Interpreting the evolution
of SARS-CoV-2 in context of Omicron
Jesse Bloom
Fred Hutch Cancer Research Center / HHMI
Slides at https://slides.com/jbloom/sars-cov-2-usg
Disclosures
- I am on the scientific advisory boards of Flagship Labs 77 and Oncorus
- I consult for Moderna
- I am an inventor on Fred Hutch licensed patents related to deep mutational scanning of viral proteins
- My lab has unfunded research collaborations with Vir Biotechnology
Outline
- Principles of viral antigenic evolution and emergence of Omicron
Outline
- Principles of viral antigenic evolution and emergence of Omicron
- Epistasis in ACE2 binding in Omicron
Outline
- Principles of viral antigenic evolution and emergence of Omicron
- Epistasis in ACE2 binding in Omicron
- Antibody escape in Omicron
Outline
- Principles of viral antigenic evolution and emergence of Omicron
- Epistasis in ACE2 binding in Omicron
- Antibody escape in Omicron
Only sometimes does virus evolution lead to changes in antigenic phenotype
- Measles virus: Does not evolve to escape immunity. People are infected at most once in their lives. A vaccine developed in the 1960s still works today.
- Influenza virus: Evolves to escape immunity. People are infected every ~5 years. The vaccine needs to be updated annually.
First let's look at another human coronavirus: CoV-229E causes common colds and has been circulating in humans since at least 1960s.
Reconstructing evolution of CoV-229E spike

We experimentally generated CoV-229E spikes at ~8 year intervals so we could study them in the lab:
- 1984
- 1992
- 2001
- 2008
- 2016
Note "ladder-like" shape of tree
Evolution of CoV-229E spike erodes neutralization by human antibody immunity
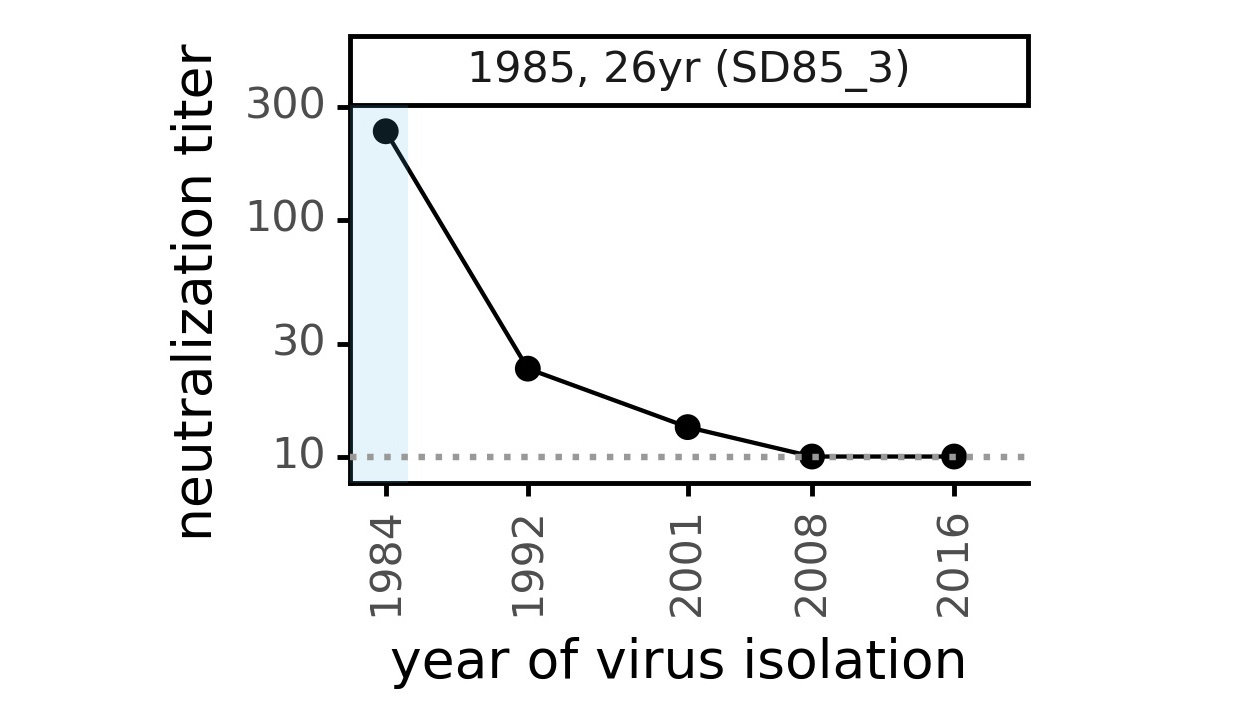
Serum collected in 1985 neutralizes virus with spike from 1984, but less effective against more recent viruses.
Viral evolution erodes antibody immunity of different people at different rates

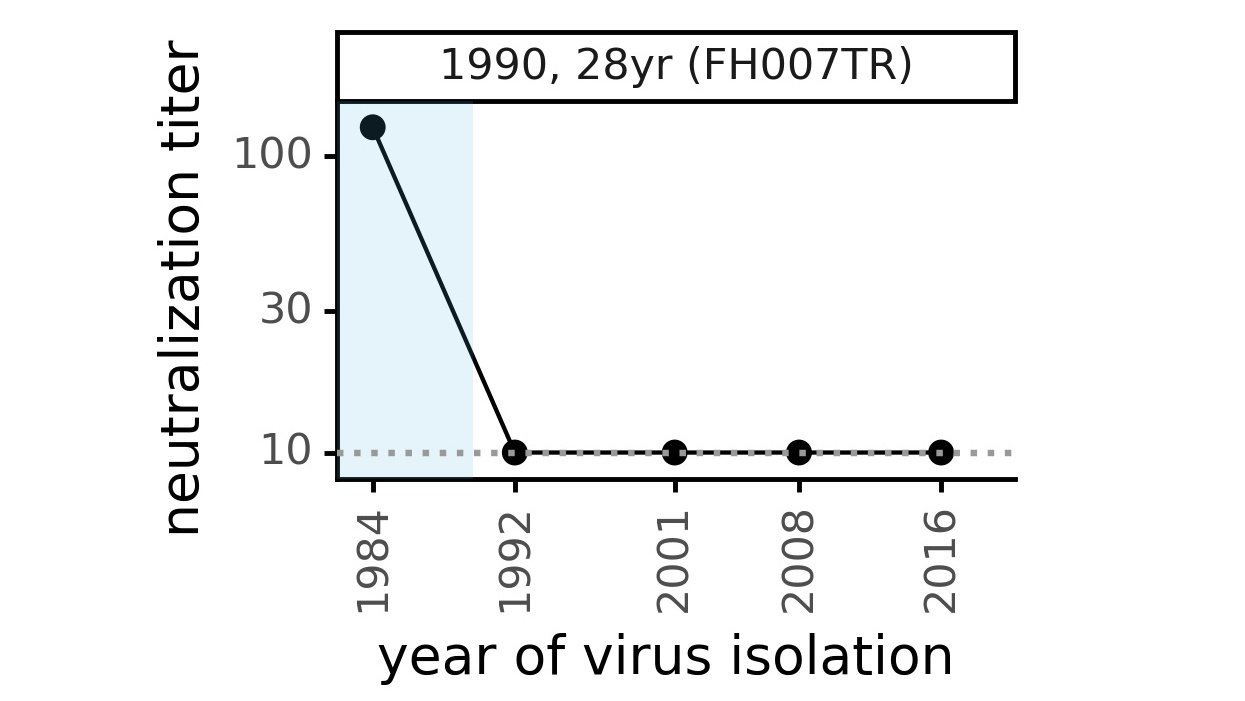
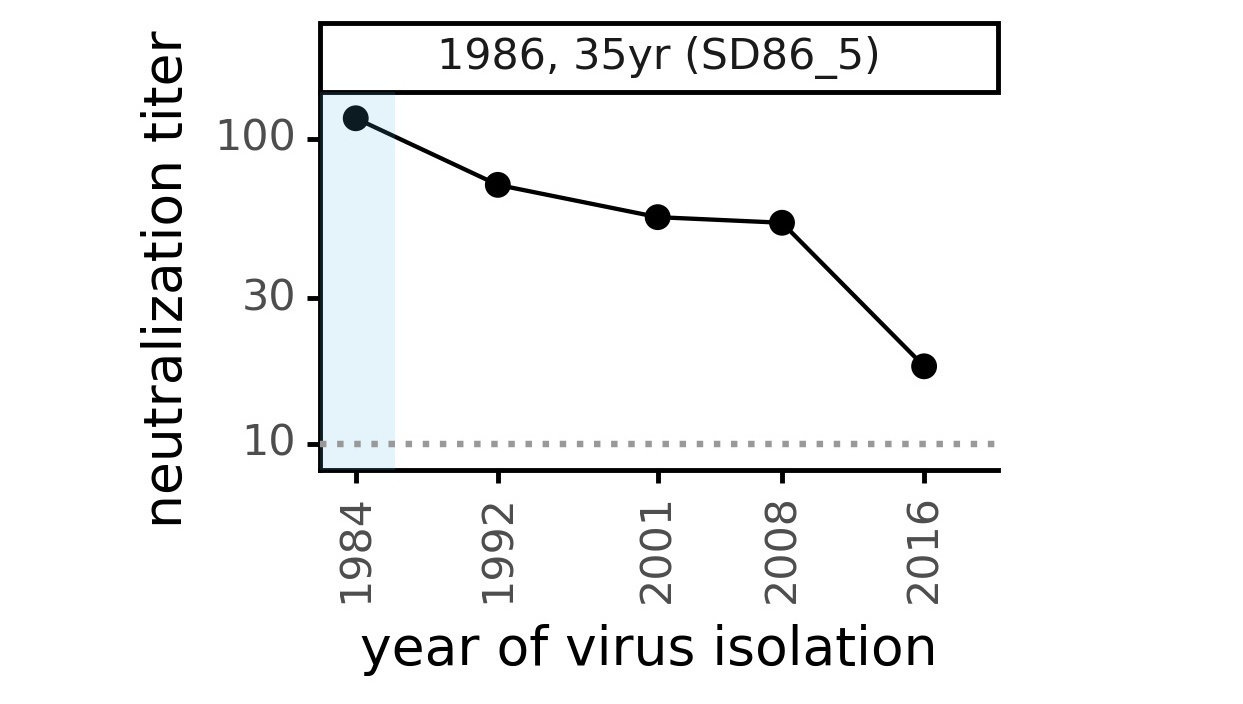
We are studying basis of these differences, as ideally vaccines would elicit more evolution-resistant sera as on the right.
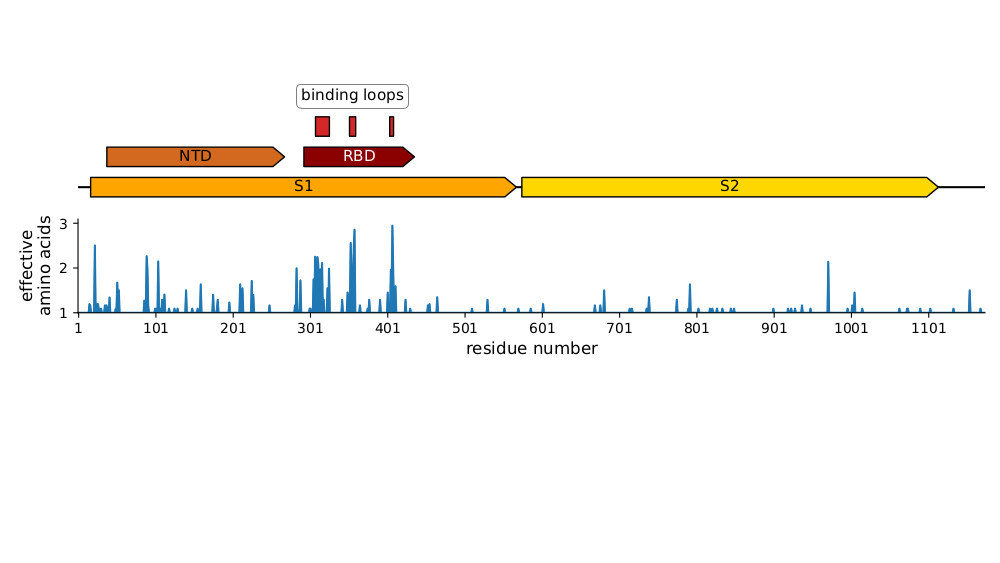
Most mutations in RBD & NTD
Plot of sequence variability across CoV-229E spike taken from Eguia, ..., Bloom, PLoS Pathogens (2021) . See also Wong, ..., Rini, Nature Communications (2017) and Li, ..., Rini, eLife (2019) for detailed structural studies of evolution in receptor-binding loops.
Similar to SARS-CoV-2 variants like Omicron except they also have cluster of mutations at S1-S2 furin cleavage site boundary.
Taken from Volz et al (2013), Kistler and Bedford (2021), and Kucharski (Twitter, 2022)
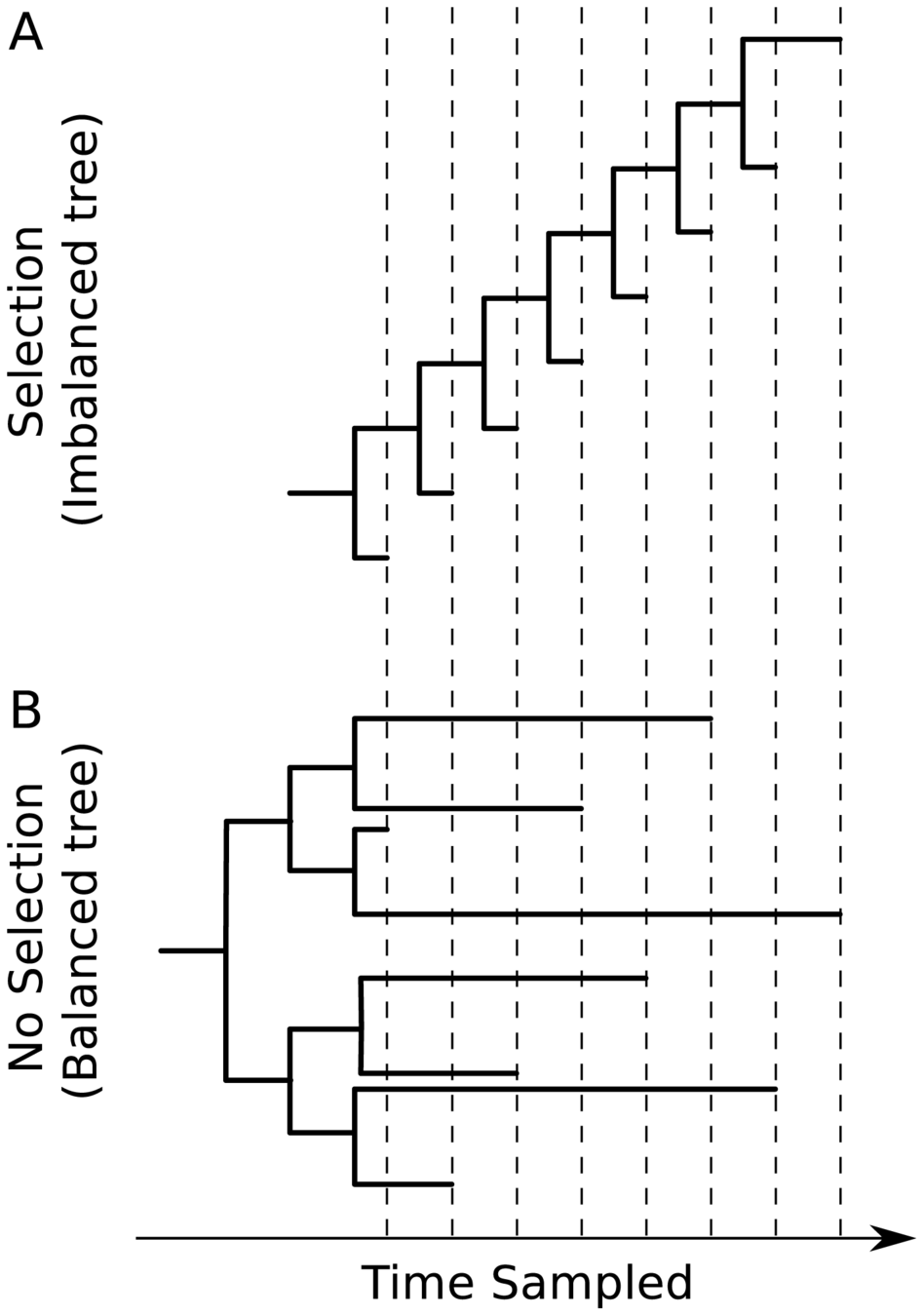
Human respiratory viruses that undergo rapid antigenic evolution tend to have a "ladder-like" tree because they evolve under "winner-take-all" paradigm where most successful variant displaces others. Viruses that evolve without antigenic selection (or have high diversity over geographic space) have more balanced trees.
CoV-229E tree looks like this
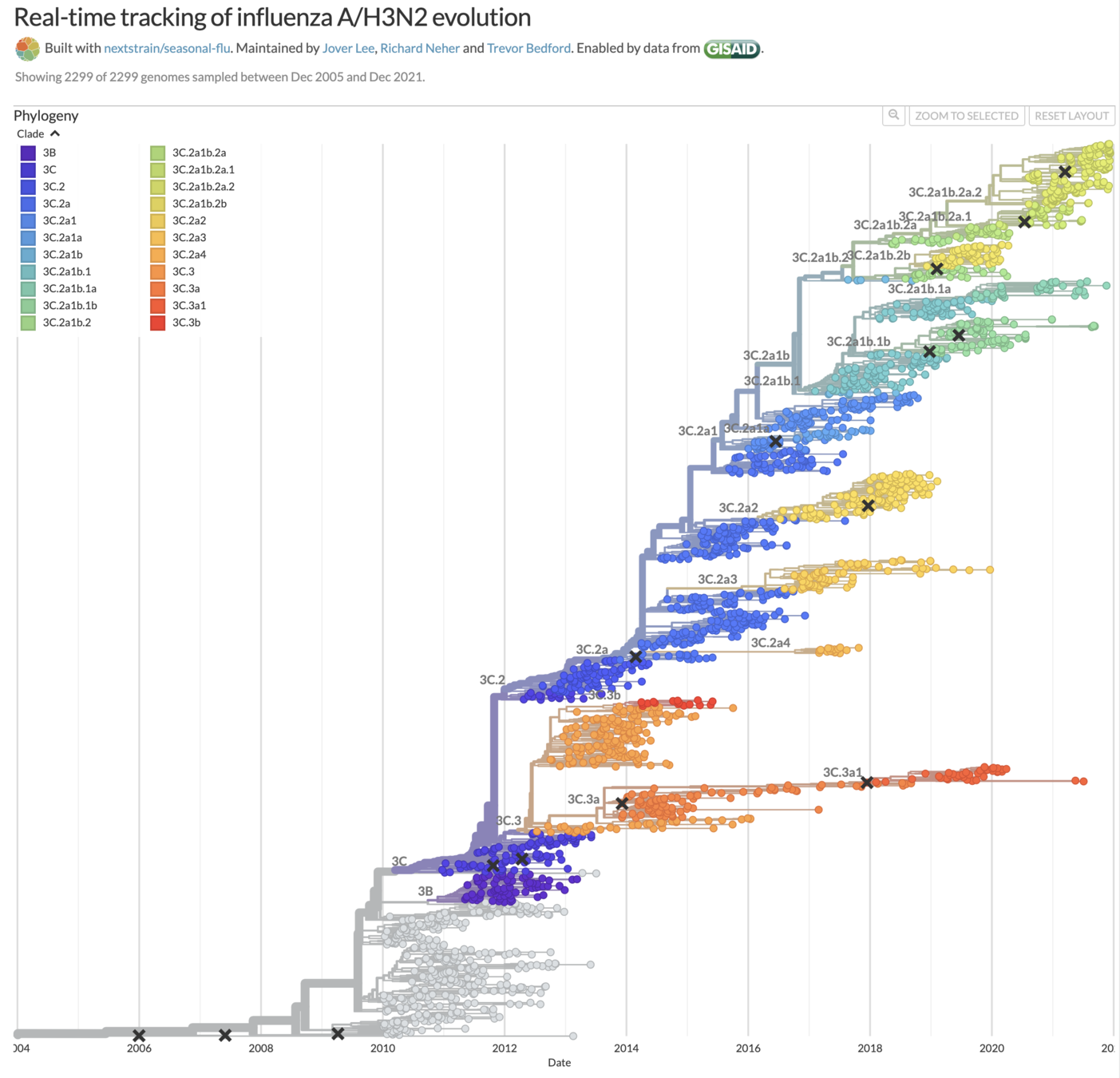
Human H3N2 influenza also tends to have ladder-like trees, and in years where it hasn't that's made vaccine strain selection difficult (vaccine strains marked with x).
Taken from Volz et al (2013), Kistler and Bedford (2021), and Kucharski (Twitter, 2022)
Human CoV-OC43 has split into two lineages, although each lineage has a ladder-like tree. Something similar has happened with influenza B.
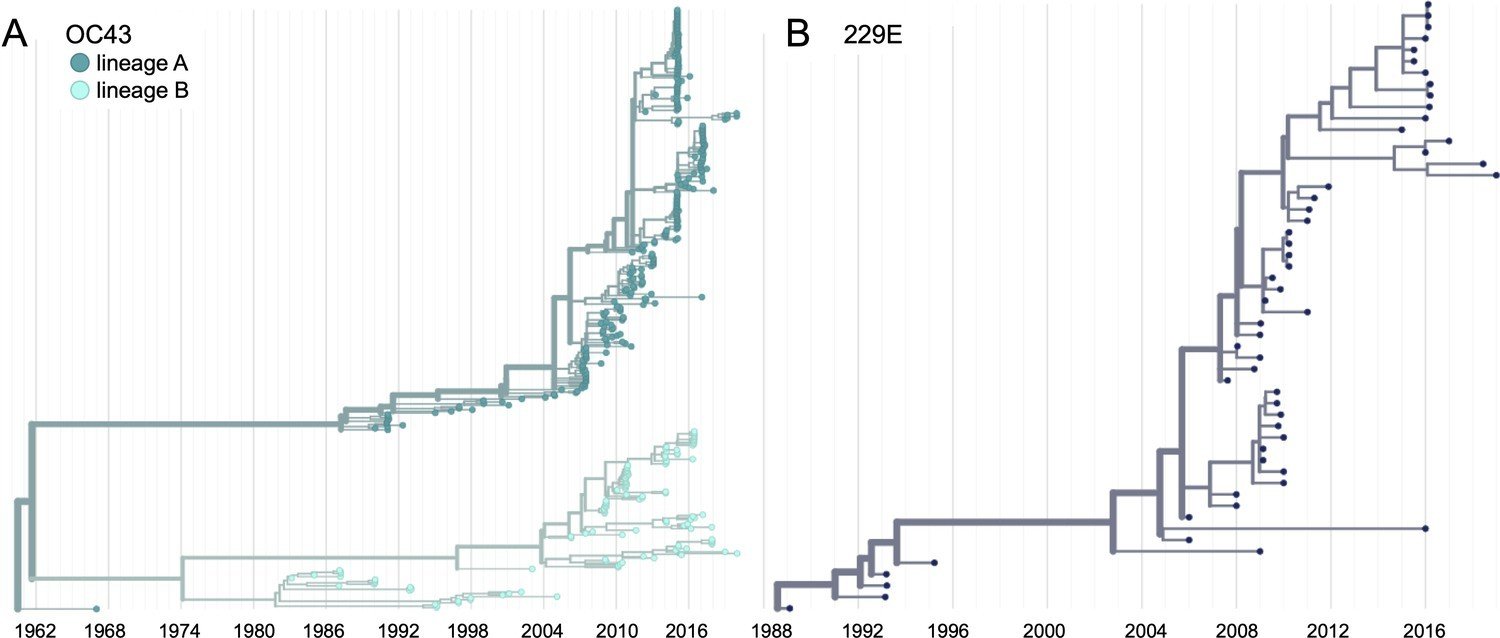
Taken from Volz et al (2013), Kistler and Bedford (2021), and Kucharski (Twitter, 2022)
Taken from Volz et al (2013), Kistler and Bedford (2021), and Kucharski (Twitter, 2022)
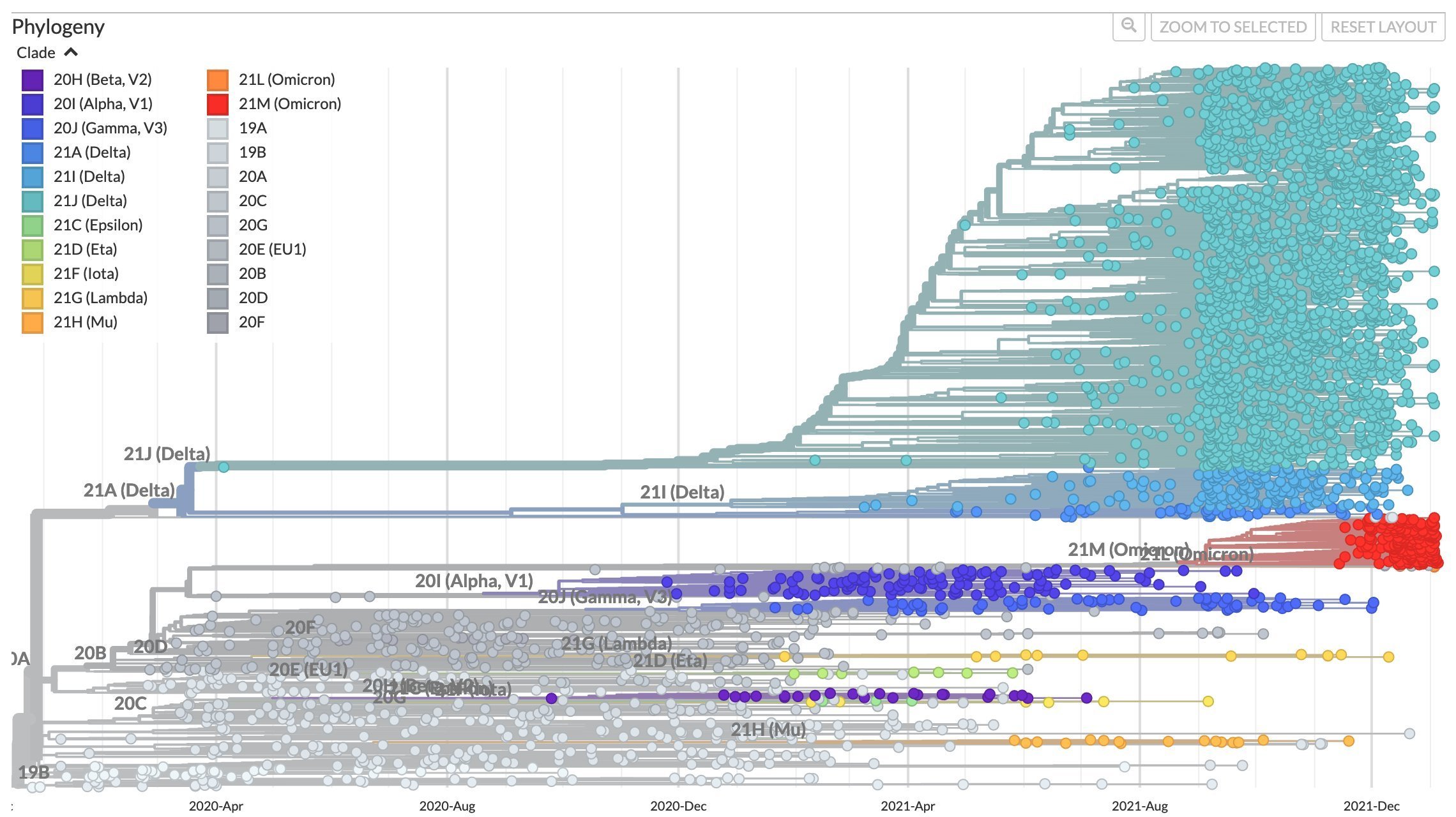
After early fixation of D614G, the SARS-CoV-2 tree has not been ladder-like
Take-aways on general principles
Other coronaviruses evolve antigenically with mutations in same regions as SARS-CoV-2 variants (exception is SARS-CoV-2 variants also have mutations is S1-S2 furin-cleavage site, probably because initial virus was poorly adapted to its furin-cleavage site).
Other antigenically evolving viruses have ladder-like phylogenetic trees, which is key to enabling vaccine updates as it provides some level of predictability (the next variant usually is descended from the last one).
So far SARS-CoV-2 has mostly defied ladder-like paradigm: Omicron not descended from Delta; Delta not descended from Alpha. However, it's still early and right now we are seeing combined effect of selection for antibody escape and increased transmissibility--presumably transmissibility will eventually plateau.
Outline
- Principles of viral antigenic evolution and emergence of Omicron
- Epistasis in ACE2 binding in Omicron
- Antibody escape in Omicron
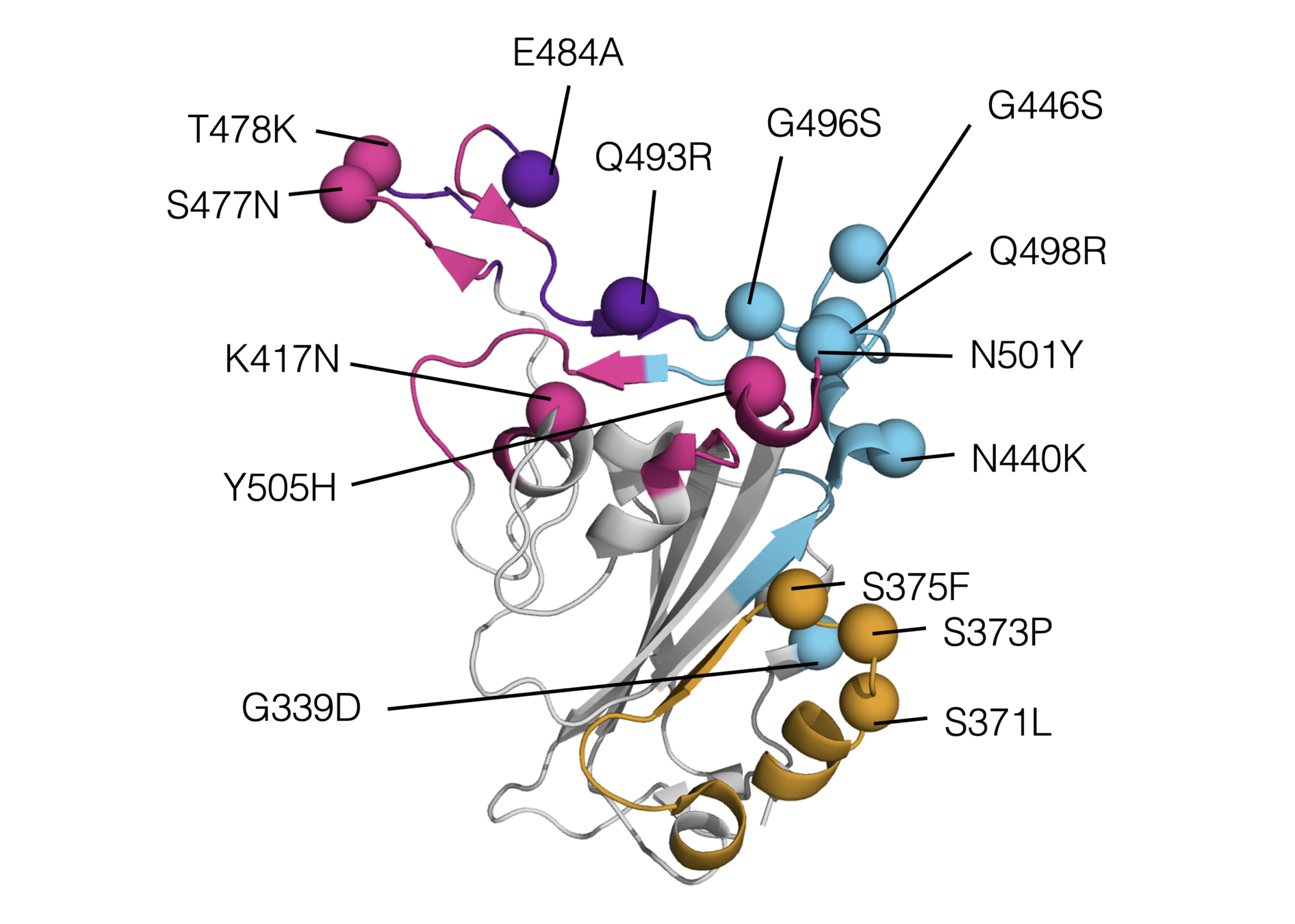
Omicron has a lot of mutations, especially in the receptor binding domain (RBD)
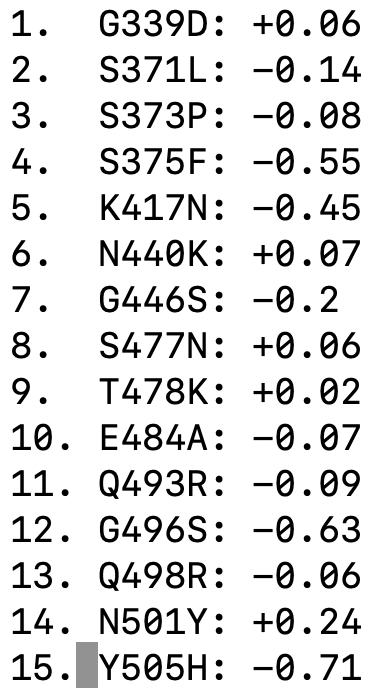
We have used deep mutational scanning to measure how all individual mutations in "original" (Wuhan-Hu-1) RBD affect ACE2 affinity: two-thirds of mutations in Omicron's RBD are individually deleterious for ACE2 affinity
The individual mutations in Omicron's RBD are predicted to have a net deleterious effect on ACE2 affinity
But Omicron actually binds to ACE2 ~2-fold better than Wuhan-Hu-1
There is favorable epistasis among mutations in Omicron RBD: they bind ACE2 better together than alone
Two studies have shown that Q498R and N501Y combine to lead to a much greater increase in affinity than either alone. See this Twitter thread for more evolutionary details.
It is also possible that there is epistasis among other combinations of mutations in Omicron, although this has not yet been experimentally elucidated.
Omicron represents first major example where the effects of combined RBD mutations could not be predicted fairly well from single mutants.
Outline
- Principles of viral antigenic evolution and emergence of Omicron
- Epistasis in ACE2 binding in Omicron
- Antibody escape in Omicron
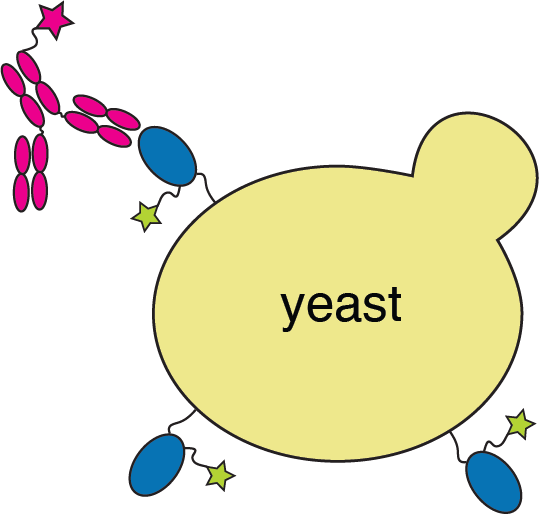
We map escape mutations by sorting for RBD variants that don't bind antibody
RBD
fluorescently labeled antibody
yeast
fluorescent tag on RBD
Escape map from a single antibody
Escape maps from lots of antibodies
See Seth Zost's thread for best summary of therapeutic antibodies and Omicron
Infection / vaccination elicit polyclonal antibodies that can bind many epitopes
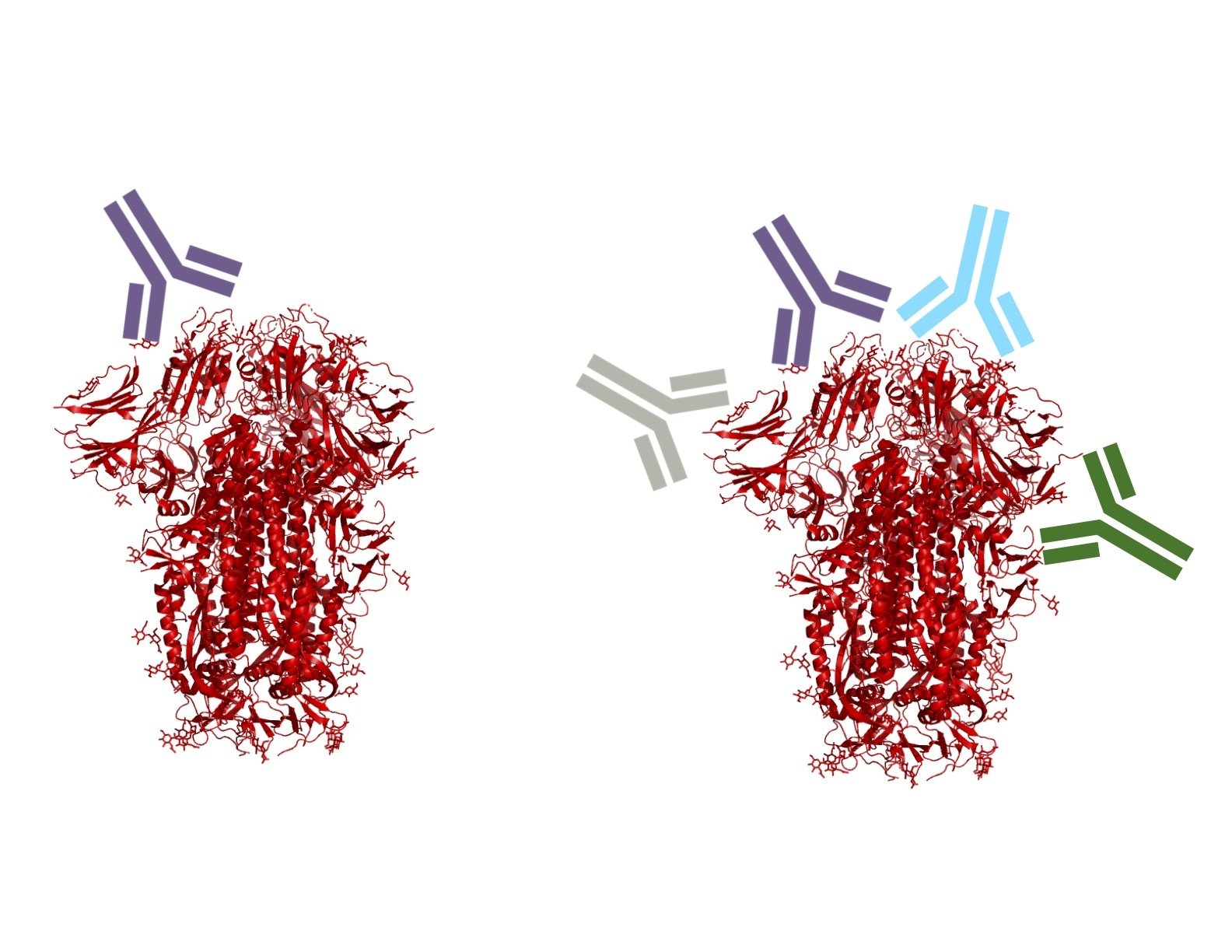
Monoclonal antibodies bind one epitope, so can usually be escaped by single mutation
Polyclonal antibodies can bind many epitopes, so often more resistant to escape
If polyclonal antibodies bind multiple epitopes, escape needs multiple mutations
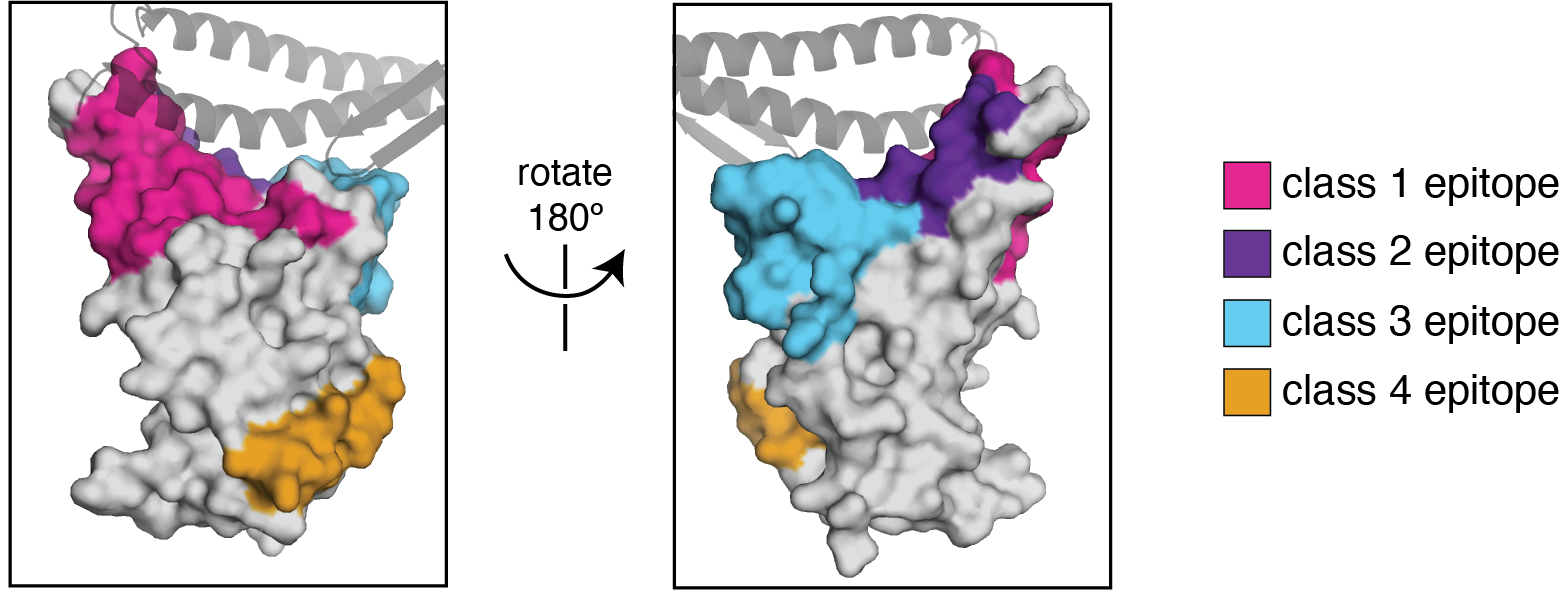
We use the antibody classification scheme from Barnes et al Nature (2020). The extension of these antibody classes to escape mapping is in Greaney et al (2021), and escape maps are available here. Class 1, 2, and 3 antibodies are often potently neutralizing, while class 4 antibodies are less neutralizing (see: Piccoli et al (2020), Dejnirattisai et al (2021), Liu et al (2020), Zost, et al (2020)).
We can visualize escape from different antibody classes
We can calculate how combinations of mutations affect polyclonal binding
Toy example of calculating escape from polyclonal mix:
https://jbloomlab.github.io/SARS2_RBD_Ab_escape_maps/mini-example-escape-calc/
Escape-calculator with deep mutational scanning data from many antibodies:
https://jbloomlab.github.io/SARS2_RBD_Ab_escape_maps/escape-calc/
Predictions correlate well with neutralization assays:
https://jbloomlab.github.io/RBD_escape_calculator_paper/neut_studies.html
See Greaney et al (2021) for paper on escape-calculator, and here for more examples on usage.
Omicron has extensive antigenic change
Omicron has a lot more escape than other variants.
Sites to watch for future antigenic mutations include 346.

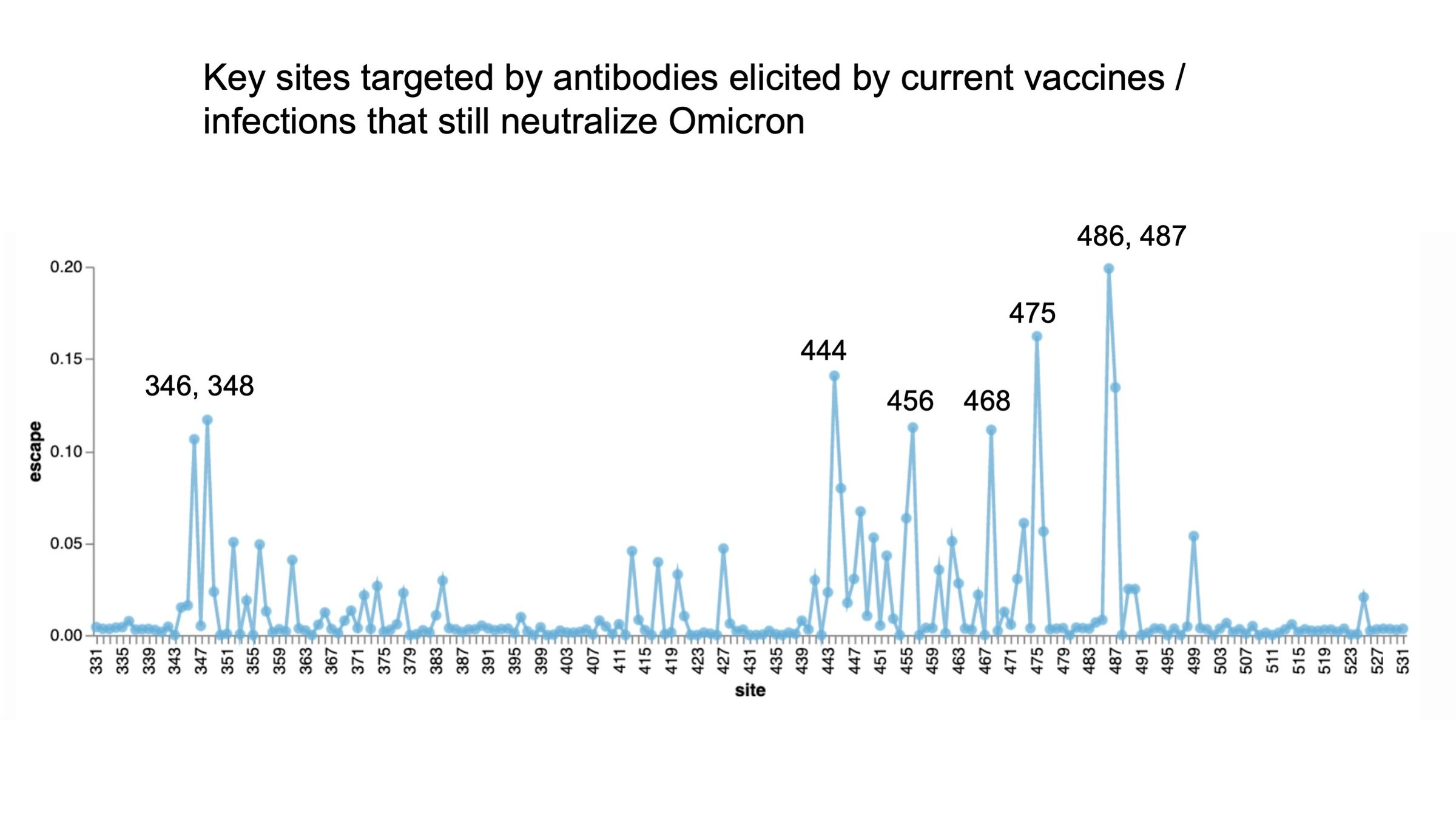
Mutations that escape antibodies elicited by SARS-CoV-2 that still neutralize Omicron represent sites of likely future change
See here for details on how to use interactive version of this plot at https://jbloomlab.github.io/SARS2_RBD_Ab_escape_maps/escape-calc/
Crowe lab (Vanderbilt)
Chu lab (Univ Wash)
Veesler lab (Univ Wash)
King lab (Univ Wash)
Li lab (Brigham & Women's)
Boeckh lab (Fred Hutch)
Alex Greninger (Univ Wash)
Nussenzweig lab (Rockefeller)
Bjorkman lab (Caltech)






Tyler Starr
Allie Greaney
Rachel Eguia
Bloom lab (Fred Hutch)


Sarah Hilton
Kate Crawford


Andrea Loes