Lentiviral-pseudotype deep mutational scanning of viral entry proteins
Jesse Bloom
Fred Hutch Cancer Center / HHMI
These slides at https://slides.com/jbloom/lentiviral-dms
Most viral vaccines induce neutralizing antibodies to the viral entry protein
All viruses have one or more entry proteins that bind receptor and then fuse with the cell membrane:
- SARS-CoV-2 spike
- influenza hemagglutinin
- HIV envelope protein
- Lassa virus glycoprotein
- Nipah virus G and F proteins
- RSV G and F proteins
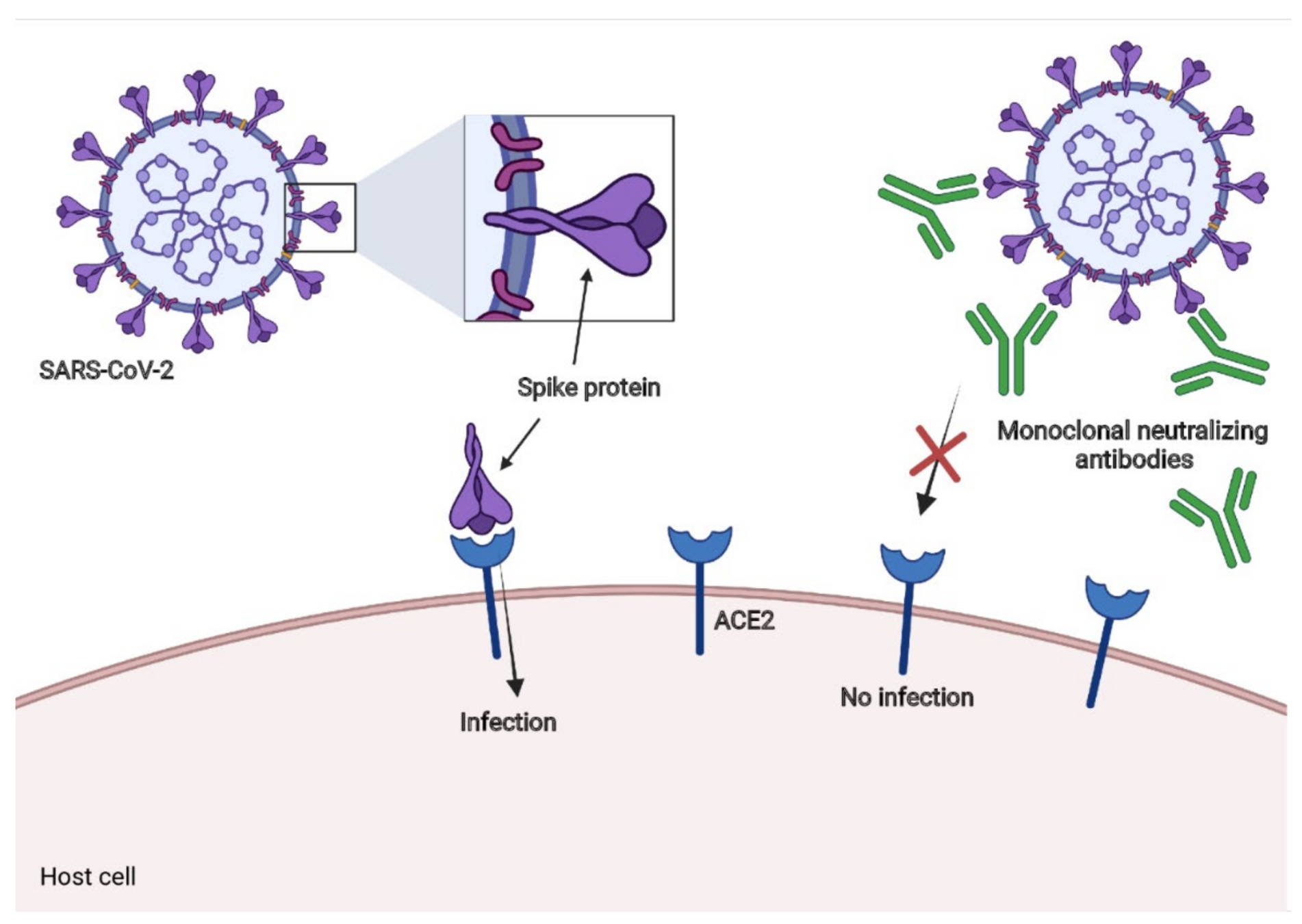
Evolution of viral entry proteins can escape antibodies and reduce vaccine efficacy
Example: new SARS-CoV-2 variants like XBB have accumulated 40 amino-acid substitutions/deletions in spike, thereby escaping all first-generation antibody drugs and greatly reducing neutralization by serum antibodies elicited by early vaccines. (Plot from Cao et al)
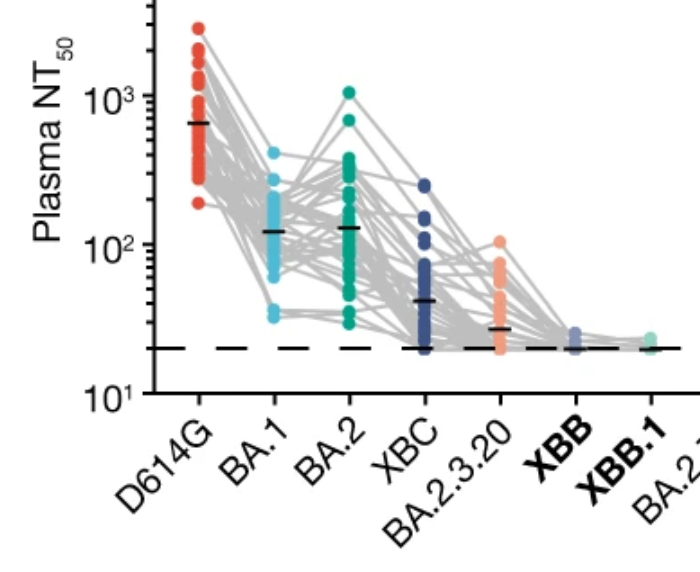
Prospective mapping of mutations?
If we could prospectively map which mutations are tolerated, and how they affect antibody escape, then we could:
- Better design/select antibodies that are resistant to escape
- Improve surveillance and vaccine updates by interpreting effects of mutations
Our lab developed yeast-display deep mutational scanning to generate this information for SARS-CoV-2 RBD

We would like a general system to produce similar information for the full entry proteins of many viruses
Lentiviral pseudotyping provides a safe way to study many viral entry proteins
- Many viruses have entry proteins amenable to lentiviral pseudotyping.
- However, traditional pseudotyping does not create genotype-phenotype link.
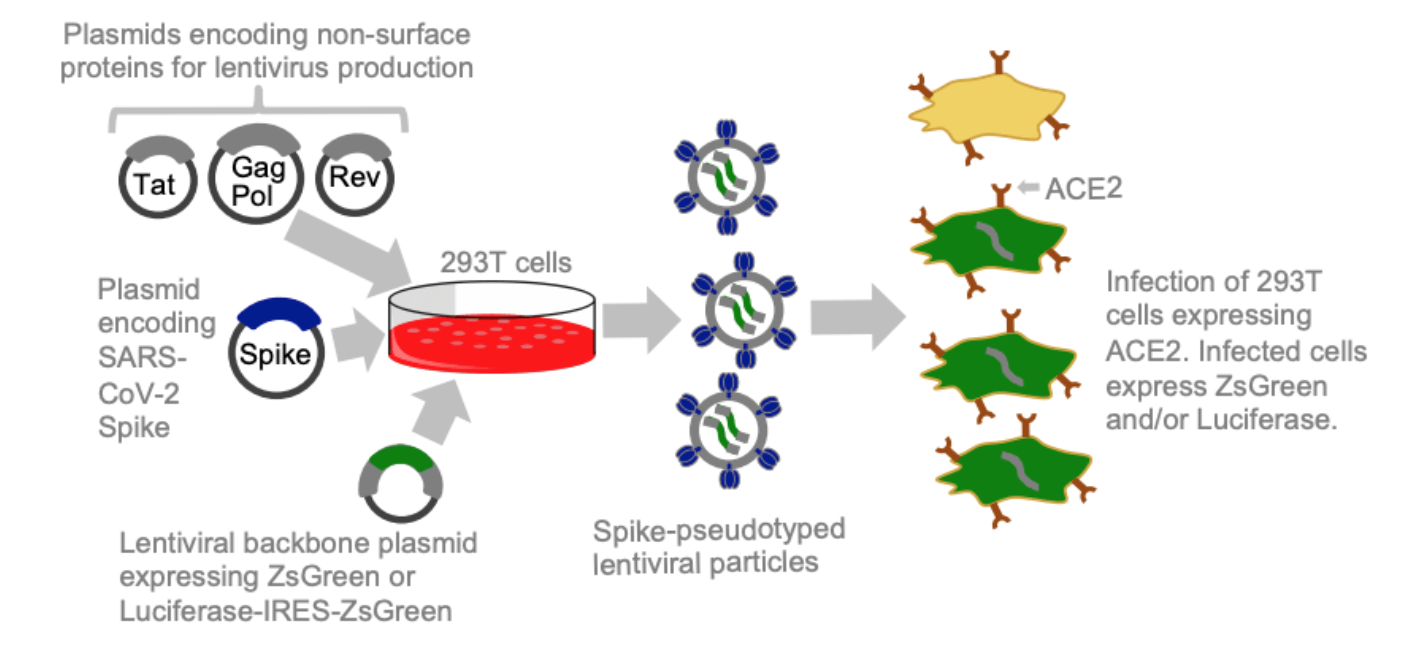
Two-step method to create genotype-phenotype linked spike-pseudotypes
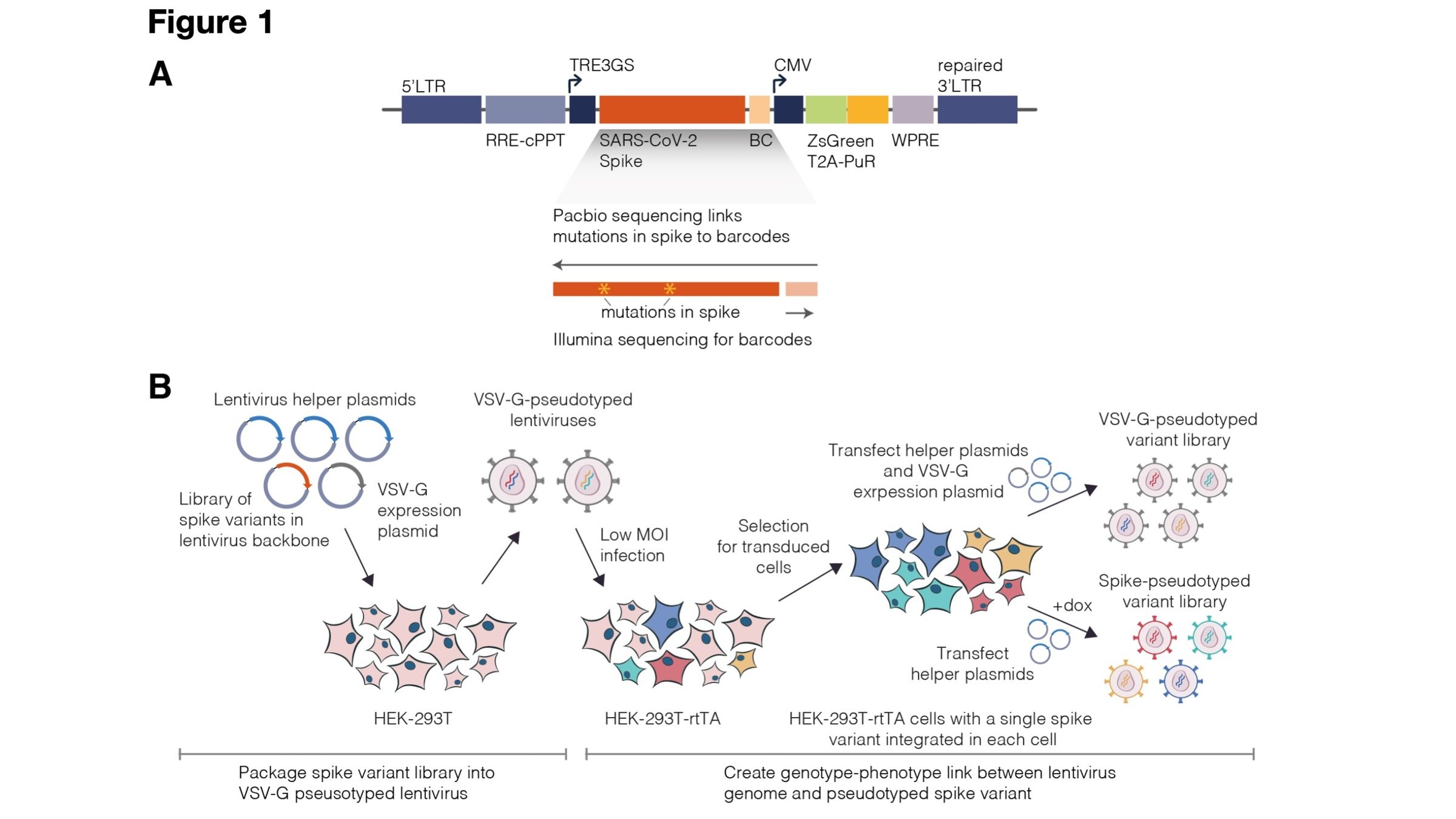
Sequencing measures relative amounts, so include neutralization standard
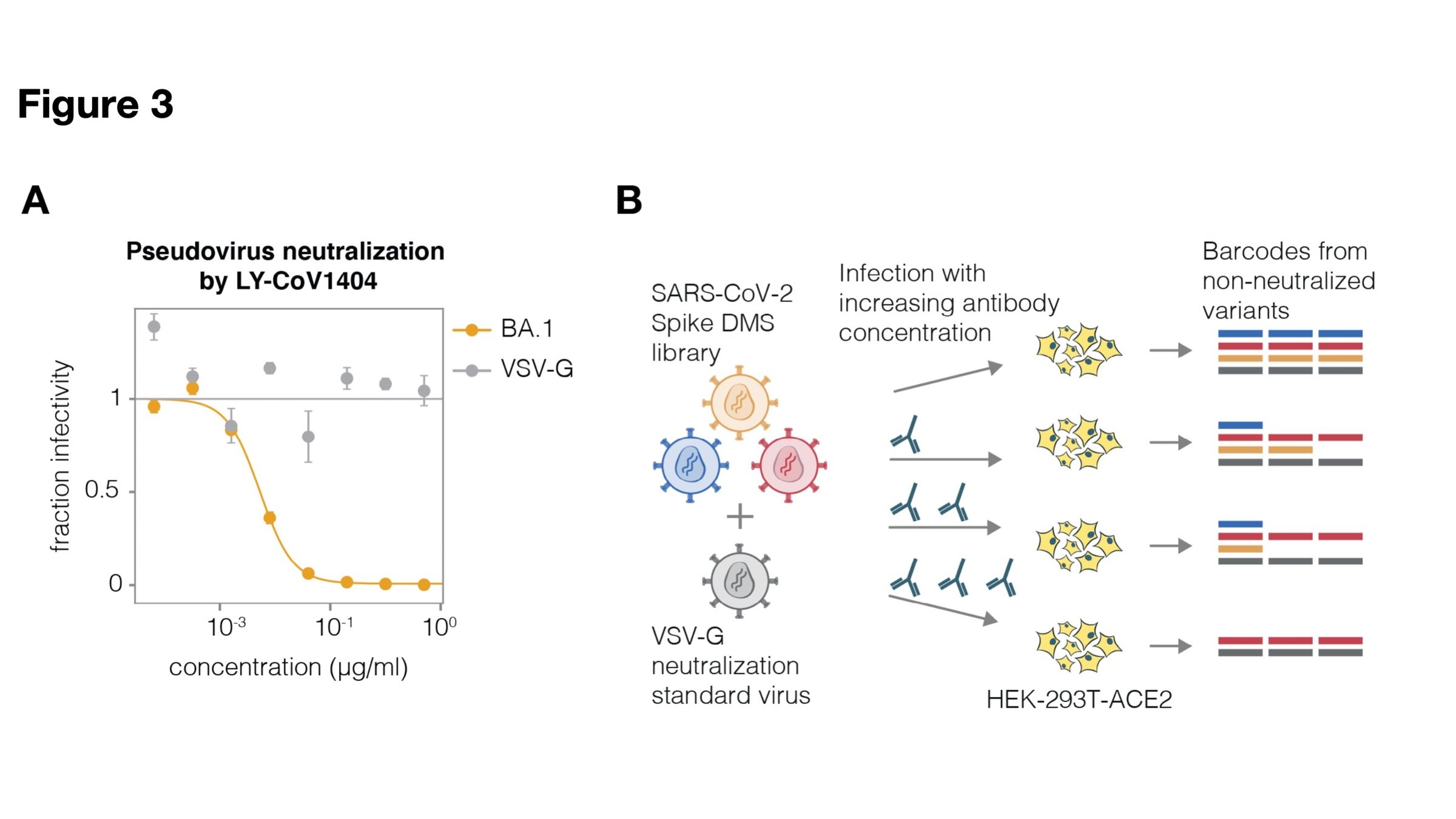
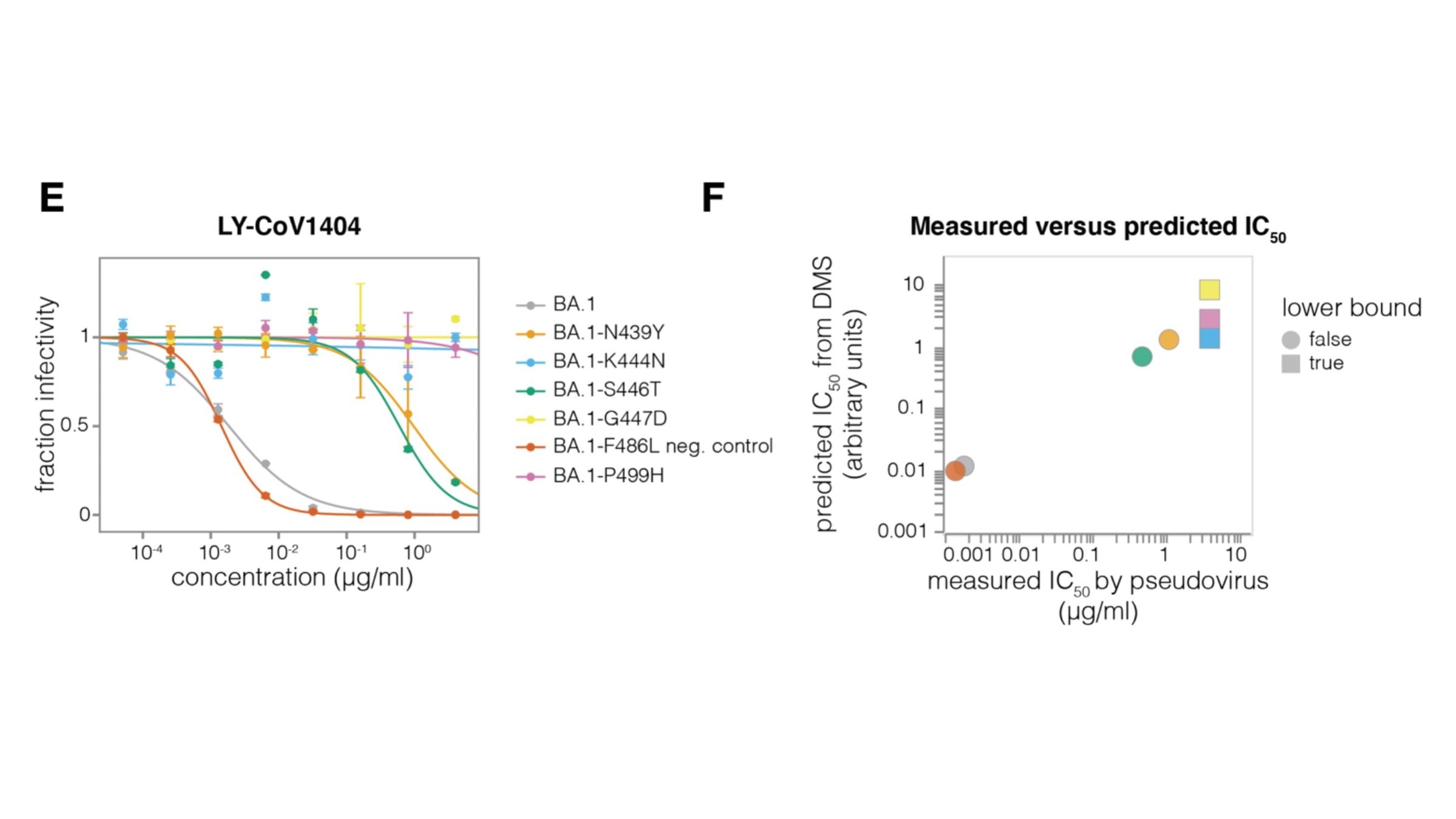
Example: RBD antibody LY-CoV1404
Validates very well
Example: S2 antibody CC67.105
This method also works for other viral entry proteins
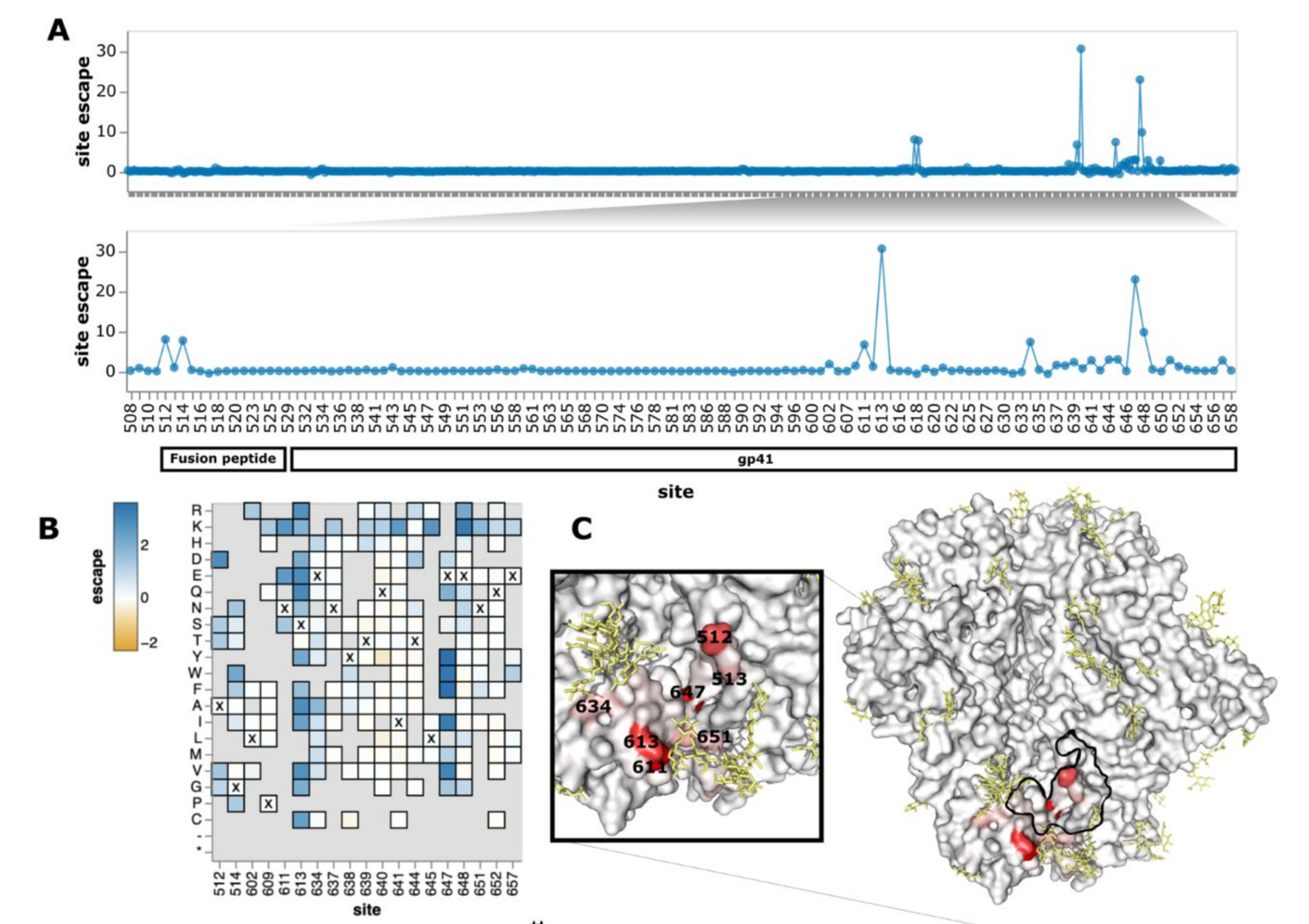
Map of how all mutations to HIV envelope (Env) affect escape from antibody PGT151
We can apply method to emerging viruses that may pose a future risk
Prospective maps of antigenic and functional effects of mutations to H5N1 HA
These maps enable real-time interpretation of viral antigenic evolution
Method should be extensible to many other viral entry proteins
- System provides safe way to do mutational studies using pseudotyped lentivirions
- We have completely or partially implemented this system for:
- SARS-CoV-2 spike
- HIV Env
- H5 influenza hemagglutinin
- Nipah virus RBP
- Lassa virus GP
- Rabies virus GP
- It could likely be applied to many other viruses, including:
- RSV
- human parainfluenza virus
- Ebola virus
- other human and potential pandemic coronaviruses and influenza viruses