Evolutionary potential of the SARS-CoV-2 receptor binding domain (RBD)
Jesse Bloom
Fred Hutch Cancer Research Center / HHMI
Slides at http://slides.com/jbloom/salzman-2020
SARS-CoV-2 virions are decorated by spike proteins that mediate viral entry
virion image from https://phil.cdc.gov/Details.aspx?pid=23312

A key portion of the spike is the receptor-binding domain (RBD)

The RBD binds ACE2 receptor on cells
Neutralizing antibodies often bind RBD & block interaction with ACE2

Mapping how mutations affect RBD is crucial for understanding virus's evolution

-Asn-Ile-Thr-Asn-Leu-
We are accustomed to mapping sequence to structure

RBD phenotypes
- protein folding
- affinity for ACE2
- binding by antibodies
Structure still matters implicitly, as it largely determines mapping from mutation to phenotype.
-Asn-Ile-Thr-Asn-Leu-
-Asn-Ile-Thr-Glu-Leu-
-Asn-Lys-Thr-Asn-Leu-
We will map change in sequence to change in biochemical phenotype
Outline
- Map how RBD mutations affect folding and ACE2 binding
Outline
- Map how RBD mutations affect folding and ACE2 binding
- Map how RBD mutations affect antibody binding
Outline
- Map how RBD mutations affect folding and ACE2 binding
- Map how RBD mutations affect antibody binding
- Use maps to predict if / how virus can escape antibodies
Outline
- Map how RBD mutations affect folding and ACE2 binding
- Map how RBD mutations affect antibody binding
- Use maps to predict if / how virus can escape antibodies

Outline
- Map how RBD mutations affect folding and ACE2 binding
- Map how RBD mutations affect antibody binding
- Use maps to predict if / how virus can escape antibodies
There are a lot of possible mutations to the SARS-CoV-2 RBD
(201 sites) X (19 amino-acid mutations per site) = 3,819 mutations
We use yeast display to enable high-throughput experiments

RBD
fluorescent ACE2
yeast
fluorescent tag on RBD
Library of yeast, each expressing different RBD mutant with identifying 16 nt barcode

Click here for details on how library is made.
For single yeast variant, it's easy to use flow cytometry to make an ACE2 binding curve
For single yeast variant, it's easy to use flow cytometry to make an ACE2 binding curve

RBD
fluorescent ACE2
yeast
(1) label yeast at different ACE2 concentrations
For single yeast variant, it's easy to use flow cytometry to make an ACE2 binding curve

RBD
fluorescent ACE2
yeast
fluorescent signal
(1) label yeast at different ACE2 concentrations
(2) measure FACS signal at each concentration
[ACE2]
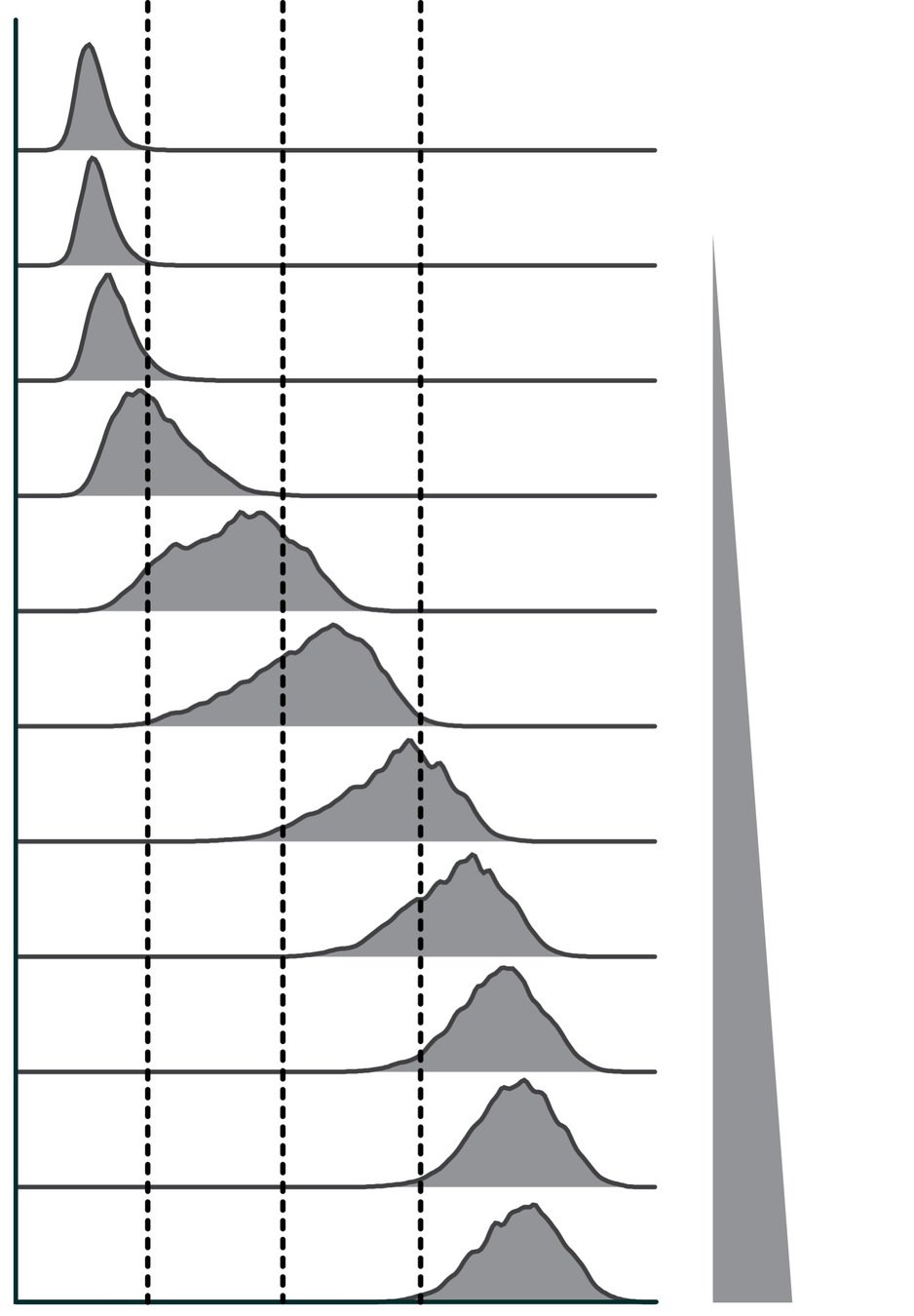
For single yeast variant, it's easy to use flow cytometry to make an ACE2 binding curve

RBD
fluorescent ACE2
yeast
fluorescent signal
(1) label yeast at different ACE2 concentrations
(2) measure FACS signal at each concentration
(3) binding curve of FACS signal versus [ACE2]
[ACE2]

Use deep sequencing to measure binding by all mutants at once
fluorescent signal
[ACE2]



(1) label yeast at different ACE2 concentrations
(2) sort into FACS bins
Deep sequence to assign RBD mutants to bins and infer dissociation curves
(3)
This strategy is TiteSeq, an elaboration of deep mutational scanning.
We similarly measure expression of folded protein for each mutant using tag on RBD

yeast
fluorescent tag on RBD
Phenotypic maps of how mutations affect RBD folding & ACE2 binding
Outline
- Map how RBD mutations affect folding and ACE2 binding
- Map how RBD mutations affect antibody binding
- Use maps to predict if / how virus can escape antibodies
We map escape mutations by sorting for RBD variants that don't bind antibody
We map escape mutations by sorting for RBD variants that don't bind antibody

We map escape mutations by sorting for RBD variants that don't bind antibody

We map escape mutations by sorting for RBD variants that don't bind antibody

We map escape mutations by sorting for RBD variants that don't bind antibody

In maps, tall letters indicate strong escape mutations
Escape map for one antibody (COV2-2499)
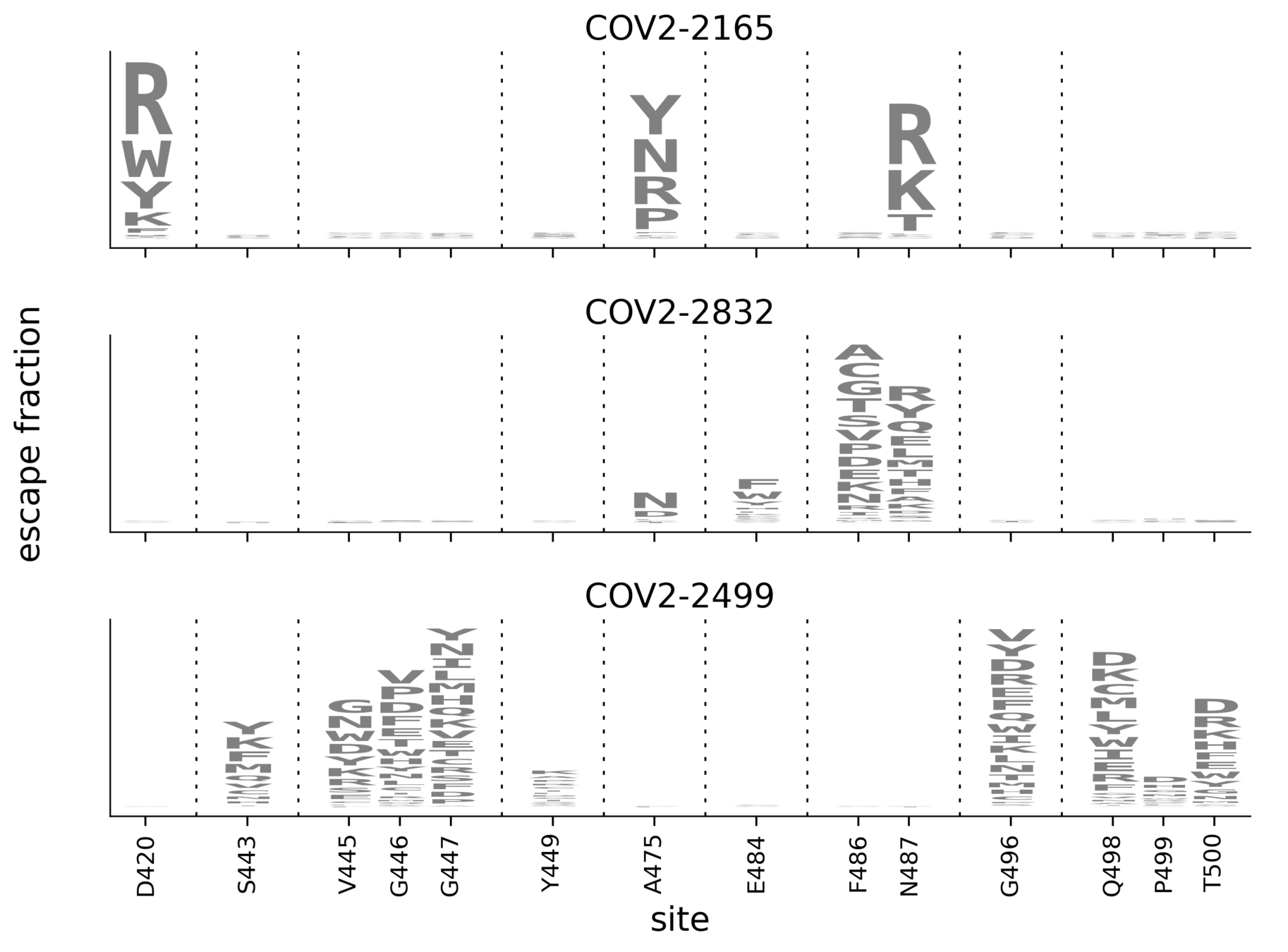
Interactive structural view of escape map (click here)
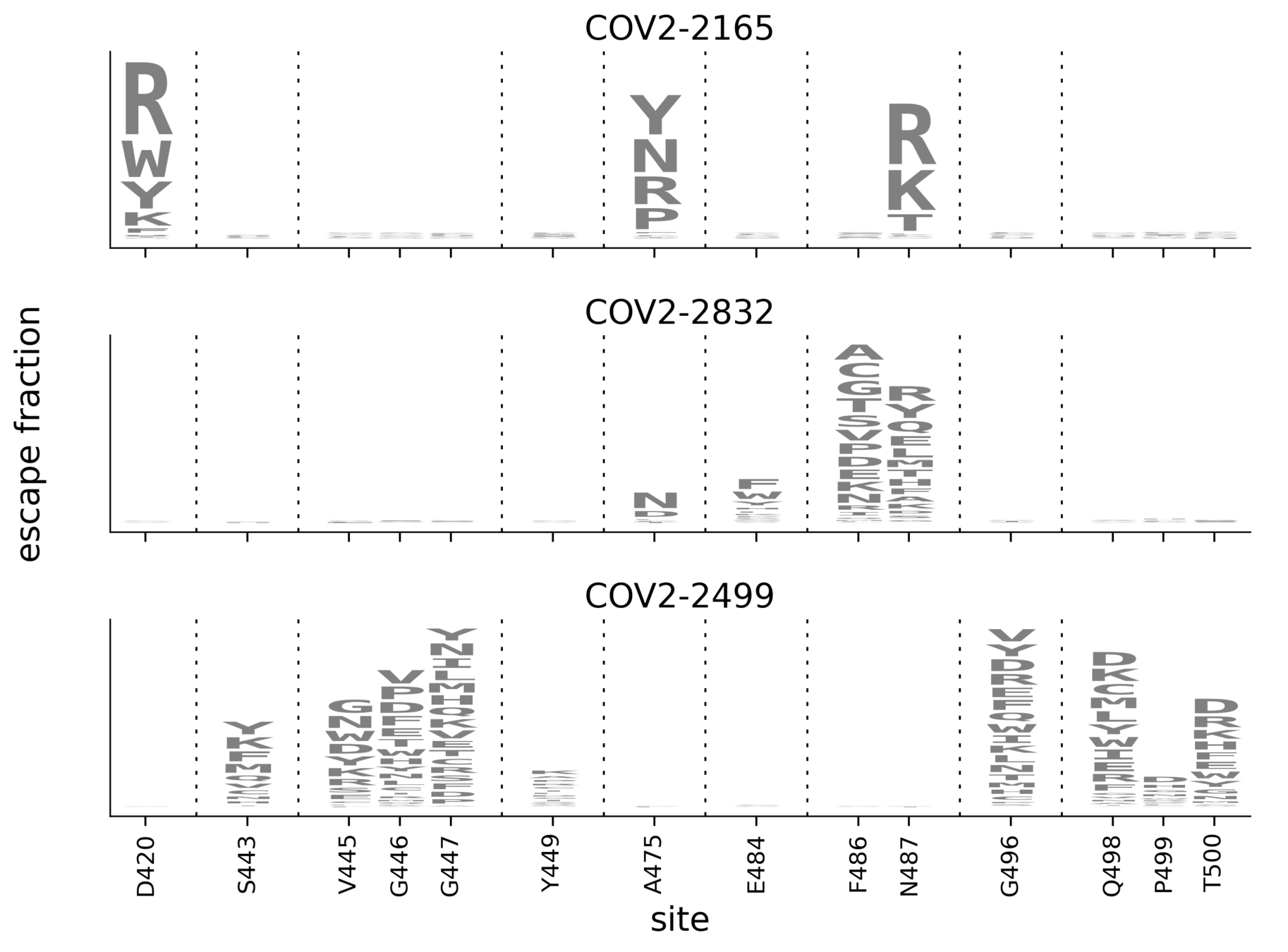
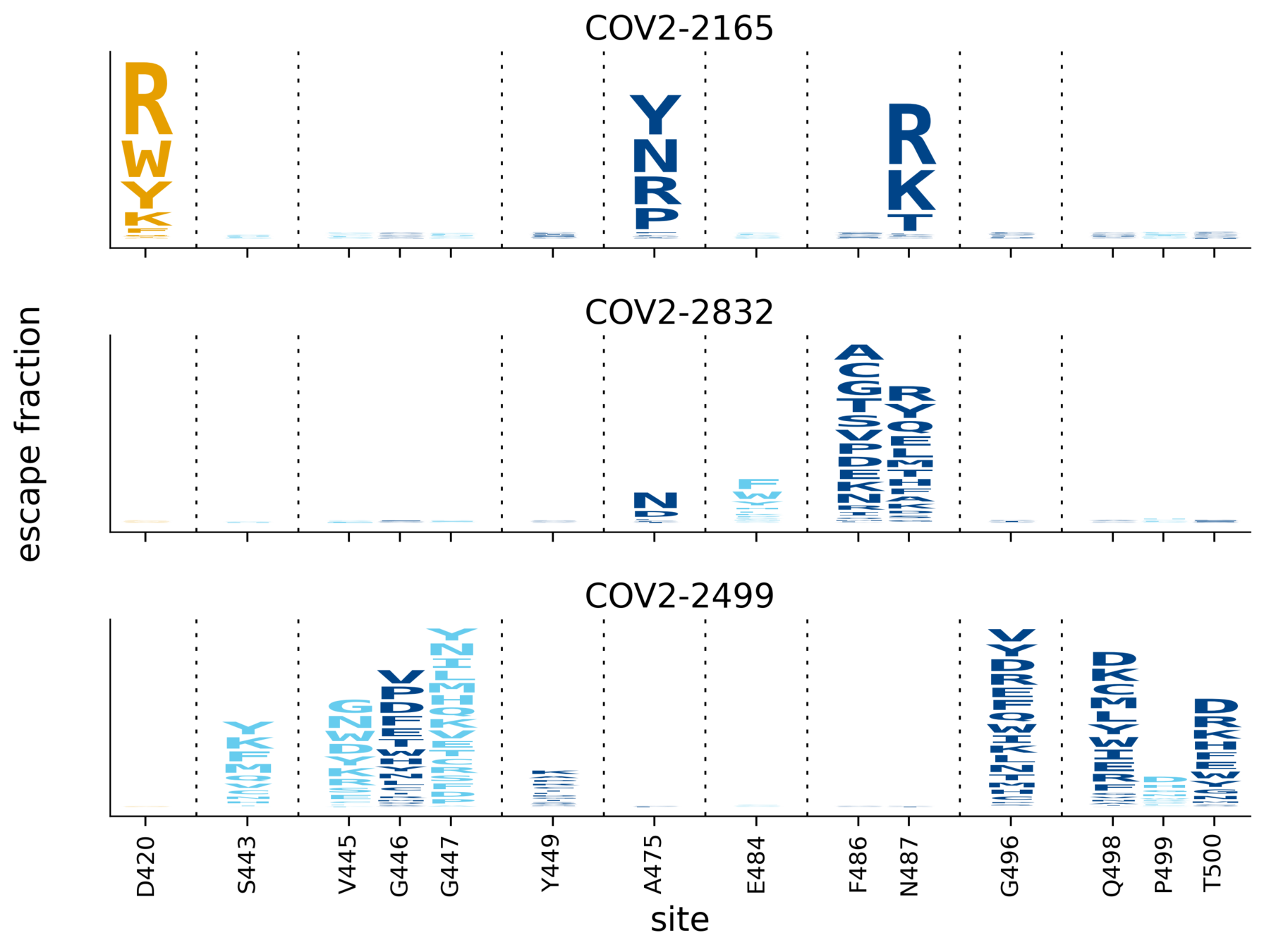
Maps easier to interpret if we color sites by region of RBD
core RBD
receptor-binding motif
ACE2-contact residue
core RBD
receptor-binding motif
ACE2-contact residue
Escape maps for three antibodies
Subtle differences between COV2-2165 & COV2-2832 (e.g., sites 486, 487) validate in neutralization assays (see here).
core RBD
receptor-binding motif
ACE2-contact residue

We can organize antibodies in space of escape mutations
Outline
- Map how RBD mutations affect folding and ACE2 binding
- Map how RBD mutations affect antibody binding
- Use maps to predict if / how virus can escape antibodies
We grew spike-expressing virus in presence of antibody
Spike-expressing VSV (Case*, Rothlauf*, ..., Whelan. Cell Host & Microbe, 2020) in a real-time cell analysis assay (Gilchuk, ..., Crowe. Immunity, 2020).
See here for more details.
Antibody COV2-2499 selected escape mutations in 5 of 16 replicates
mutation
G446D
Q498R
count
3
2
mutation
G446D
Q498R
count
3
2
Escape map has many mutations, why were these two selected?
mutation
G446D
Q498R
count
3
2
Only some mutations accessible by single-nucleotide changes
mutation
G446D
Q498R
count
3
2
Many mutations deleterious for ACE2 binding, but ones selected in virus aren't


ACE2 binding
weak strong
Selected mutations are single-nt changes that escape but retain ACE2 binding
effect on escape
ACE2 affinity

effect on escape
ACE2 affinity
Phenotypic maps also explain mutations selected by other antibodies
Antibody COV2-2165 did not select any escape mutations in repeated trials
0 escape mutants in 56 attempts
Initially appear to be many escape mutations from this antibody
But only a few are accessible by single-nucleotide changes
Accessible mutations impair folding or ACE2 binding, explaining why virus can't escape


RBD expression
ACE2 binding

RBD expression /
ACE2 binding
weak strong
Phenotypic maps explain if / how virus escapes antibodies
Mapping escape from antibodies in current clinical use
Escape map for REGN10933, one of the two antibodies in Regeneron cocktail
Regeneron used viral selections to identify a handful of escape mutations
Red indicates escape mutations identified by Regeneron in Baum, ..., Kyratsous. Science (2020)
Our approach maps these mutations plus all others that escape REGN10933
Red indicates escape mutations identified by Regeneron in Baum, ..., Kyratsous. Science (2020)
Useful to have these complete maps for surveillance of emerging viral mutations

Conclusions
Conclusions
- We can create phenotypic maps of how all mutations to SARS-CoV-2 RBD affect folding, function, and antigenicity
Conclusions
- We can create phenotypic maps of how all mutations to SARS-CoV-2 RBD affect folding, function, and antigenicity
- These maps are useful for understanding evolution under antibody pressure
Conclusions
- We can create phenotypic maps of how all mutations to SARS-CoV-2 RBD affect folding, function, and antigenicity
- These maps are useful for understanding evolution under antibody pressure
- These maps are useful for viral surveillance



Tyler Starr
Allie Greaney
Sarah Hilton
Bloom lab (Fred Hutch)



James Crowe
Seth Zost
Pavlo Gilchuk
Crowe lab (Vanderbilt)



Tyler Starr
Allie Greaney
Sarah Hilton
Bloom lab (Fred Hutch)



James Crowe
Seth Zost
Pavlo Gilchuk
Crowe lab (Vanderbilt)



Tyler Starr
Allie Greaney
Sarah Hilton
Bloom lab (Fred Hutch)
Bloom lab (Fred Hutch)
- Kate Crawford
- Adam Dingens
- Andrea Loes
- Rachel Eguia
Crowe lab (Vanderbilt)
- Elad Binshtein
Whelan lab (Wash U)
- Sean Whelan
- Paul Rothlauf
- Zhuoming Liu
King lab (Univ Wash)
- Neil King
- Dan Ellis
Veesler lab (Univ Wash)
- David Veesler
- Alexandra Walls




These slides: