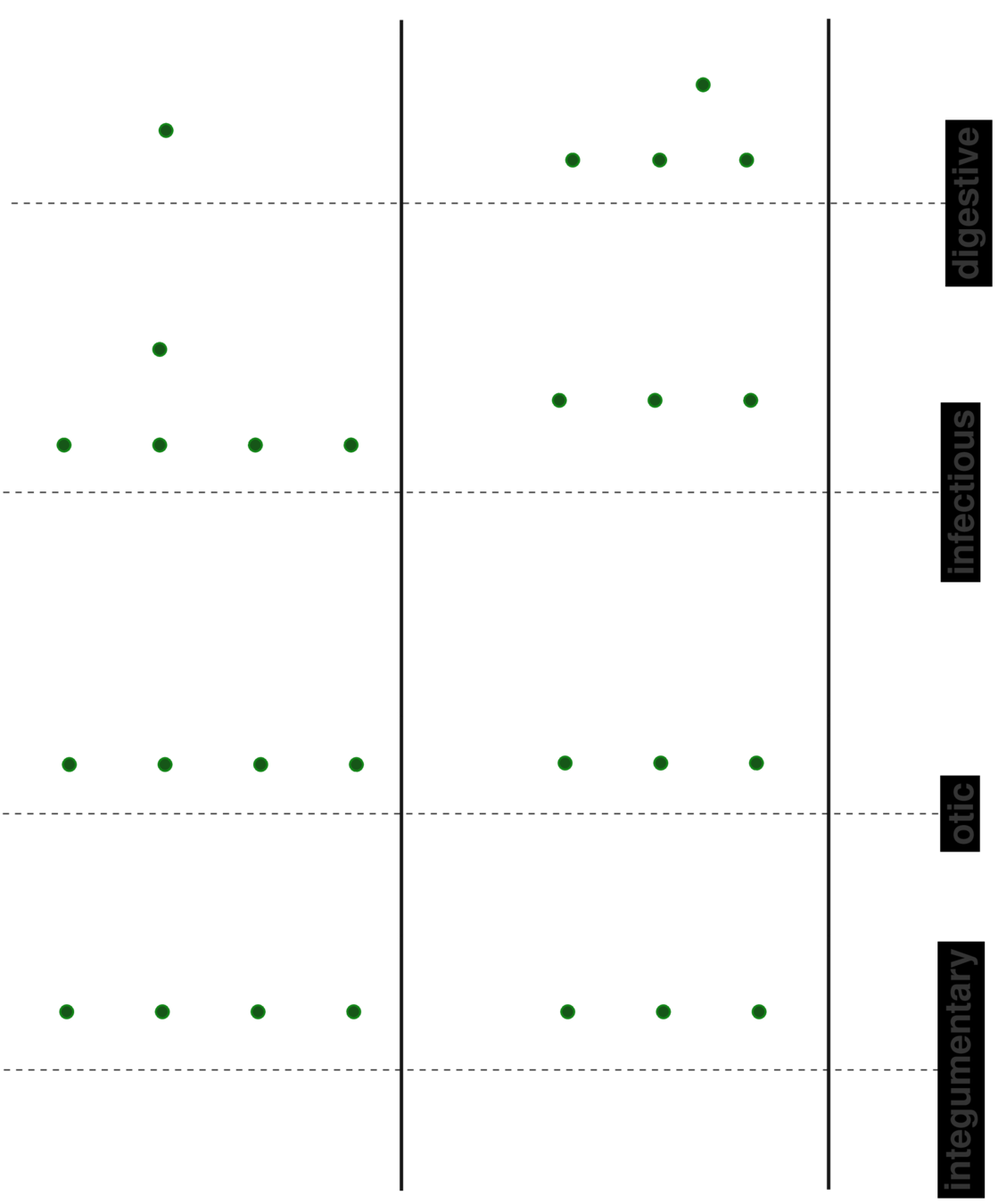
Specialized HMM models from code sequences
Model control and case cohorts seprately
given a new test case, compute likelihood of sample arising from case models vs control models
sequence likelihood defect
Huang, Yi, Victor Rotaru, and Ishanu Chattopadhyay. "Sequence likelihood divergence for fast time series comparison." Knowledge and Information Systems 65, no. 7 (2023): 3079-3098.
ZeD Lab: Predictive Screening from Comorbidity Footprints



Nature Medicine
JAHA
CELL Reports
Science Adv.
The ZCoR Approch: Rapidly Re-targettable
| ZED performance | Competition | |
|---|---|---|
| Autism | >80% AUC at 2 yrs | "obvious" |
| Alzheimer's Disease | ~90% AUC | 60-70% AUC |
| Idiopathic Pulmonary Fibrosis | ~90% AUC | NA |
| MACE | ~80% AUC | ~70% AUC |
| Bipolar Disorder | ~85% AUC | NA |
| CKD | ~85% AUC | NA |
| Cancers (Prostate, Bladder, Uterus, Skin) | ~75-80% AUC | Low |
Predictions at the Point-of-Diagnosis
Can my patient continue taking anti-fibrotics over long term?
Digital Twins for Health trajectories
}
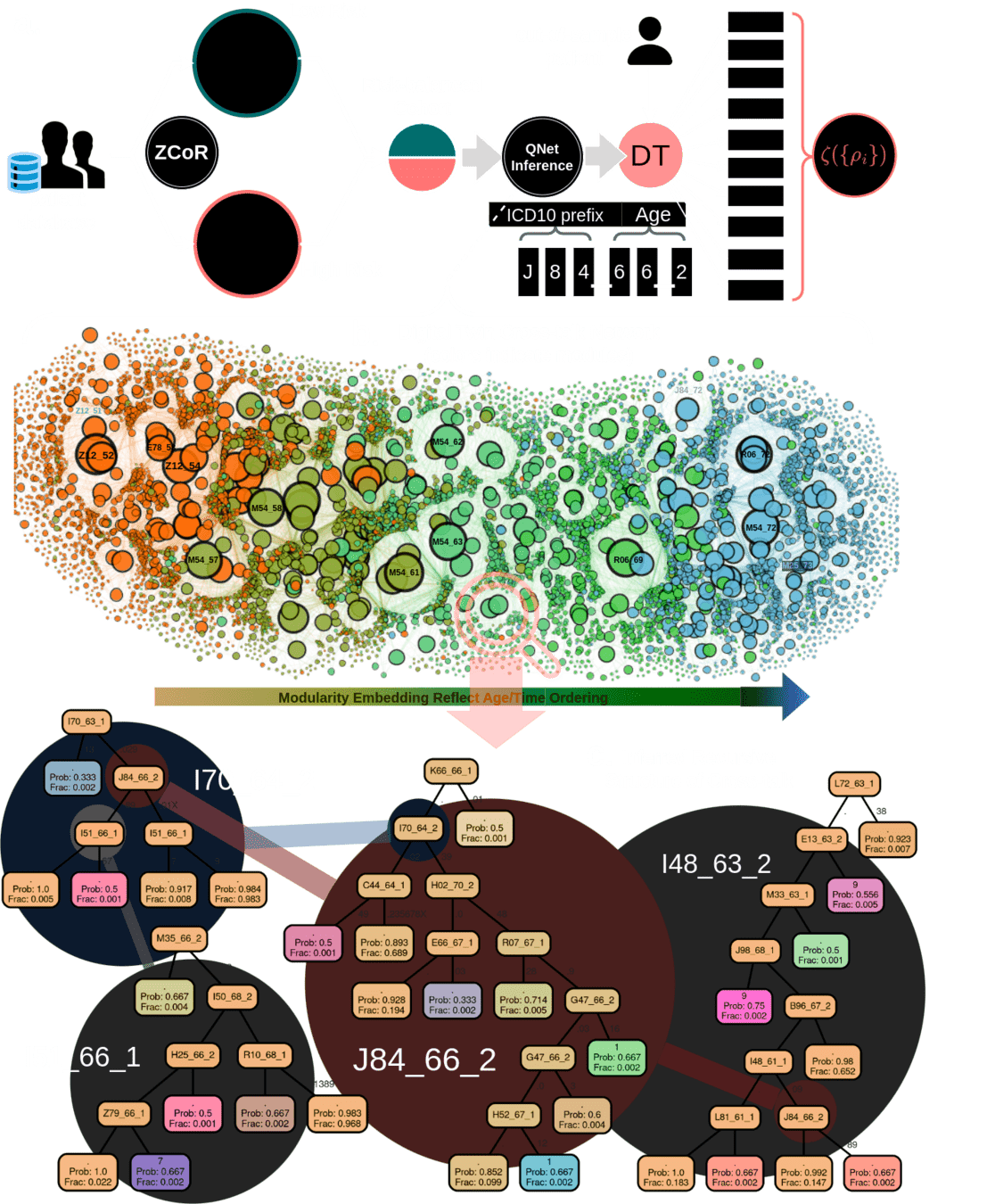
1M parameters
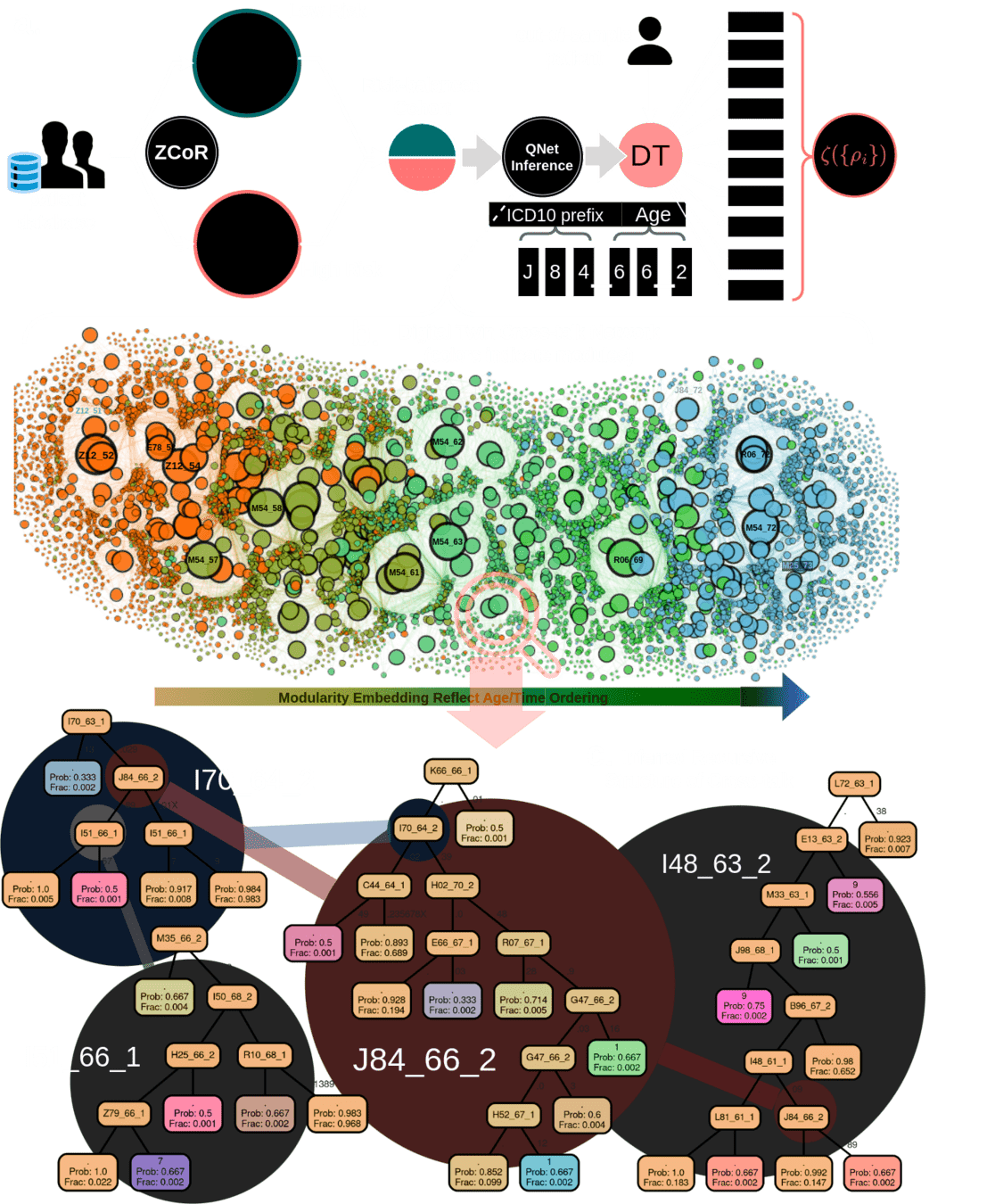
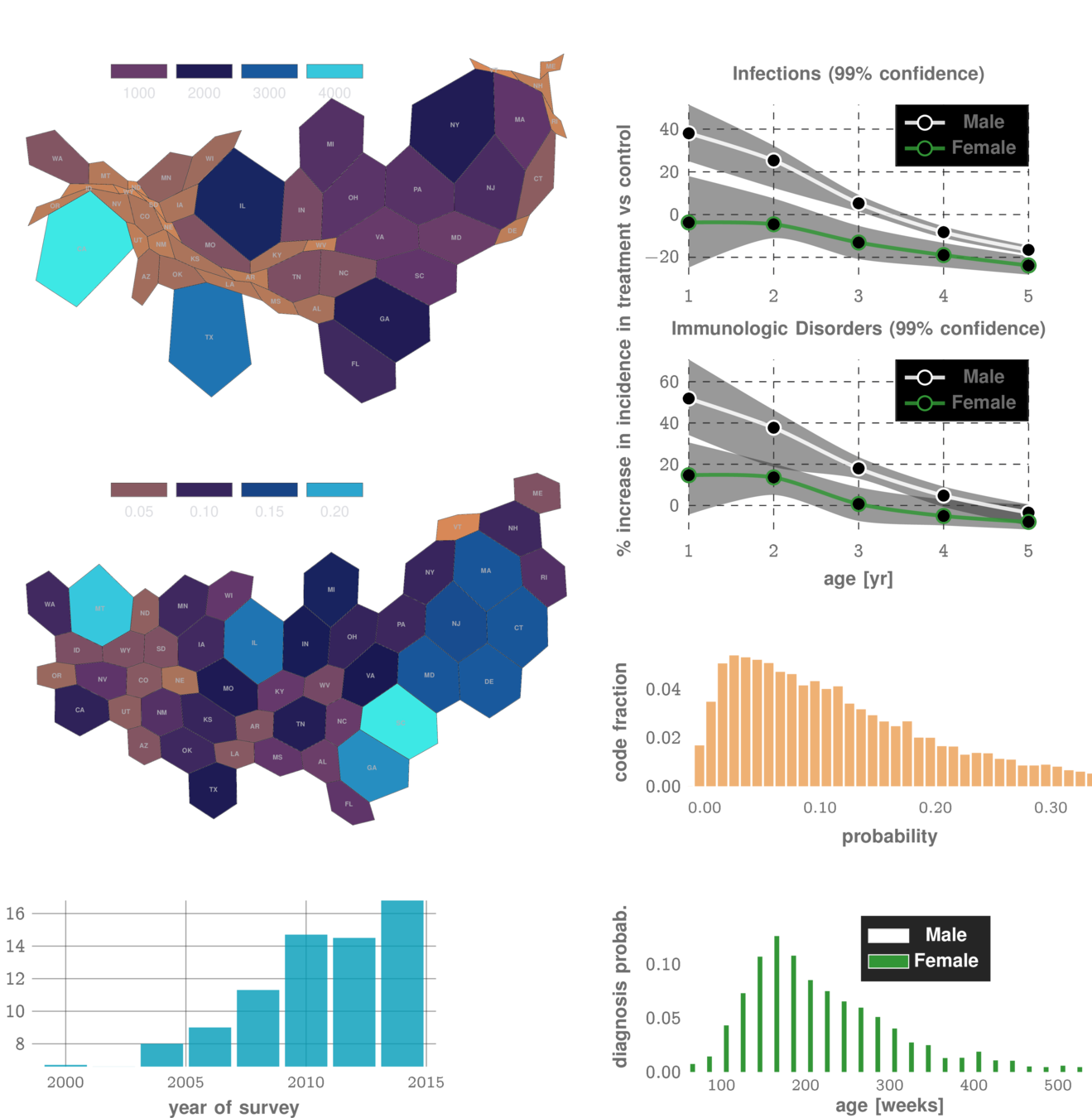
1M parameters
Predicts disorders across the disease specturm
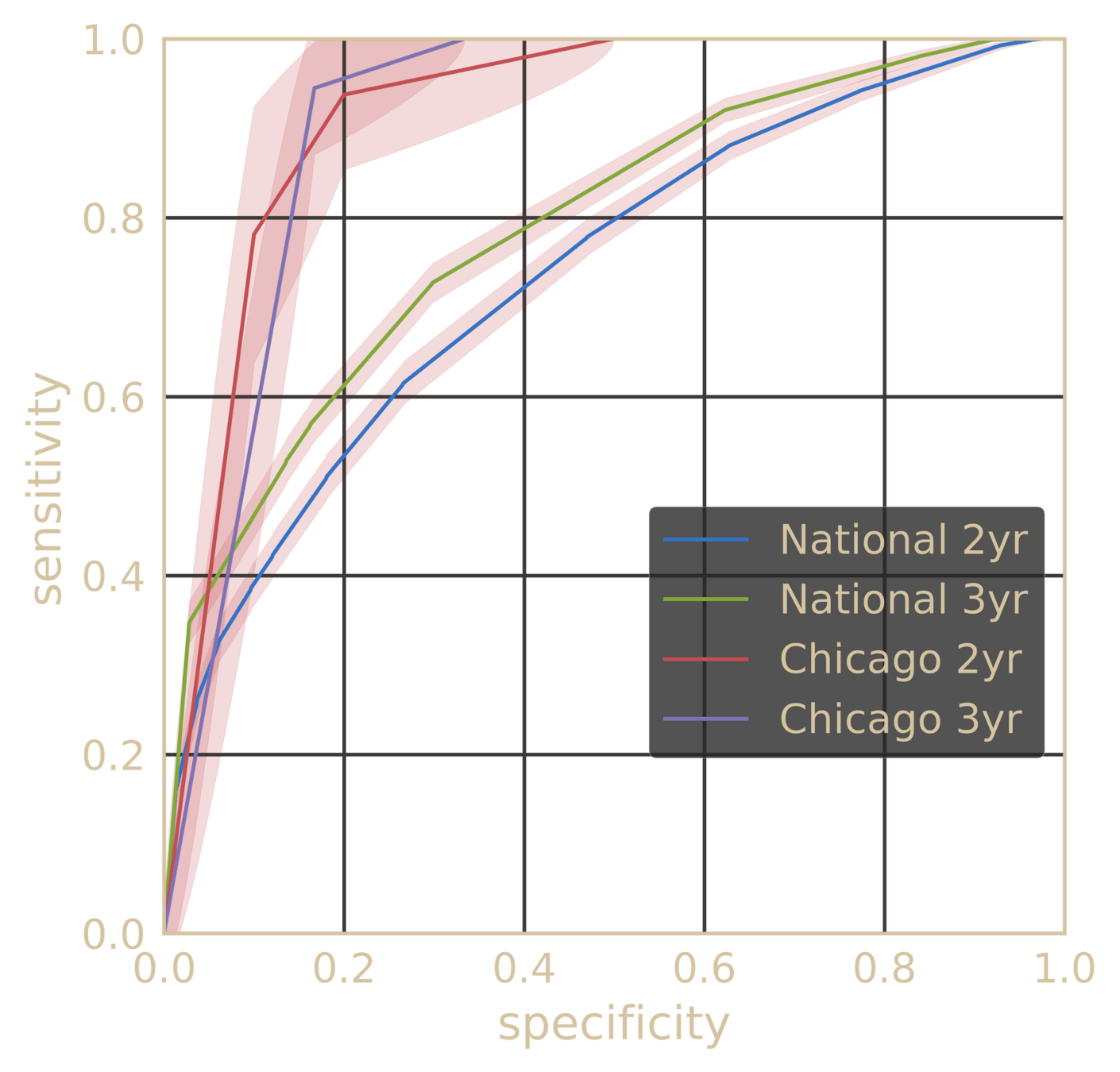
Pre-empting Effectiveness of Antifibrotics at the point of diagnosis
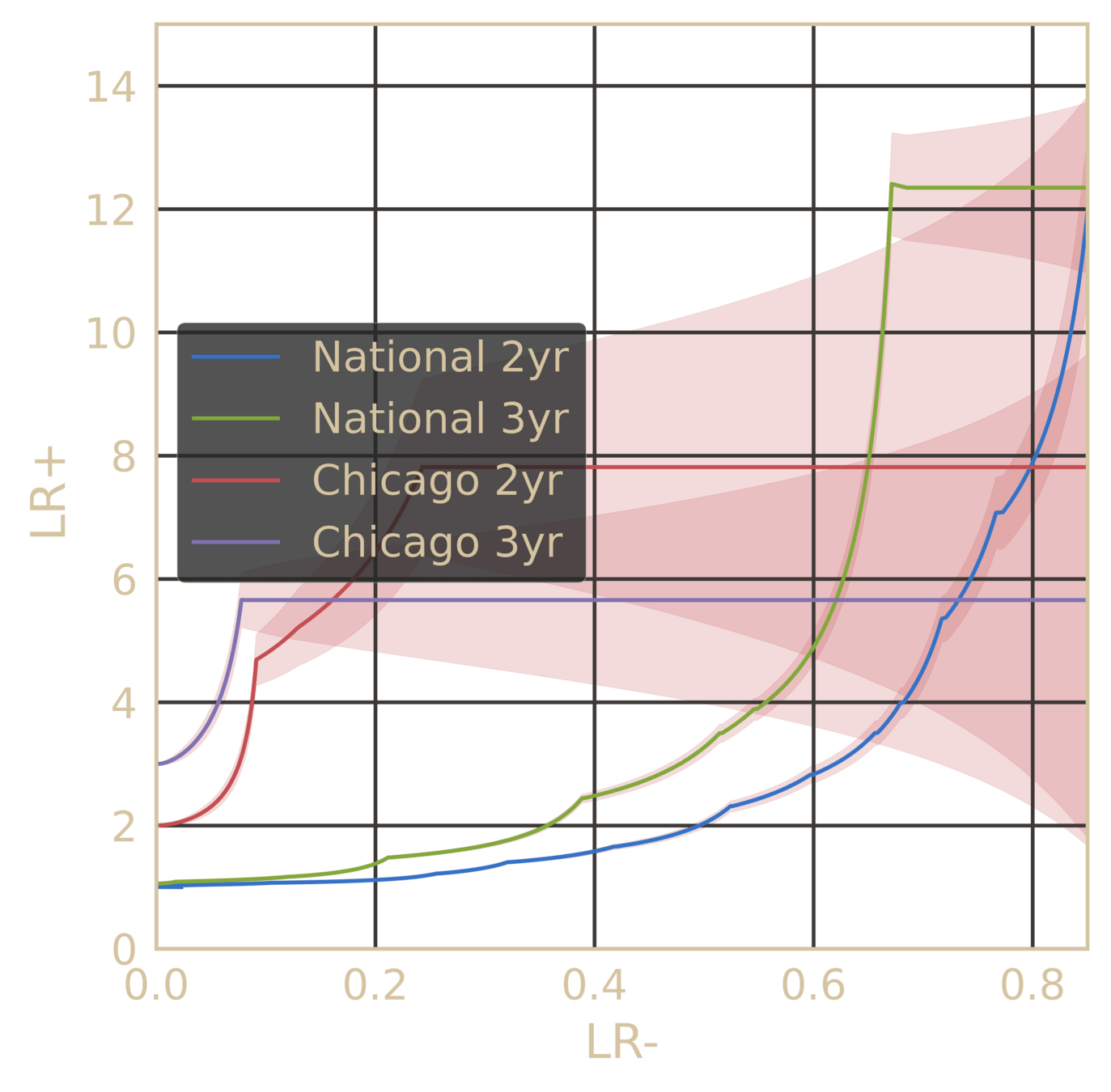
~78% AUC
26-32 out of 100 discontinued
4-5 out of 100 discontinued
Prognosis at Point-of-Diagnosis
- Optimizing Management
Patient Journey
- Continuous Risk Monitoring
Early Diagnosis
- Universal Screening
- Cohort Selection
Reduce screen failure rates
Holistic health surveillance
Predict antifibrotics continuation
improve outcomes
Summary
3
2
1
ishanu@uchicago.edu
@ishanu_ch
1 in 59
Autism Spectrum Disorder
36
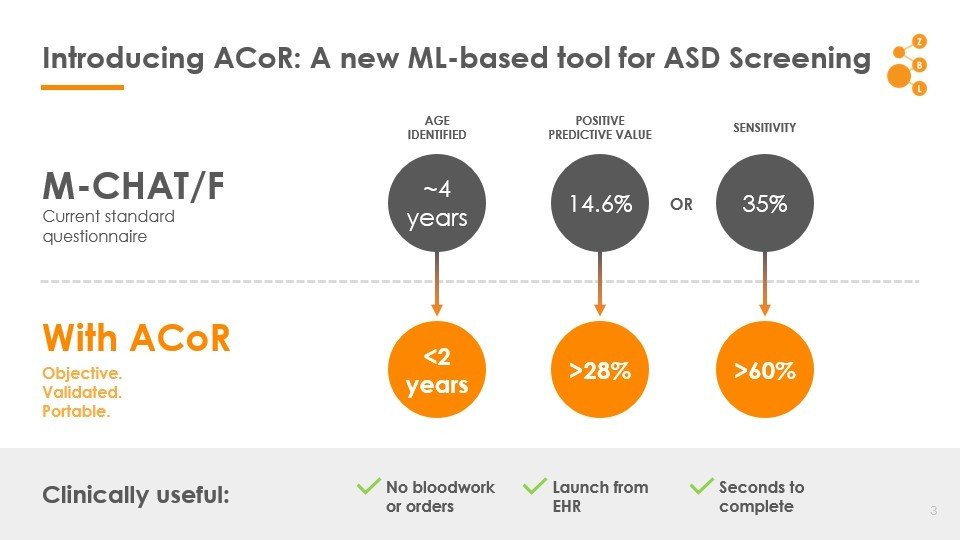
Autism Co-morbid Risk (ACoR) Score
Data: Onishchenko etal. Science Advances 2021
>5 Million in US. >13 Million in next 10 years
Alzheimer's Disease and Related Dimentia
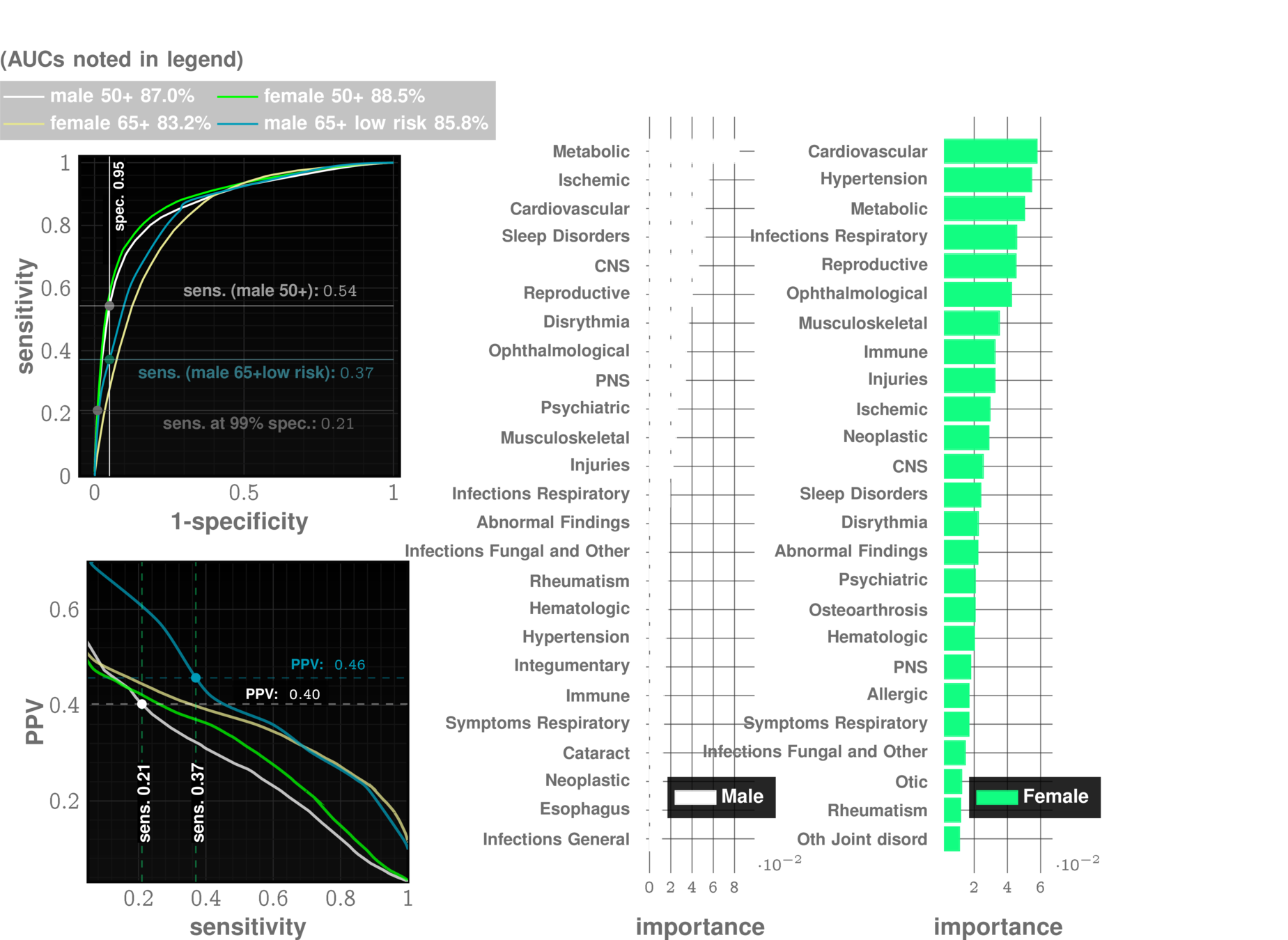
MOCA, Blood Tests
Current Practice:
state of art with EHR:
~67% AUC*
ZCoR: ~87%
Application to Suicide Attempts and Ideation (SISA) , PTSD*:
perhaps surprising connection between mood disorders and physiological comorbidities
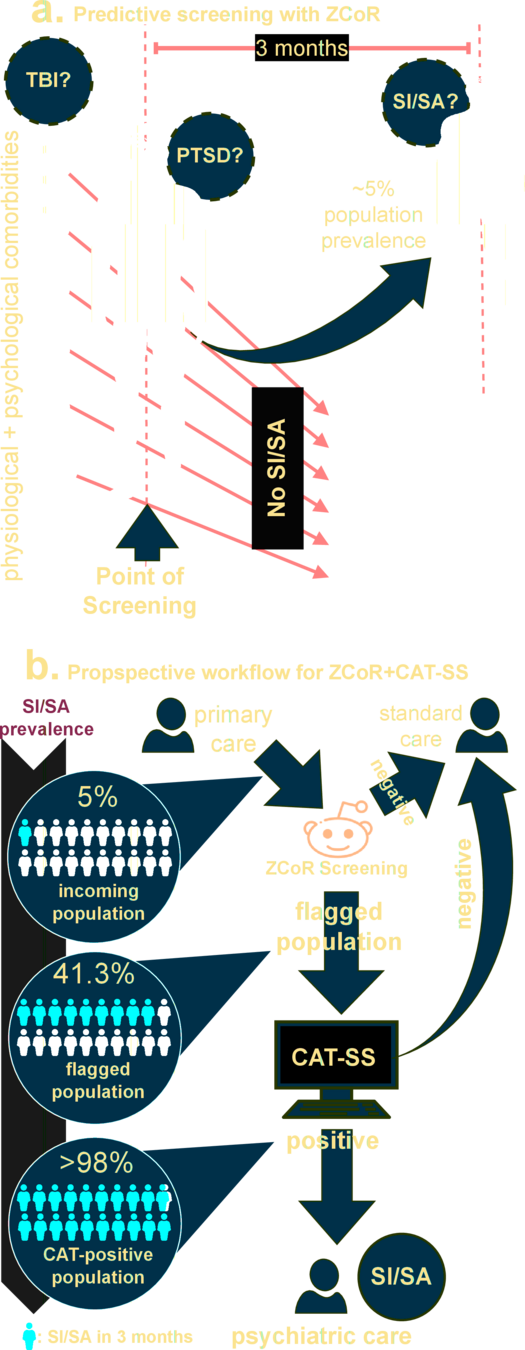

Gibbons RD, Kupfer D, Frank E, Moore T, Beiser DG, Boudreaux ED. Development of a Computerized Adaptive Test Suicide Scale-The CAT-SS. J Clin Psychiatry. 2017 Nov/Dec;78(9):1376-1382. doi: 10.4088/JCP.16m10922. PMID: 28493655.
* in press
Medicine is poised to enter a transformative era, ushered by the emergence of sophisticated Artificial Intelligence (AI) models.
Enable more holistic approaches to medicine, where predictive patterns can be rapidly recognized and exploited

Research Direction 2
Uncovering A Digital Twin of the Maturing Human Microbiome

Sizemore, Nicholas, Kaitlyn Oliphant, Ruolin Zheng, Camilia R. Martin, Erika C. Claud, and Ishanu Chattopadhyay. "A digital twin of the infant microbiome to predict neurodevelopmental deficits." Science Advances 10, no. 15 (2024): eadj0400.

ishanu chattopadhyay
Nicholas Sizemore
Kaitlyn Oliphant
Erika Claud

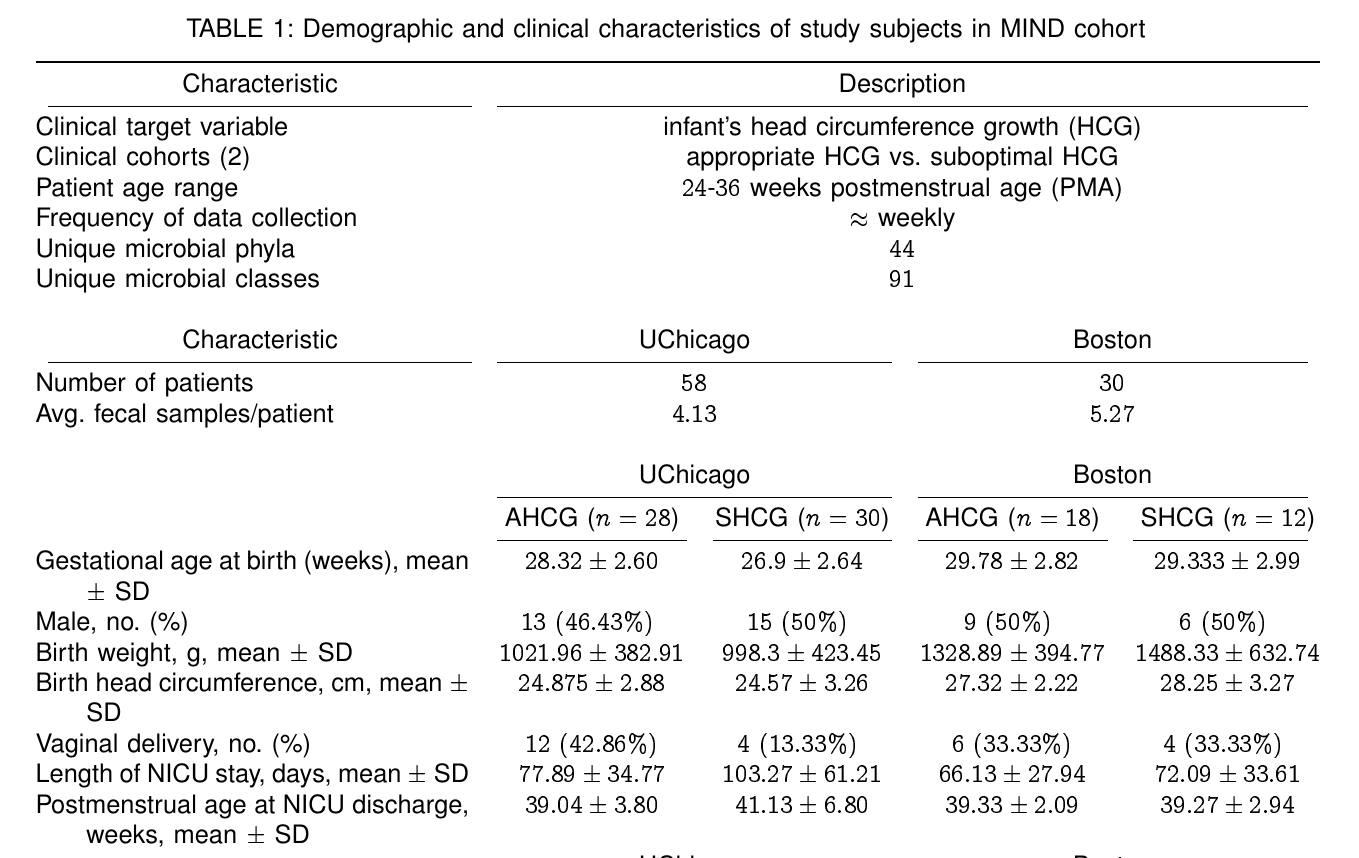
THE PROBLEM
Can microbial assay from gut actionably
pre-empt developmental markers?
Assuming a 1000 species ecosystem, and 1 successful experiment every day to discern a single two-way relationship, we would need 1,368 years to go through all possibilities. If we look for 3 way interactions, we would need 454,844 years

Can we predict the next pandemic?
Can we predict future mutations? Can we define the "edge of emergence"?

Digital Twins for complex systems

Chattopadhyay, Ishanu, Kevin Wu, Jin Li, and Aaron Esser-Kahn. "Emergenet: Fast Scalable Pandemic Risk Assessment of Influenza A Strains Circulating In Non-human Hosts." (2023). Under Review in Nature
PREEMPT
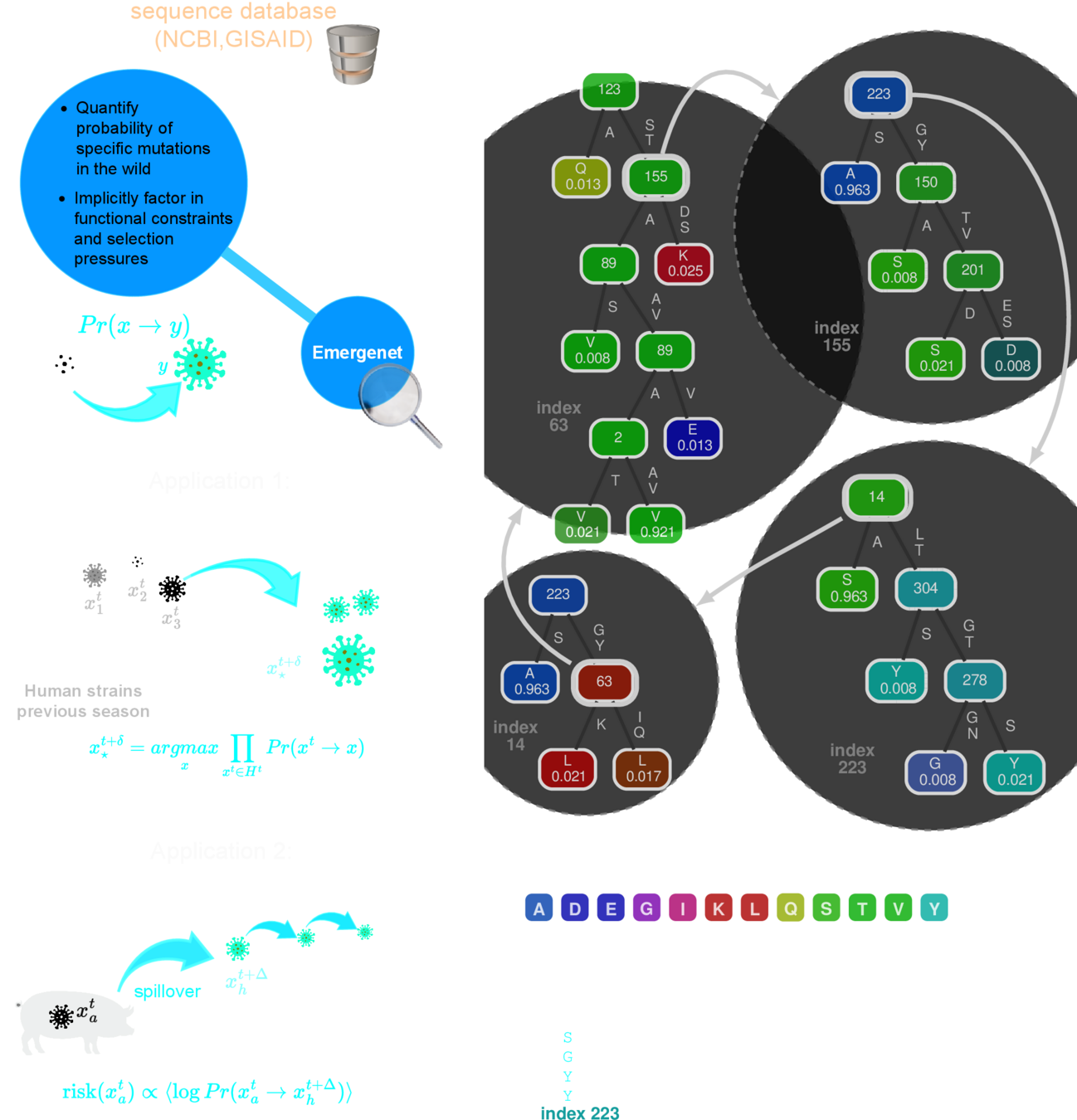


Q-Net
recursive forest
A Math Solution to a Hard Biological Problem
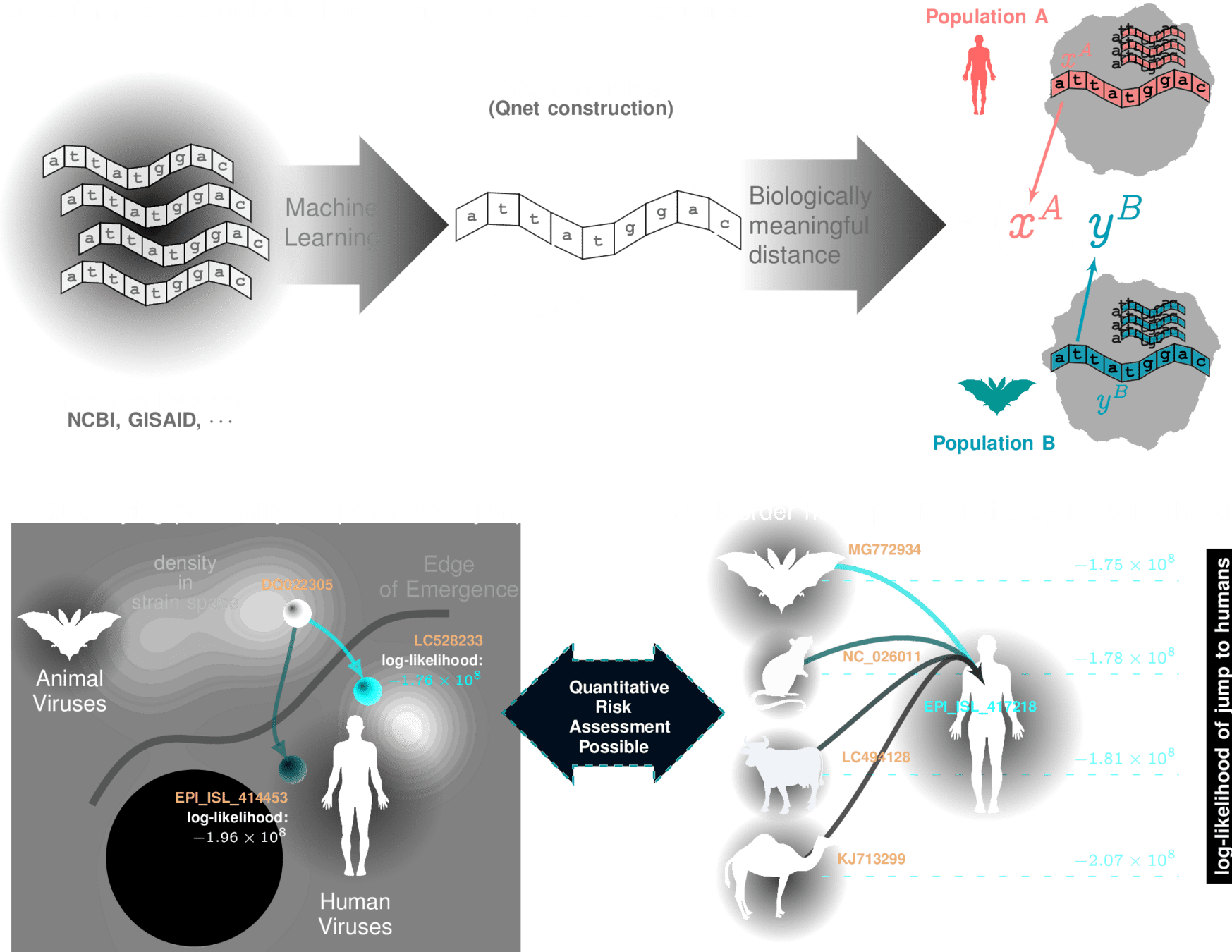
we can tell if new strain will adapt to humans
Influenza Risk Assessment Tool (IRAT) scoring for animal strains
slow (months), quasi-subjective, expensive
*https://www.cdc.gov/flu/pandemic-resources/monitoring/irat-virus-summaries.htm
24 scores in 14 years
~10,000 strains collected annually
CDC
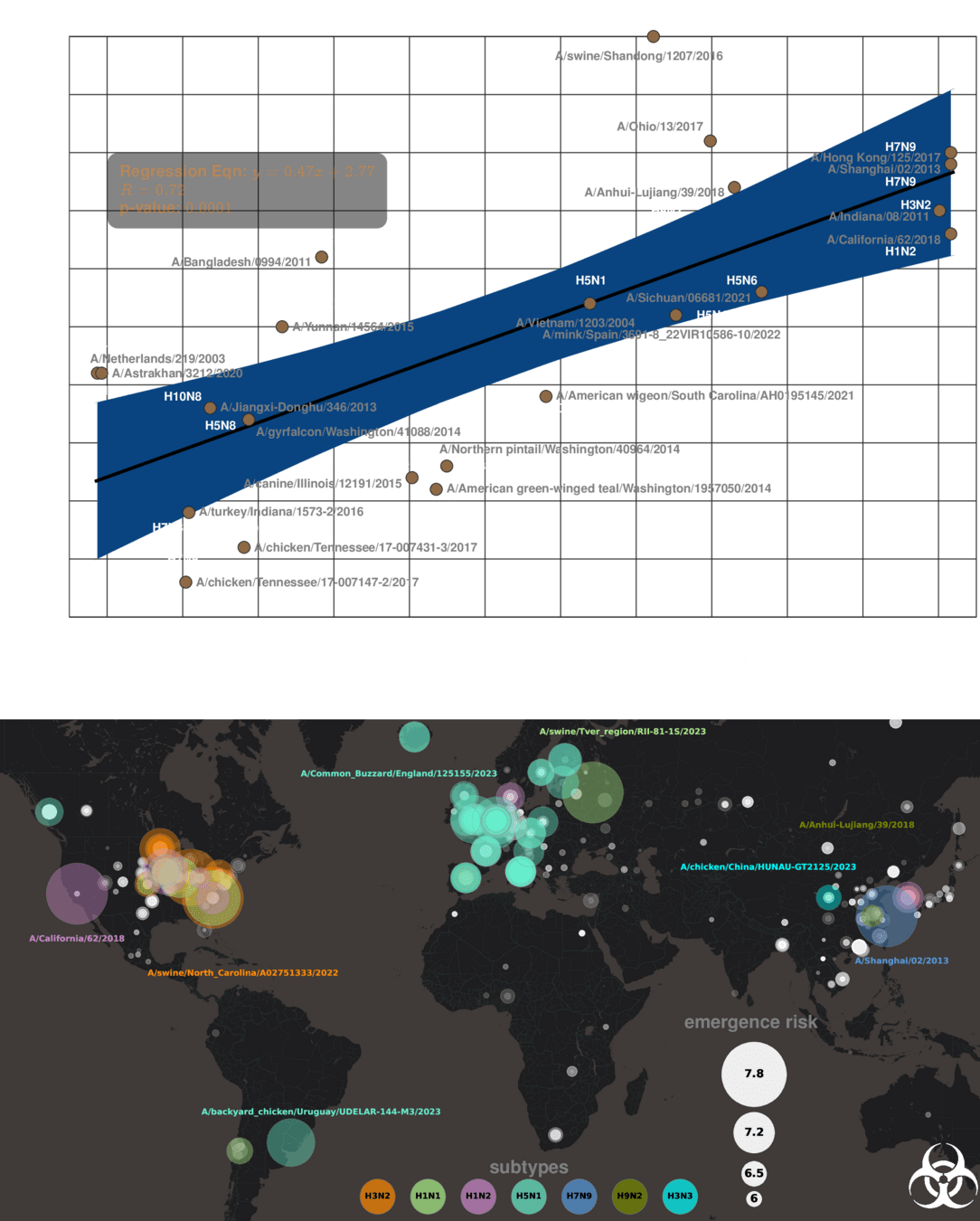
Emergenet time: 1 second

Stamping Out the Next Pandemic **Before** The First Human Infection
BioNorad
Apply the same "tech" to the microbiome modeling problem

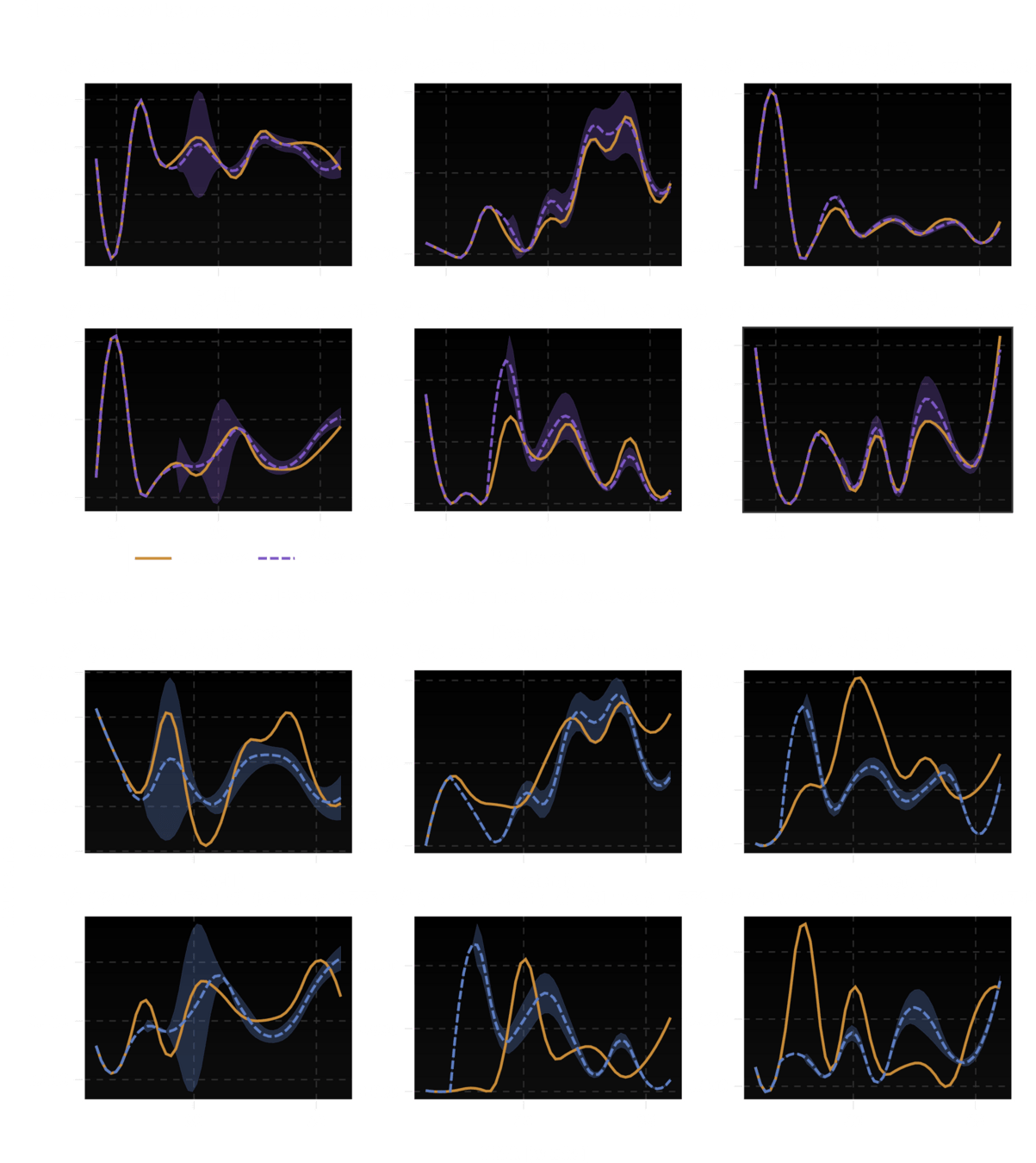
Ability to "fill in" missing data is equivalent to making trajectory forecasts
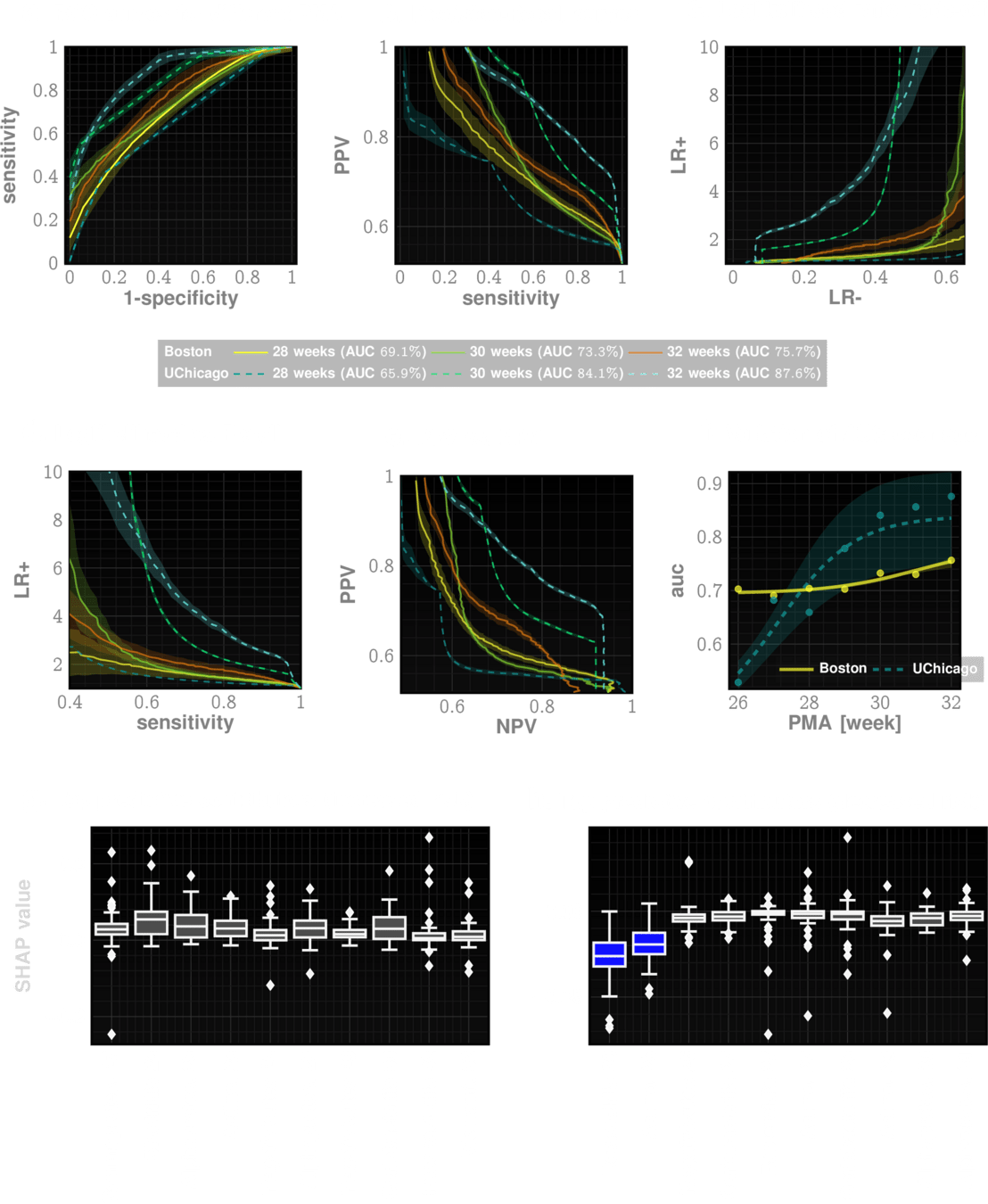



Our risk measure is highly predictive and actionable

Which entities are most predictive?
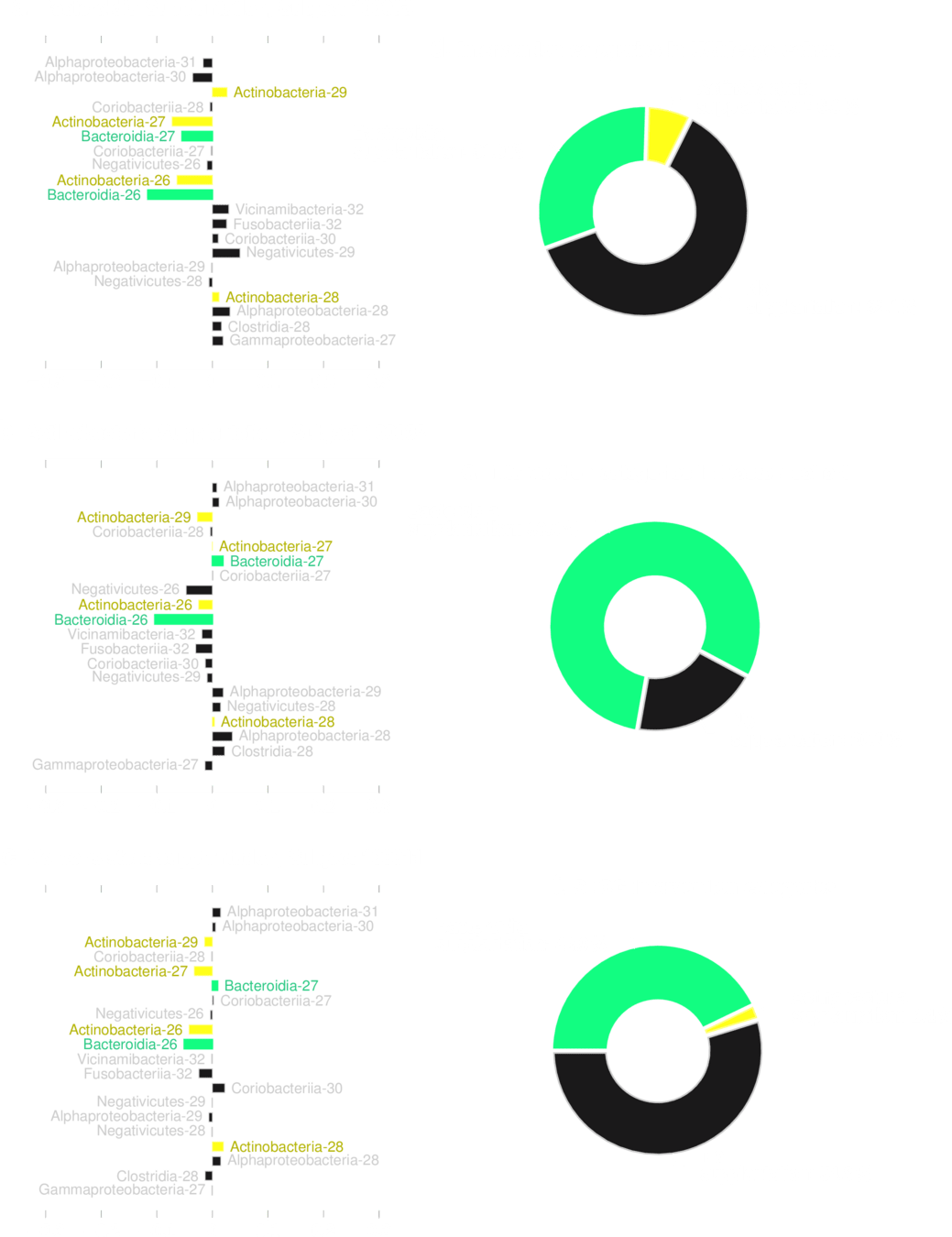
No transplantation is guaranteed to work reliably
Predicted to reduce
risk reliably
Predicted to reduce
risk reliably
Future
Answer the question: "what is a healthy microbiome?"
Explicit supplantation profiles that are tuned to individual ecosystems
Bioreactor experiments
Research Summary
Transform bio-surveillance
Transform modeling of complex systems
Transform early diagnosis
Democratize AI unleashing its power for social good
"Empowering Minds, Transforming Futures: AI Innovation with Purpose."
Enabling technology to reduce human suffering
Enabling the Vulnerable
To Have a Voice
Early Life Interventions to maximize human potential
Health Equity
"Fusing Innovation with Compassion:
Advancing AI Research for the Greater Good."
Q&A
ishanu chattopadhyay
Digital Twin of the Human Microbiome
University of Chicago Medicine

Nicholas Sizemore
Kaitlyn Oliphant
Erika Claud

pip install qbiome
import qbiome
from qbiome.data_formatter import DataFormatter
from qbiome.quantizer import Quantizer
from qbiome.qnet_orchestrator import QnetOrchestrator
from qbiome.forecaster import Forecaster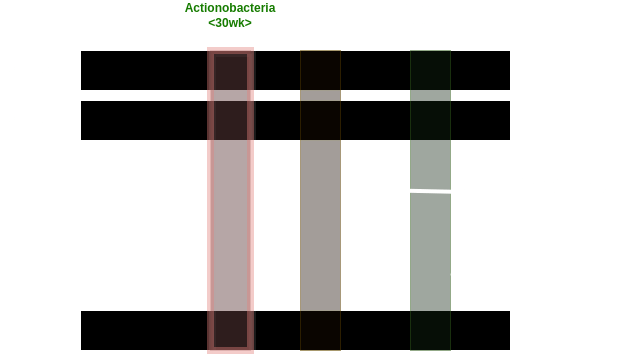
This is a general method!



E-Net
recursive forest
E-distance
a biologically informed, adaptive distance between strains
This distance is "special"
smaller distances imply a quatitatively high probability of spontaneous jump
$$J \textrm{ is the Jensen-Shannon divergence }$$
Sanov's Theorem & Pinsker's Inequality
Theorem
stable profile \(x_{h}\), "well-adapted" \(\Rightarrow Pr(x_h\rightarrow x_h) \approx 1 \)
For "new" profile \(x_{a}\), \( \displaystyle \theta(x_{a},x_{h}) \approx 0 \)
Assume:
Then, we have:
we can tell if new profile will be stable
A Math Solution to a Hard Biological Problem
Biology-aware Perturbations to "reconstruct" missing data
sample
?
Can i meaningfully perturb aundunce values?
Can we fill them in if they are missing?
Risk of Time-stamped Microbial Profile to lead to Developmental Deficit
The Zero Profile

Q-net inferred with typical patients

Q-net inferred with patients with neurodevelopmental deficit
initial microbiome
current microbiome
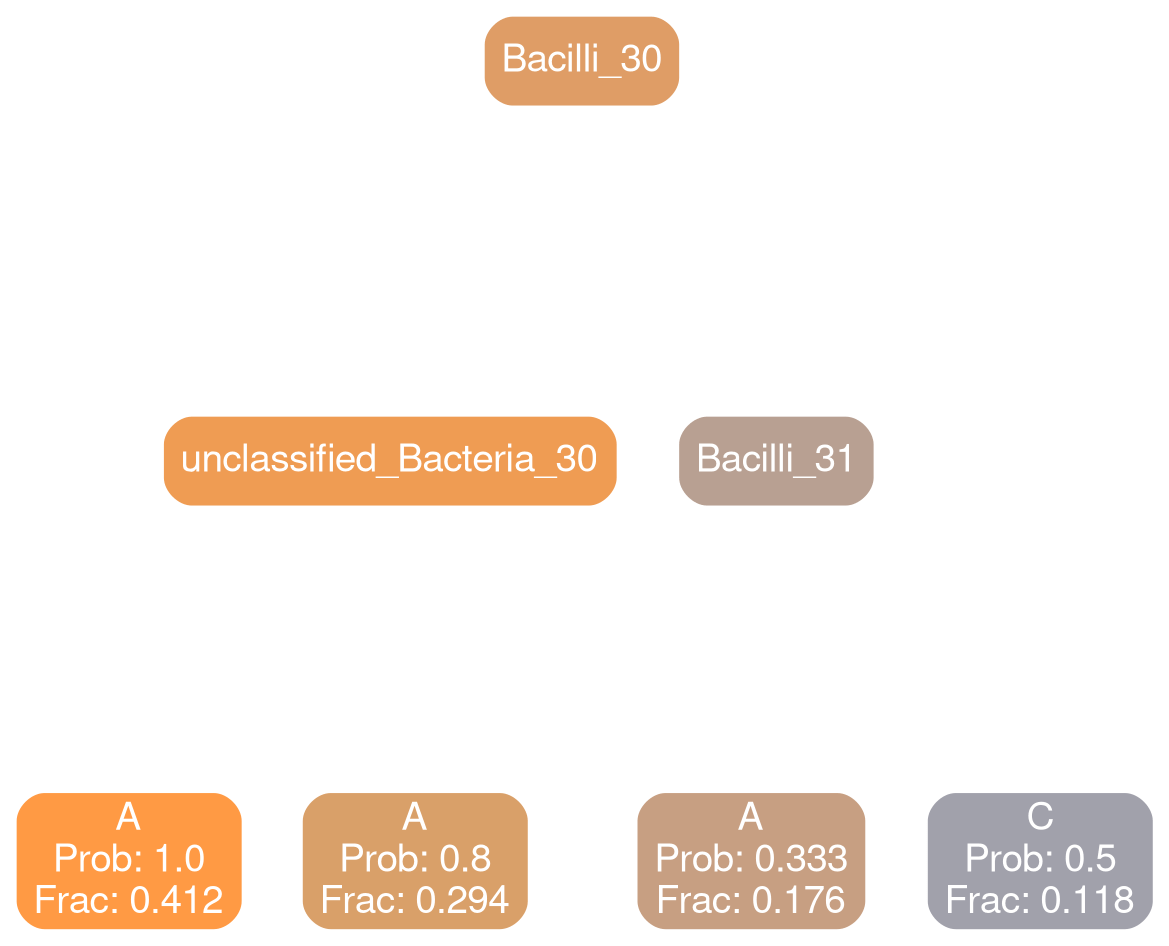
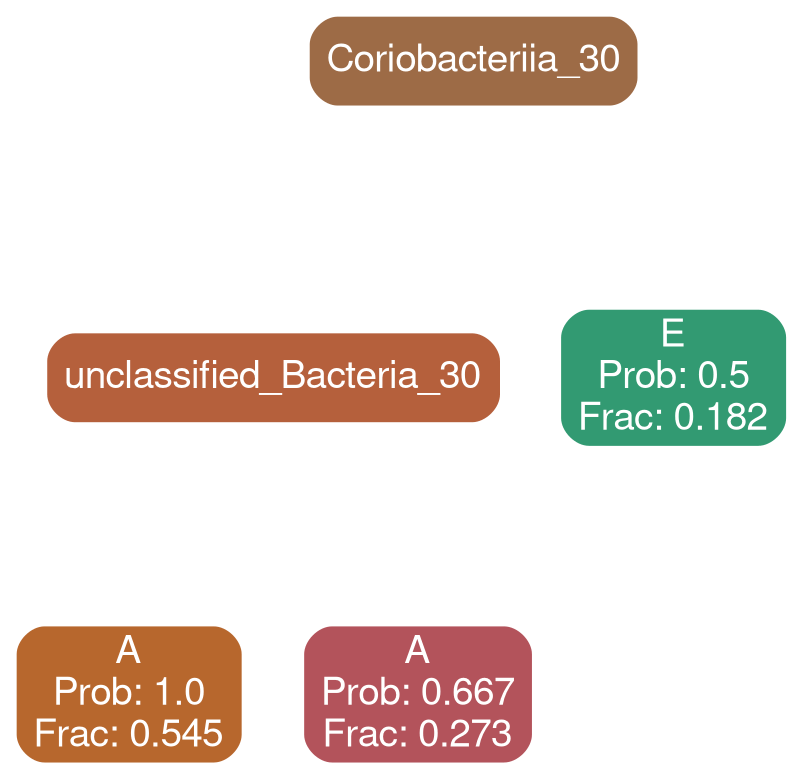
Actinobacteria 30
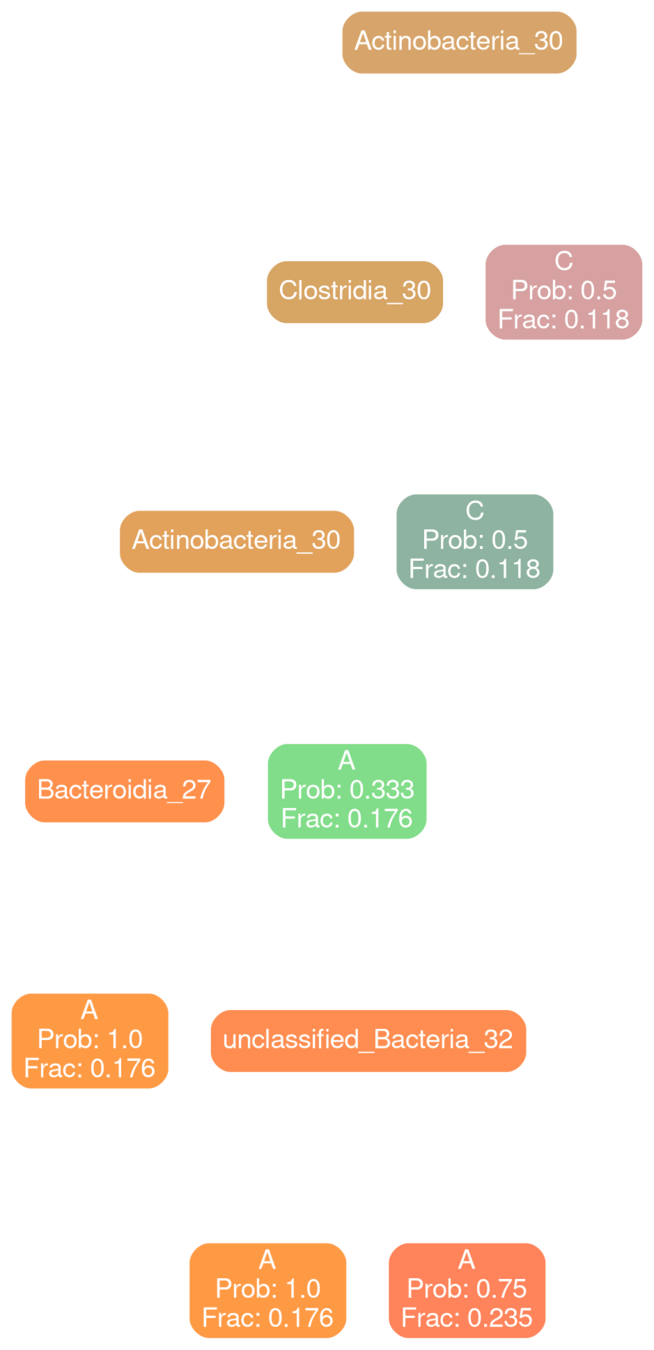
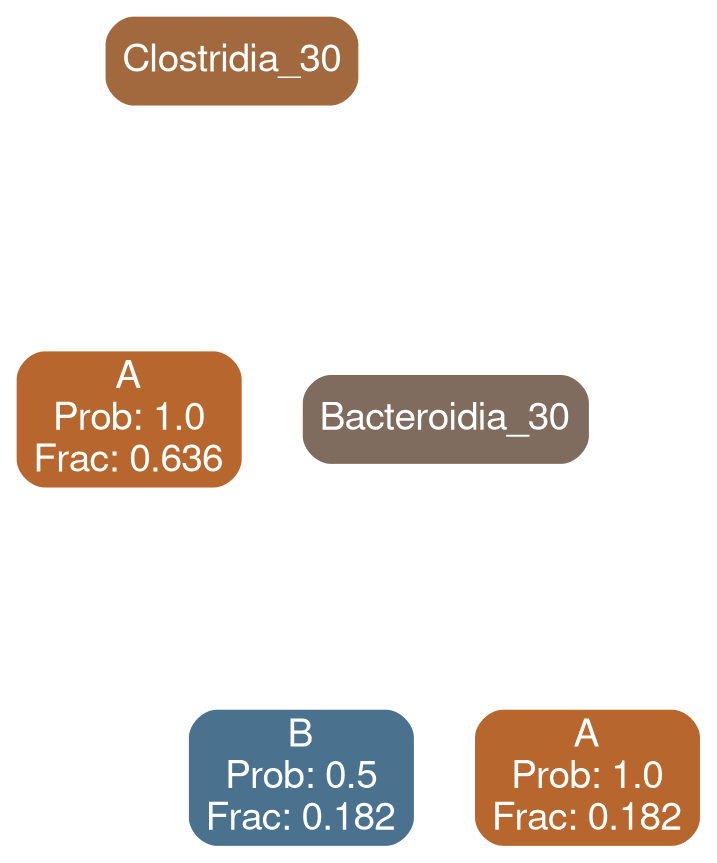
Bacilli 30
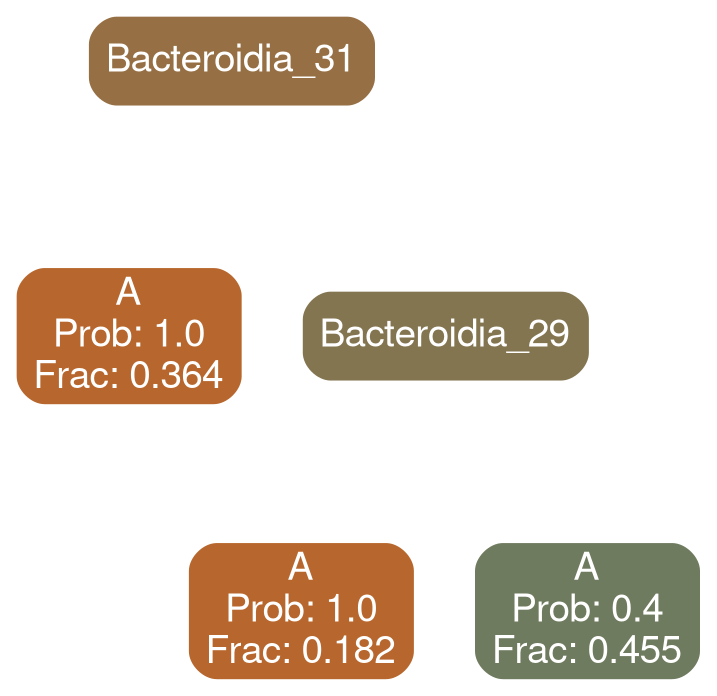
Bacteroidia 30
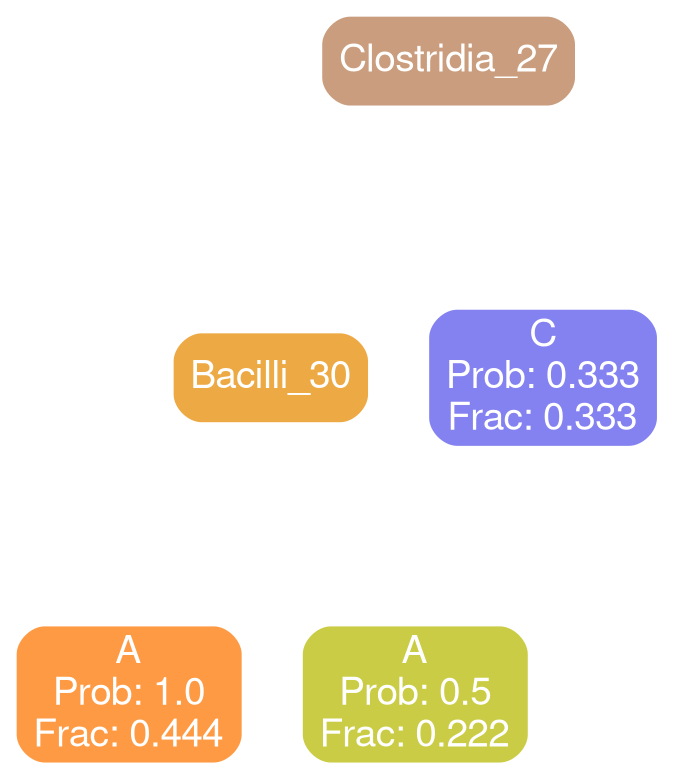
Coriobacteria 32
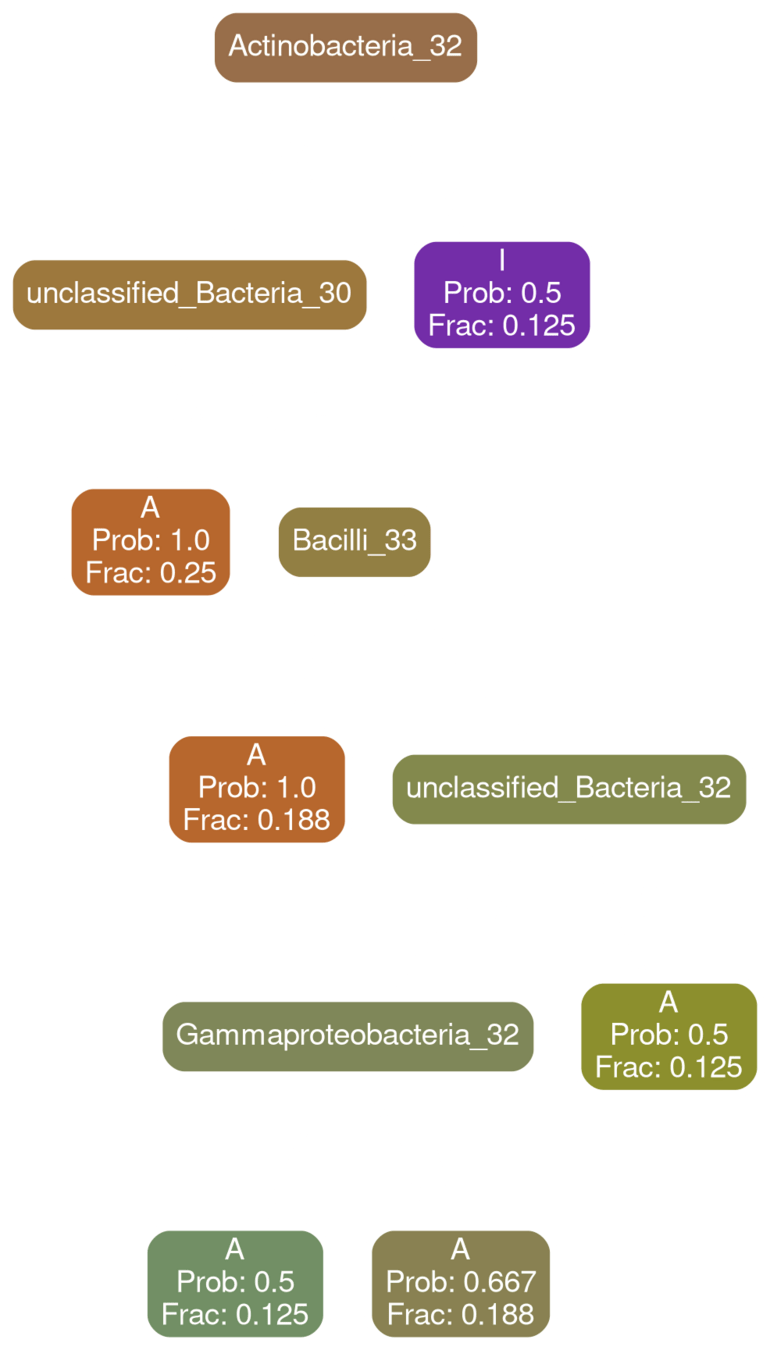
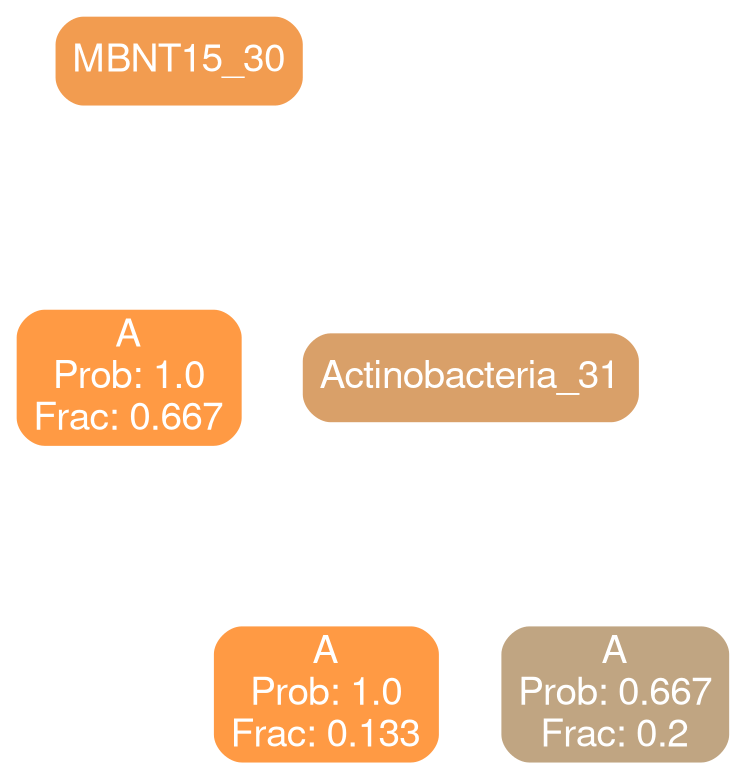
Gammaproteobacteria 32
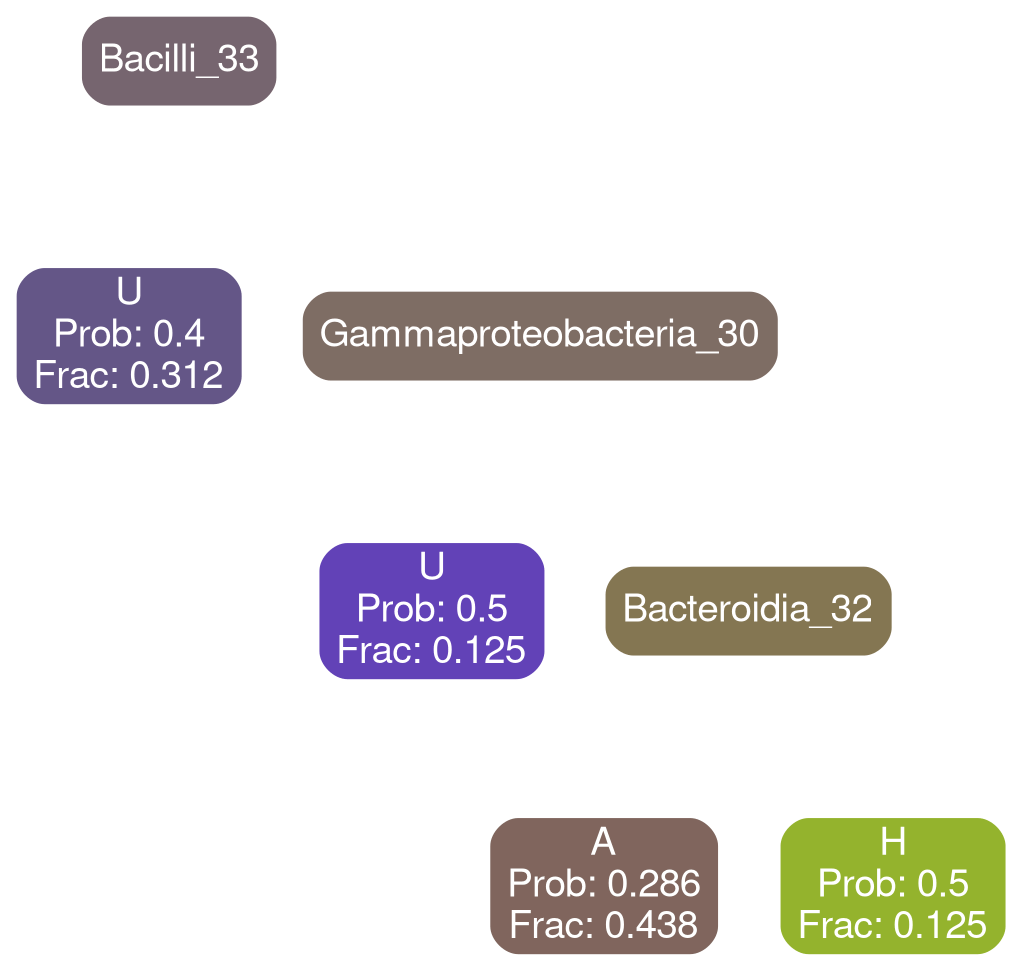
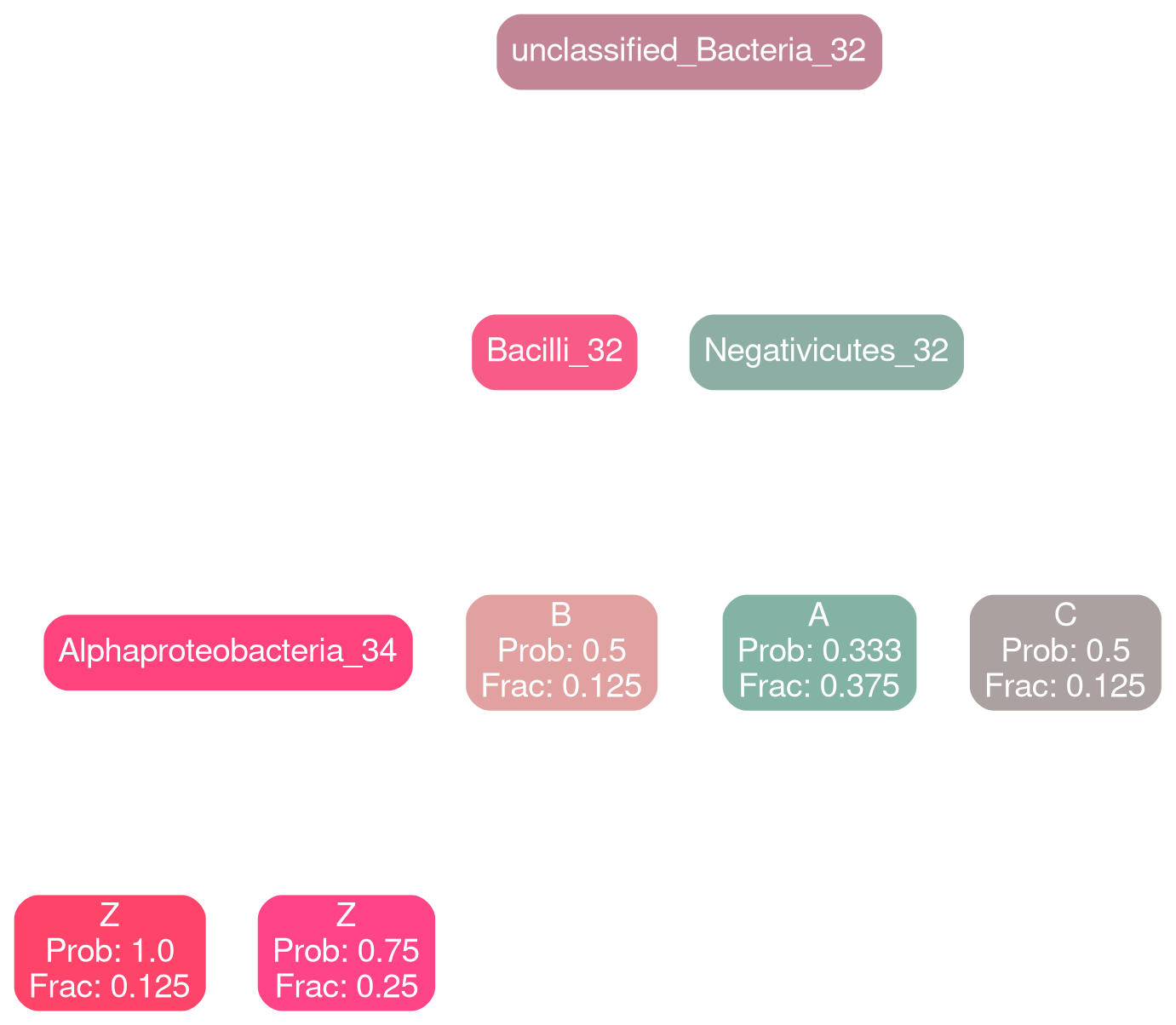
AHCTG
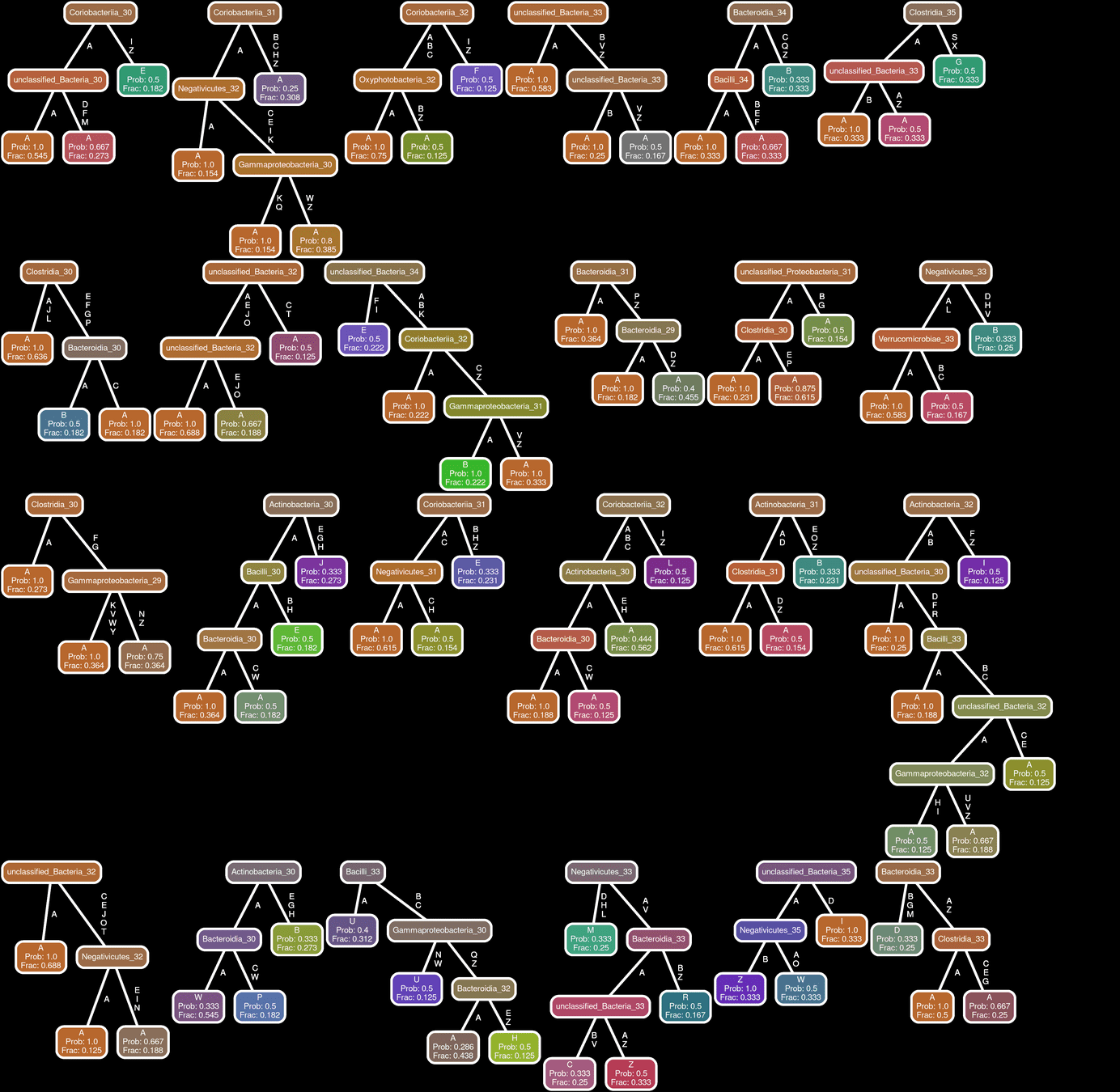
SHCTG
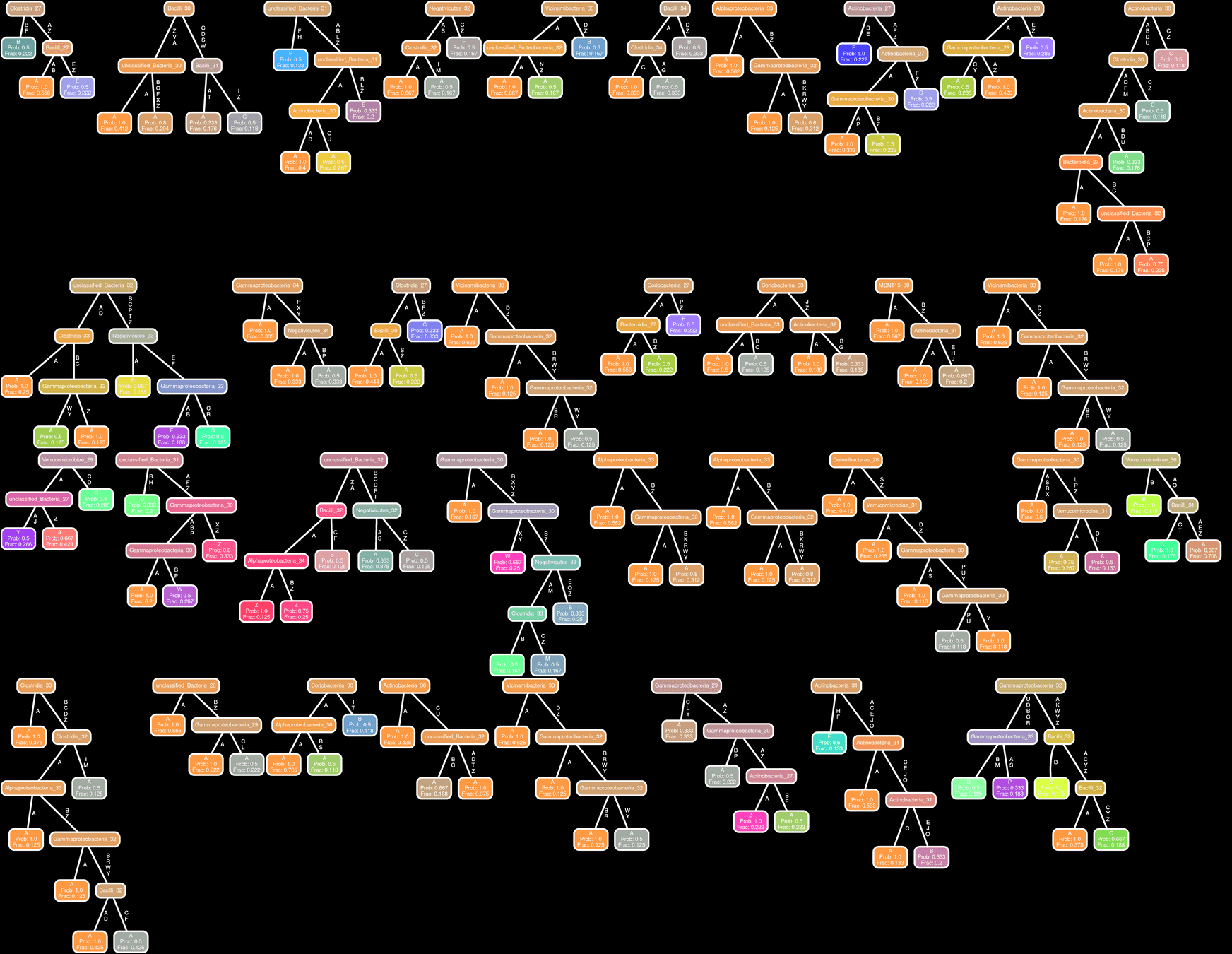
All patients All Entities
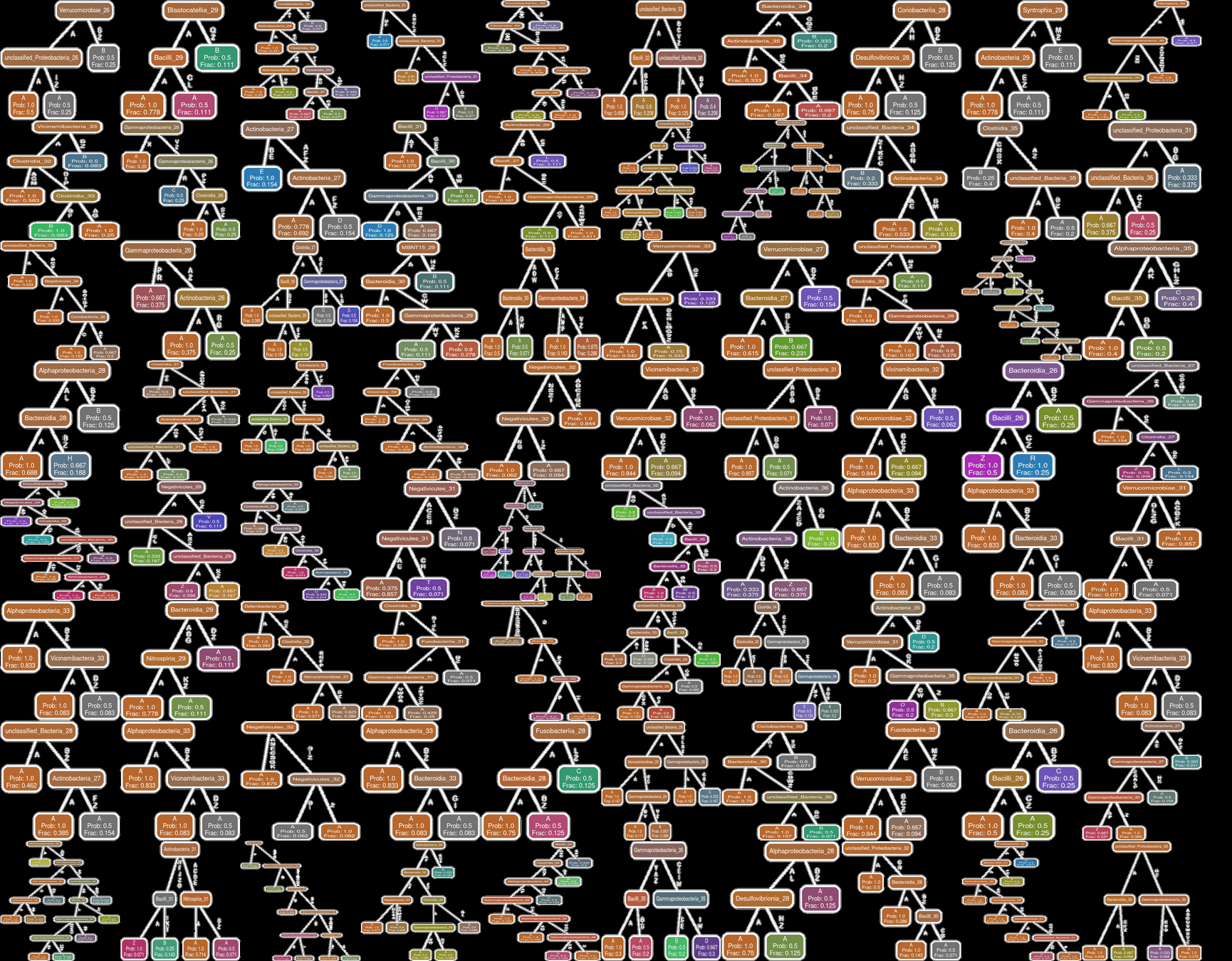
Feeding Variables added
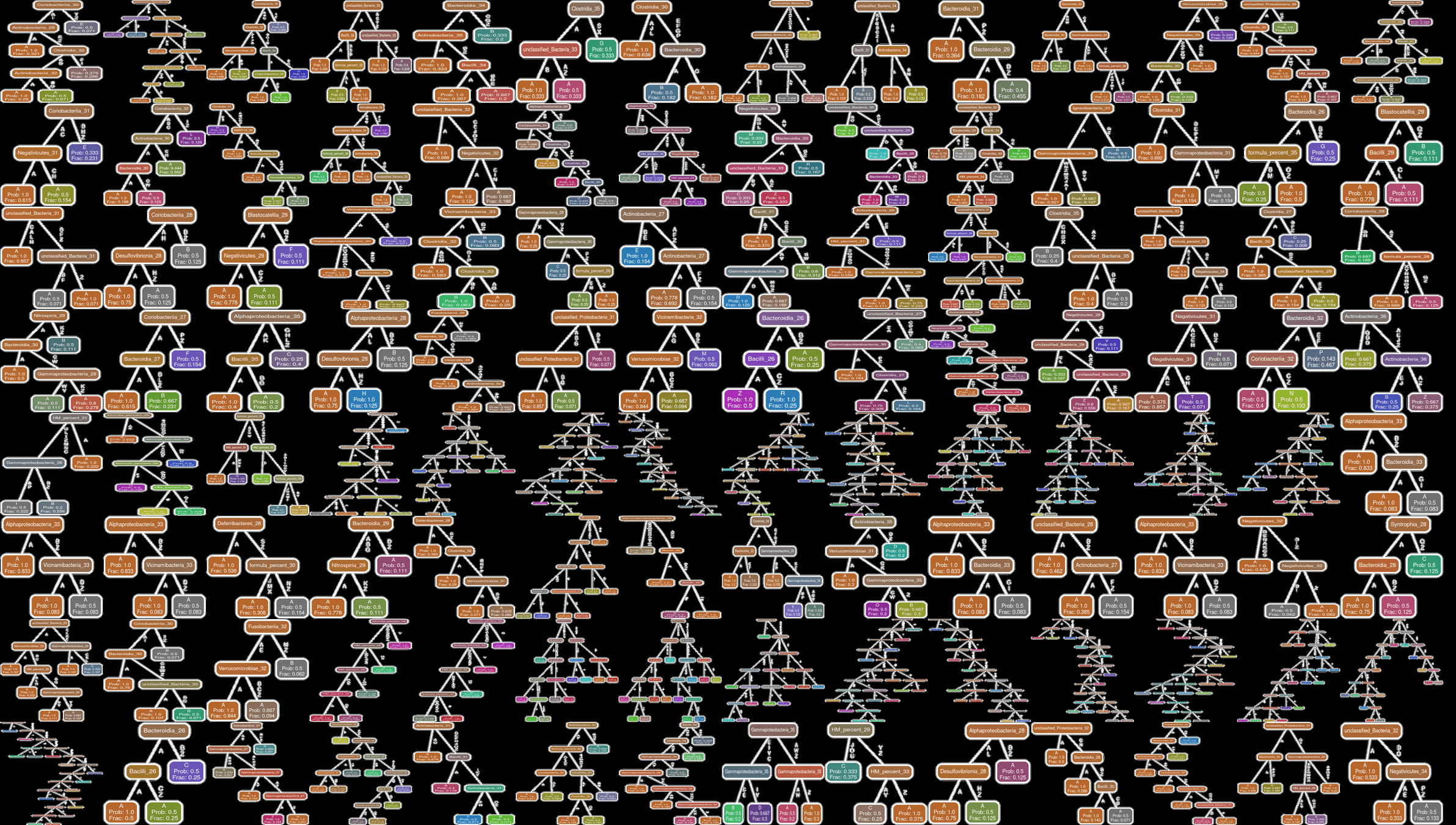

Ability to "fill in" missing data is equivalent to making trajectory forecasts

Our risk measure is highly predictive and actionable

Which entities are most predictive?

Supplantation MUST be personalized
Supplantation MUST be personalized
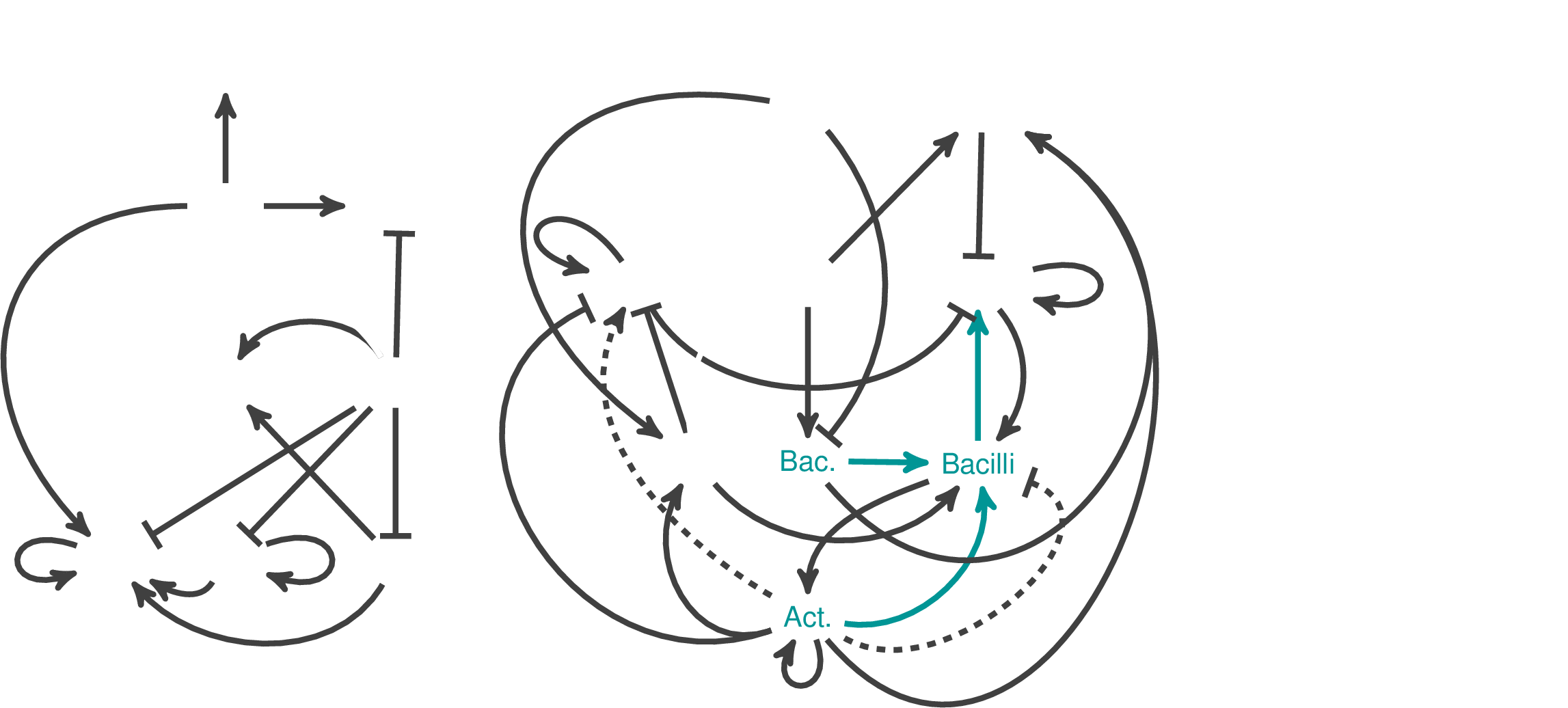
Network Interpretations?
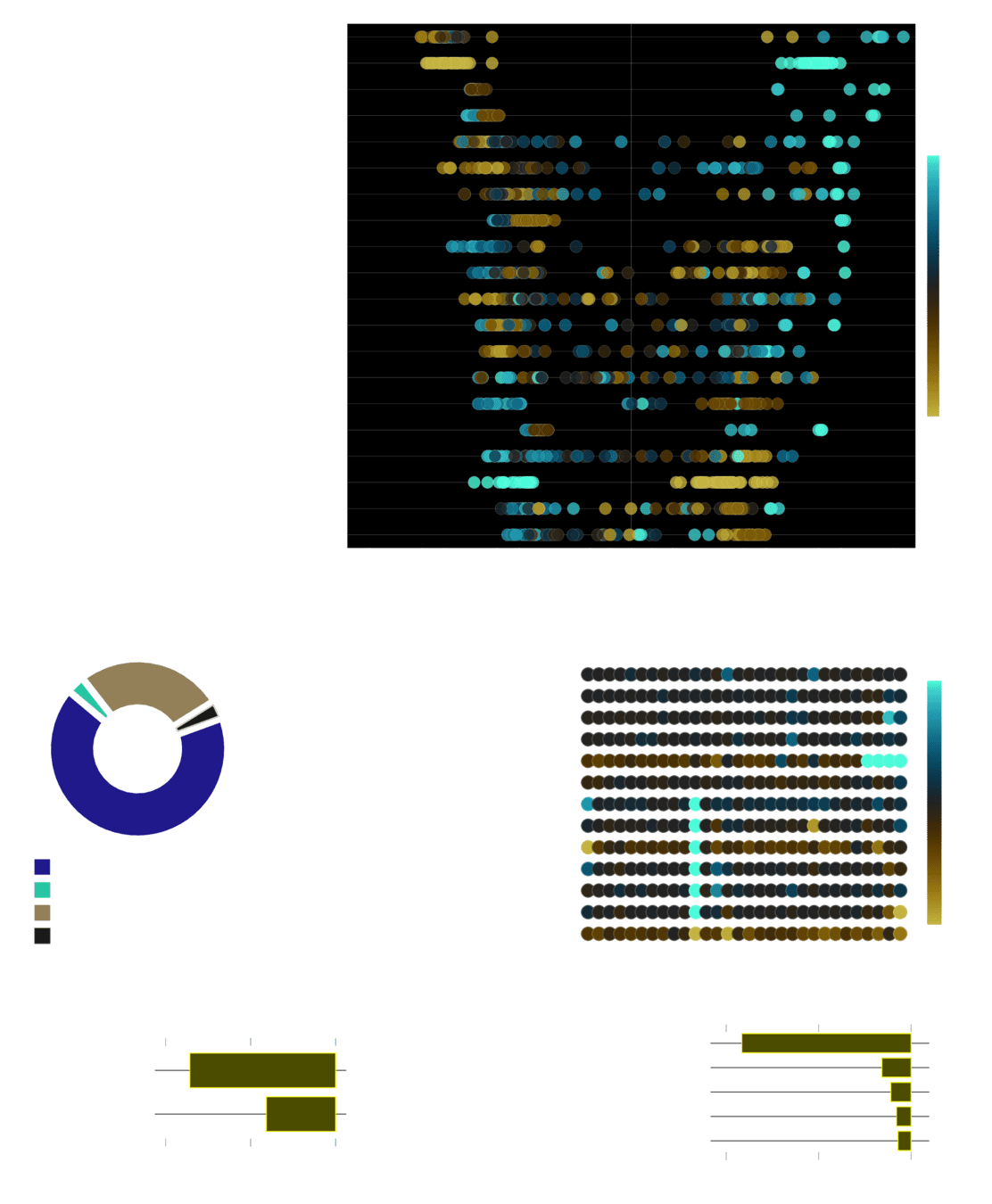
Effect of Clinical Variables
Influenza Risk Assessment Tool (IRAT) scoring for animal strains
Can we replicate IRAT scores*?
slow (months), quasi-subjective, expensive
*https://www.cdc.gov/flu/pandemic-resources/monitoring/irat-virus-summaries.htm

genomic analysis
receptor binding
animal
transmission
antivirals available
population immunity
human infections
animal
hosts
global prevalence
antigenic novelty
disease severity