Using an escape calculator to assess antigenic impacts of SARS-CoV-2 mutations
Jesse Bloom
Fred Hutch Cancer Center / HHMI
Deep mutational scanning can map all RBD mutations that escape antibody binding
RBD
fluorescently labeled antibody
yeast
fluorescent tag on RBD
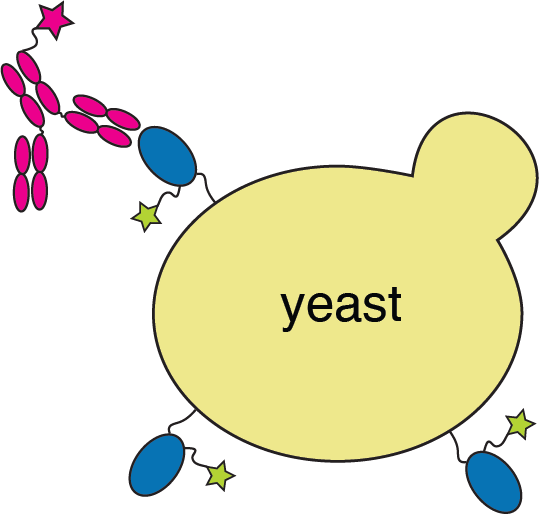
Experiments combine flow cytometry and deep sequencing of a library of yeast expressing all RBD mutants
For interactive escape map, see: https://jbloomlab.github.io/SARS-CoV-2-RBD_MAP_LY-CoV555/
Escape map from one antibody (LY-CoV555, ie bamlanivimab). Peaks indicate sites where mutations escape binding.
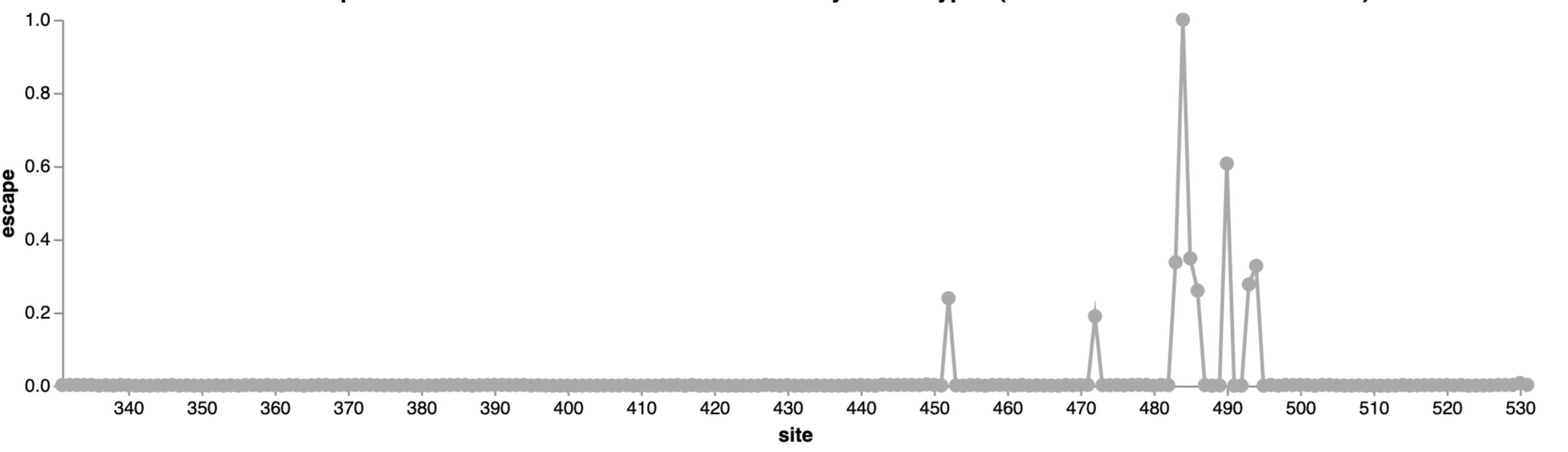
484
452
490
Infection / vaccination elicit polyclonal antibodies
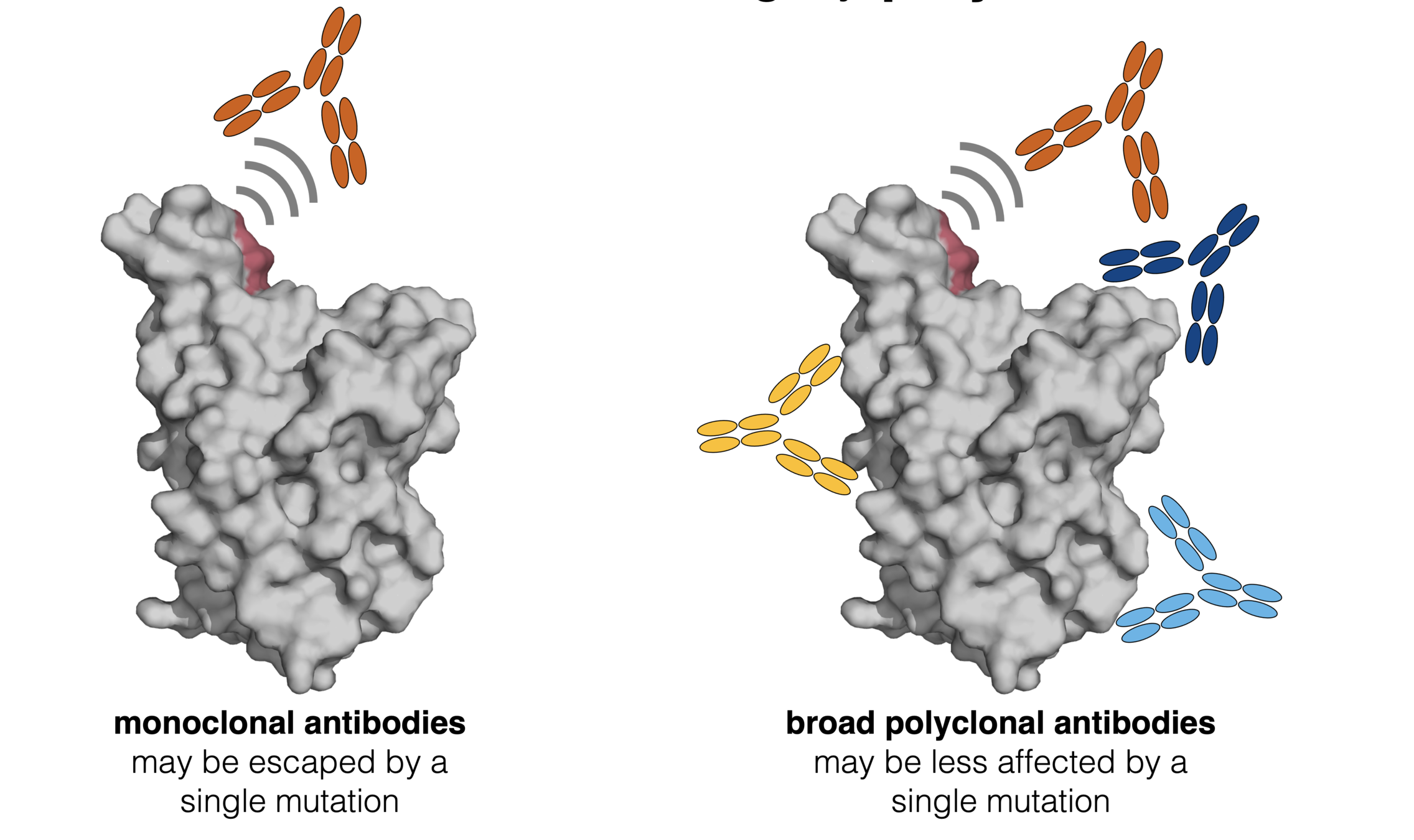
How do mutations affect polyclonal antibodies? First, consider an equal mix of three monoclonal antibodies.

Interactive version of this mini example is at https://jbloomlab.github.io/SARS2_RBD_Ab_escape_maps/mini-example-escape-calc/
LY-CoV555 is escaped at both sites 484 and 490, so mutating either site has same overall effect
Average escape across all antibodies

Mutating site 484 or 490 eliminates neutralization by antibody LY-CoV555, as reflected in thick black line showing average
Interactive version of this mini example is at https://jbloomlab.github.io/SARS2_RBD_Ab_escape_maps/mini-example-escape-calc/
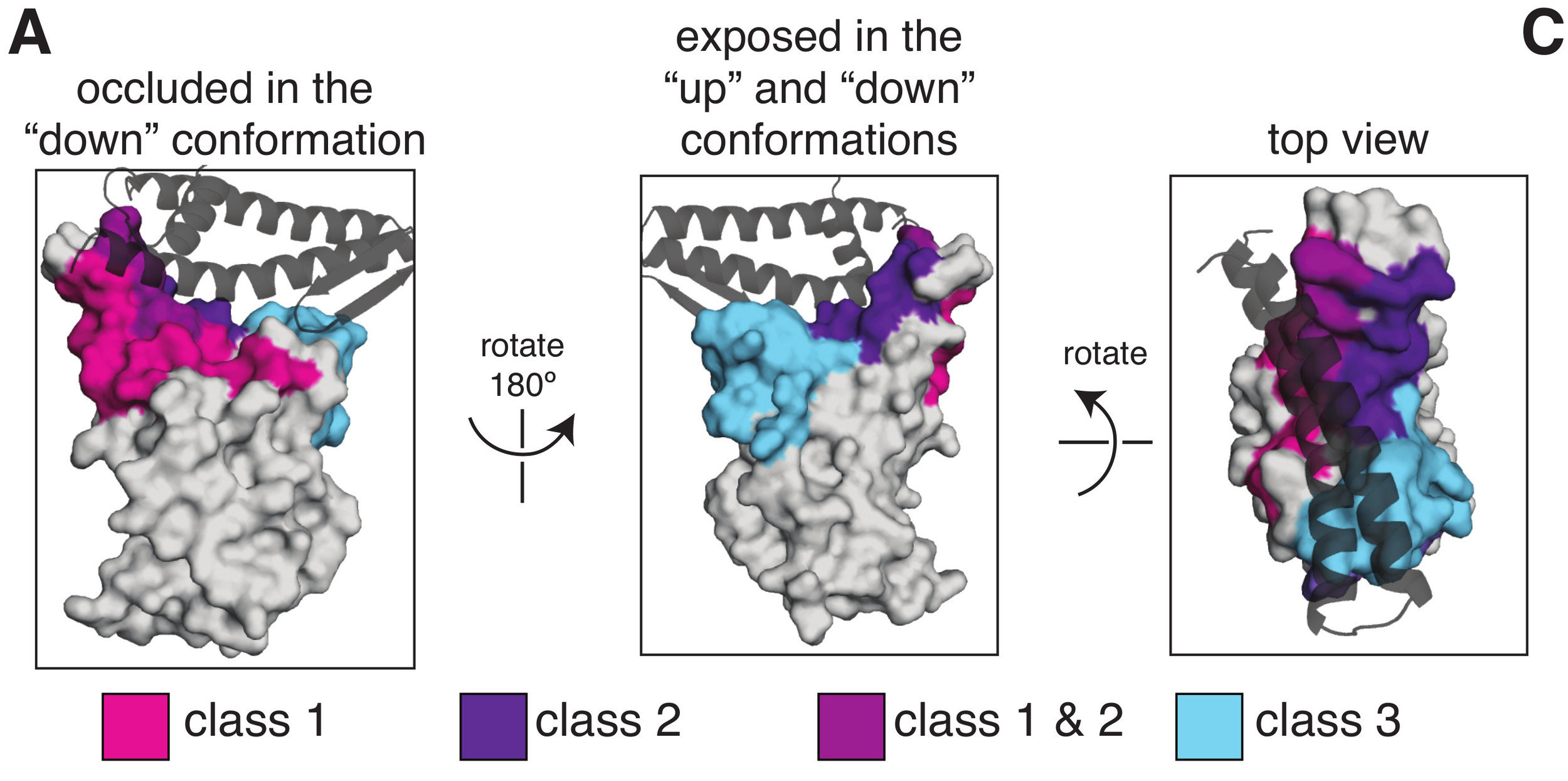
Note this approach is a fine-grained method for defining epitopes, instead of simple discrete classification
Antibody-escape calculator extends this principle to deep mutational scanning data for ~1,500 different human antibodies
Escape calculator is described in Greaney et al (2022), and is available at https://jbloomlab.github.io/SARS2_RBD_Ab_escape_maps/escape-calc/
36 antibodies mapped by Tyler Starr & Allie Greaney in Bloom lab, from early SARS-CoV-2 strains
Antibody-escape calculator extends this principle to deep mutational scanning data for ~1,500 different human antibodies
Escape calculator is described in Greaney et al (2022), and is available at https://jbloomlab.github.io/SARS2_RBD_Ab_escape_maps/escape-calc/
36 antibodies mapped by Tyler Starr & Allie Greaney in Bloom lab, from early SARS-CoV-2 strains
1,522 (!) antibodies mapped by Sunney Xie, Richard Cao, Fanchong Jian, et al at Peking University. From early strains, BA.1, & patients with prior SARS-CoV-1 infection. See here.
Using the interactive escape calculator
https://jbloomlab.github.io/SARS2_RBD_Ab_escape_maps/escape-calc/
We also provide a Python module for batch calculation of escape scores
Module can be used to antigenically score sequences
Here is pilot nextstrain tree with escape scores of BA.2 and descendants created in collaboration with Trevor Bedford.