Interpreting the evolution of SARS-CoV-2
Jesse Bloom
Fred Hutch Cancer Center / HHMI
These slides at https://slides.com/jbloom/sars2-uw-immuno
Disclosures
I am on the scientific advisory boards of Apriori Bio and Oncorus
I have consulted on topics related to viral evolution for Moderna and Merck
I am an inventor on Fred Hutch licensed patents related to deep mutational scanning of viral proteins
My lab has unfunded research collaborations with Vir Biotechnology
Outline
- Conceptual principles of viral antigenic evolution
Outline
- Conceptual principles of viral antigenic evolution
- SARS-CoV-2 receptor-binding domain (RBD)
Outline
- Conceptual principles of viral antigenic evolution
- SARS-CoV-2 receptor-binding domain (RBD)
- Interpreting evolution of the SARS-CoV-2 RBD
Outline
- Conceptual principles of viral antigenic evolution
- SARS-CoV-2 receptor-binding domain (RBD)
- Interpreting evolution of the SARS-CoV-2 RBD

The Faroe Islands


"Measles had not prevailed on the Faroes since 1781, then it broke out early in April 1846."
"Of the 7782 inhabitants, about 6000 were taken with measles."
"Of the many aged people still living in the Faroes who had measles in 1781, not one was attacked the second time."
Panum is describing immune memory, which provides lifelong protection from measles.
But we are repeatedly infected by some other viruses. Typical person infected by H3N2 influenza ~5 years. Why?
History offers natural experiment with influenza like Panum's study of measles
From 1918 to 1957, H1N1 influenza circulated in humans.
In 1957, H1N1 disappeared from humans due to new H2N2 pandemic.
In 1977, old H1N1 strain from ~1954 was accidentally re-released (probably during misguided vaccine trial) and caused a global pandemic.


"One boy from Hong Kong had a transient febrile illness from 15 to 18 January. On Sunday 22 January, three boys were in the college infirmary… 512 boys (67%) spent between three and seven days away from class."

"Of about 130 adults who had some contact with the boys, only one, a house matron, developed similar symptoms."
Influenza infection elicits multi-decade immunity similar to measles - but only if virus is evolutionarily frozen
Only some human RNA respiratory viruses evolve to escape immunity
Why some viruses evolve to escape immunity while others don't is a deep question outside scope of this talk. See here for some possible explanations.
Rate of viral antigenic evolution
Measles
Influenza
Rate of viral antigenic evolution
Measles
Where do human coronaviruses fall on this spectrum?
Coronaviruses
???
???
Influenza
Rate of viral antigenic evolution
Measles
Coronaviruses
???

???
Initially, people speculated SARS-CoV-2 would evolve slowly. But we weren't so sure...
Influenza
We decided to study another human coronavirus: CoV-229E causes common colds and has been circulating in humans for a long time.
Reconstructing evolution of CoV-229E spike

We experimentally generated CoV-229E spikes at ~8 year intervals so we could study them in the lab:
- 1984
- 1992
- 2001
- 2008
- 2016
Note "ladder-like" shape of tree
Evolution of CoV-229E spike erodes neutralization by human antibody immunity
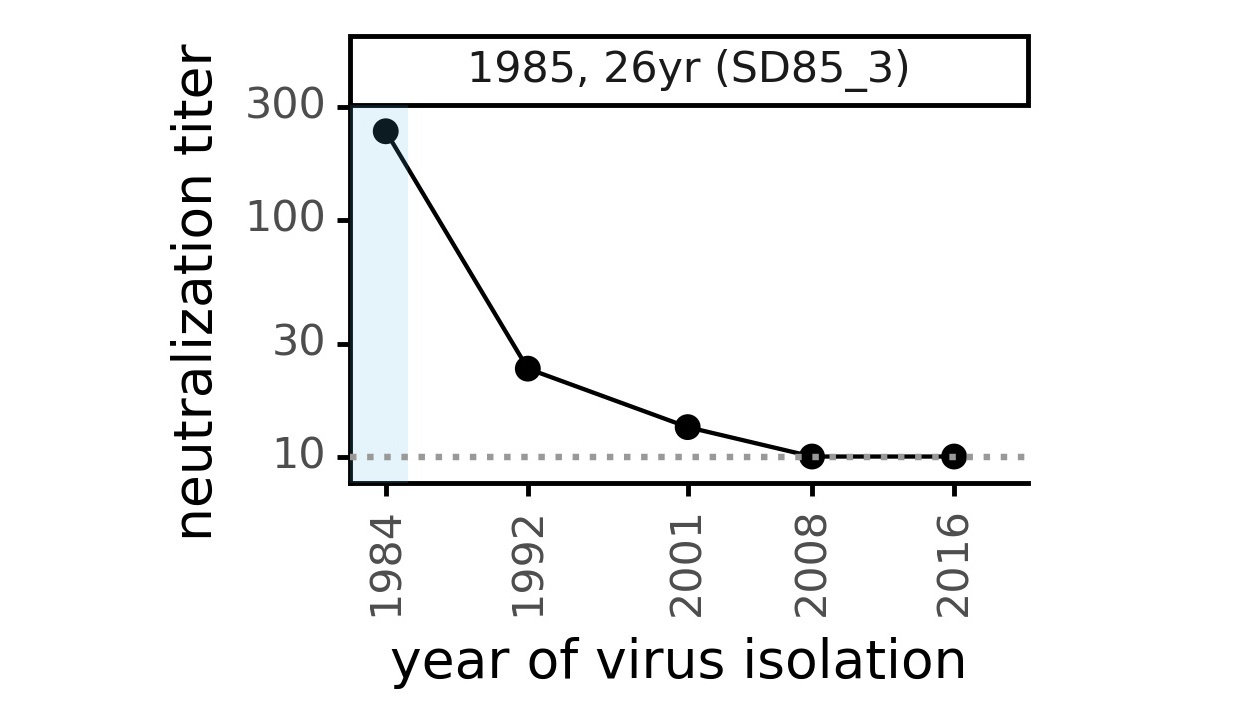
Serum collected in 1985 neutralizes virus with spike from 1984, but less effective against more recent viruses.
Viral evolution erodes antibody immunity of different people at different rates

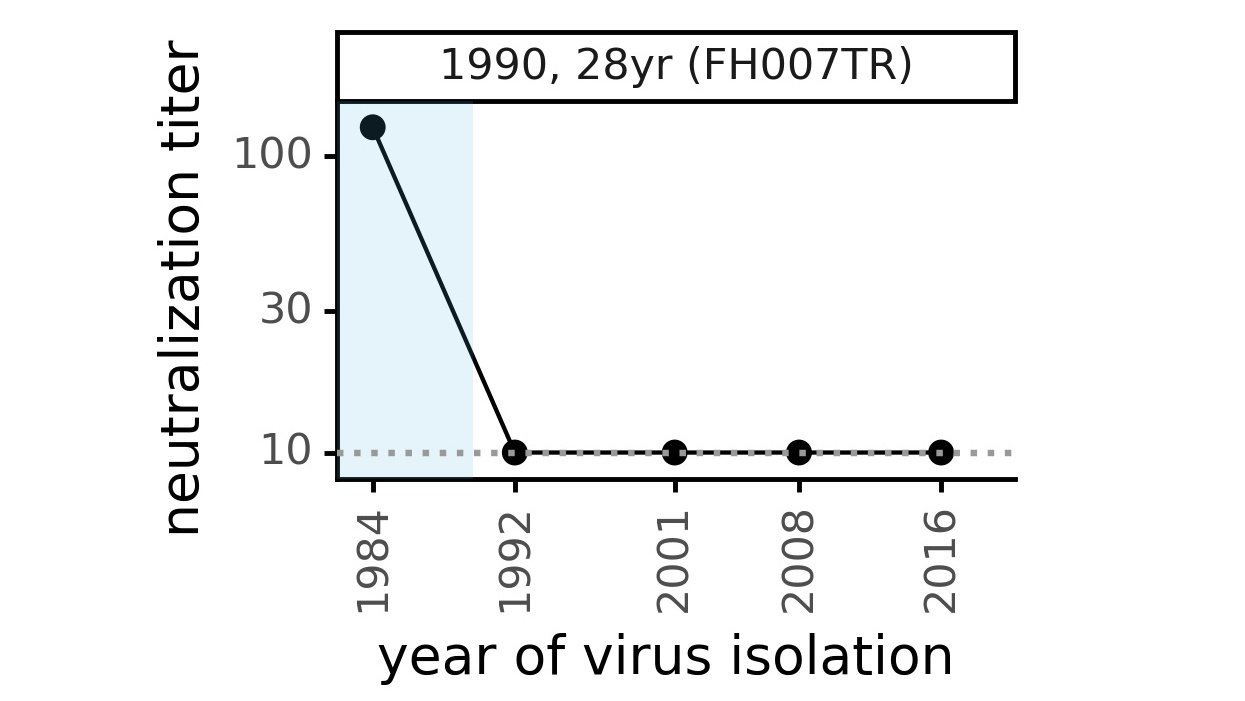
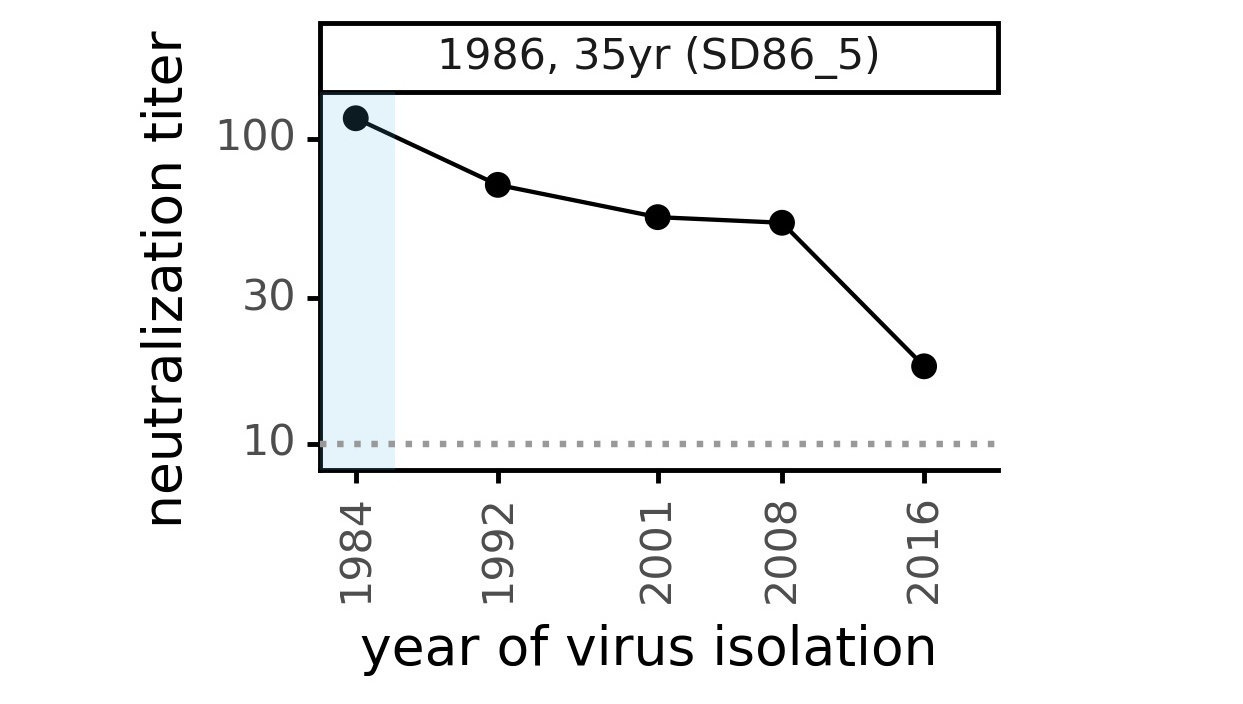
We are studying basis of these differences, as ideally vaccines would elicit more evolution-resistant sera as on the right.
Rate of viral antigenic evolution
Measles
CoV-229E
CoV-OC43
SARS-CoV-2
Most human coronaviruses undergo rapid antigenic evolution
Influenza
Phylogenetic tree shape & vaccine strategy
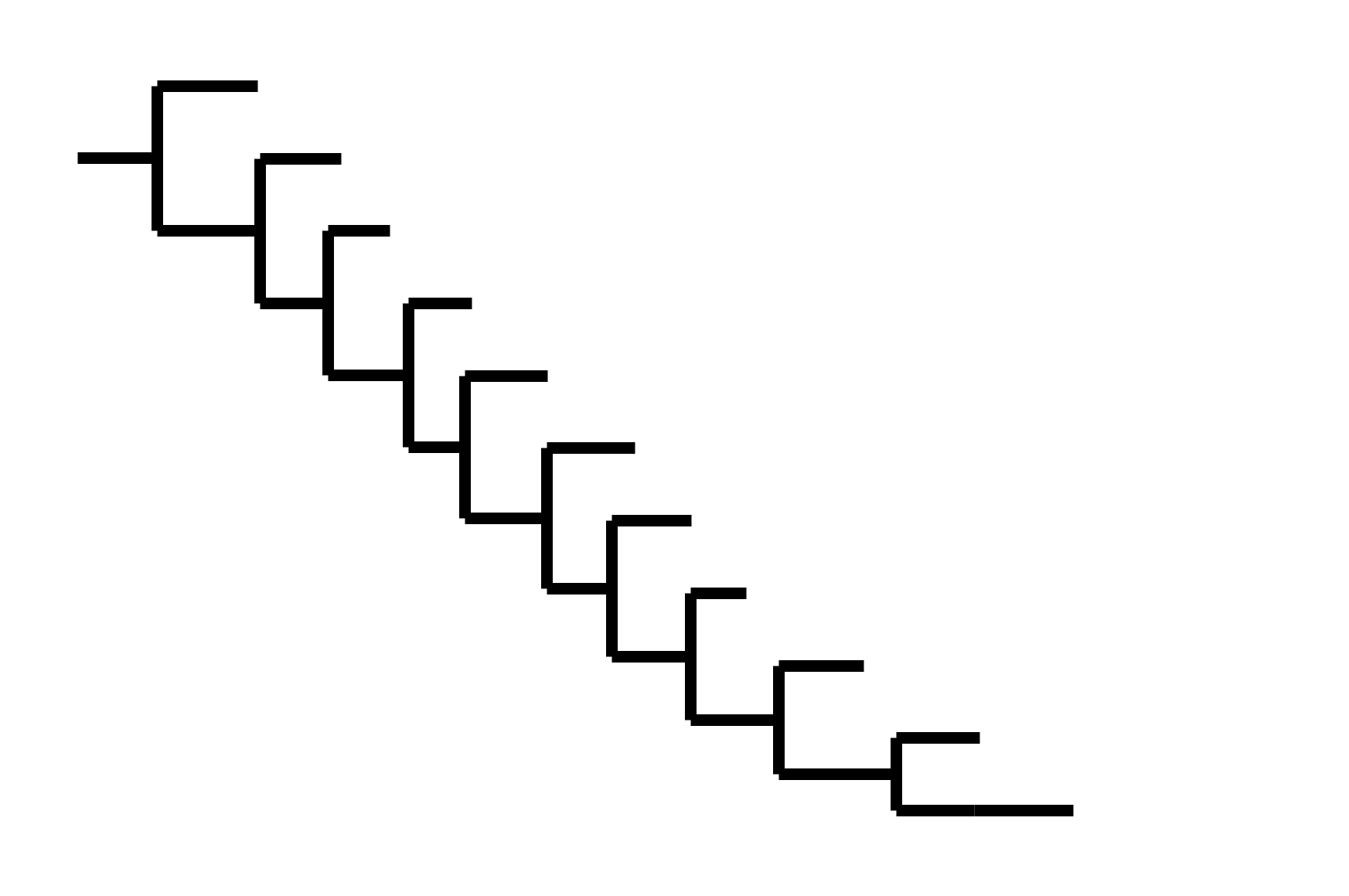
CoV-229E has ladder-like tree:
- new variants displace old ones
- new variants descend from recent successful ones
Human influenza A evolves this way too. It's theoretically possible to pick single well-matched vaccine strain.
Phylogenetic tree shape & vaccine strategy
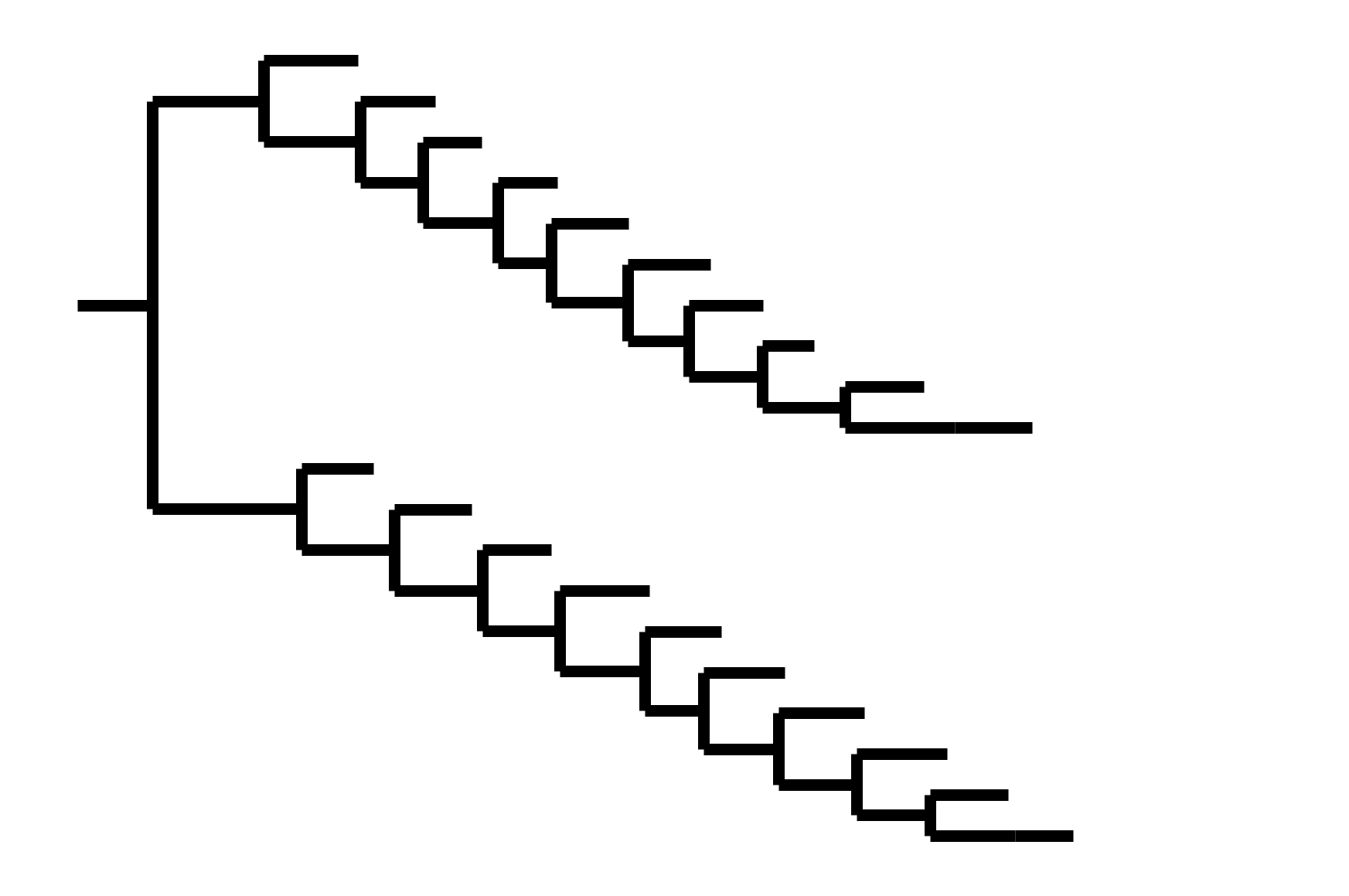

CoV-229E has ladder-like tree:
- new variants displace old ones
- new variants descend from recent successful ones
Human influenza A evolves this way too. It's theoretically possible to pick single well-matched vaccine strain.
CoV-OC43 split into two ladder-like lineages. Influenza B evolves this way too. It's theoretically possible to pick well-matched bivalent vaccine.
Phylogenetic tree shape & vaccine strategy

CoV-229E has ladder-like tree:
- new variants displace old ones
- new variants descend from recent successful ones
Human influenza A evolves this way too. It's theoretically possible to pick single well-matched vaccine strain.

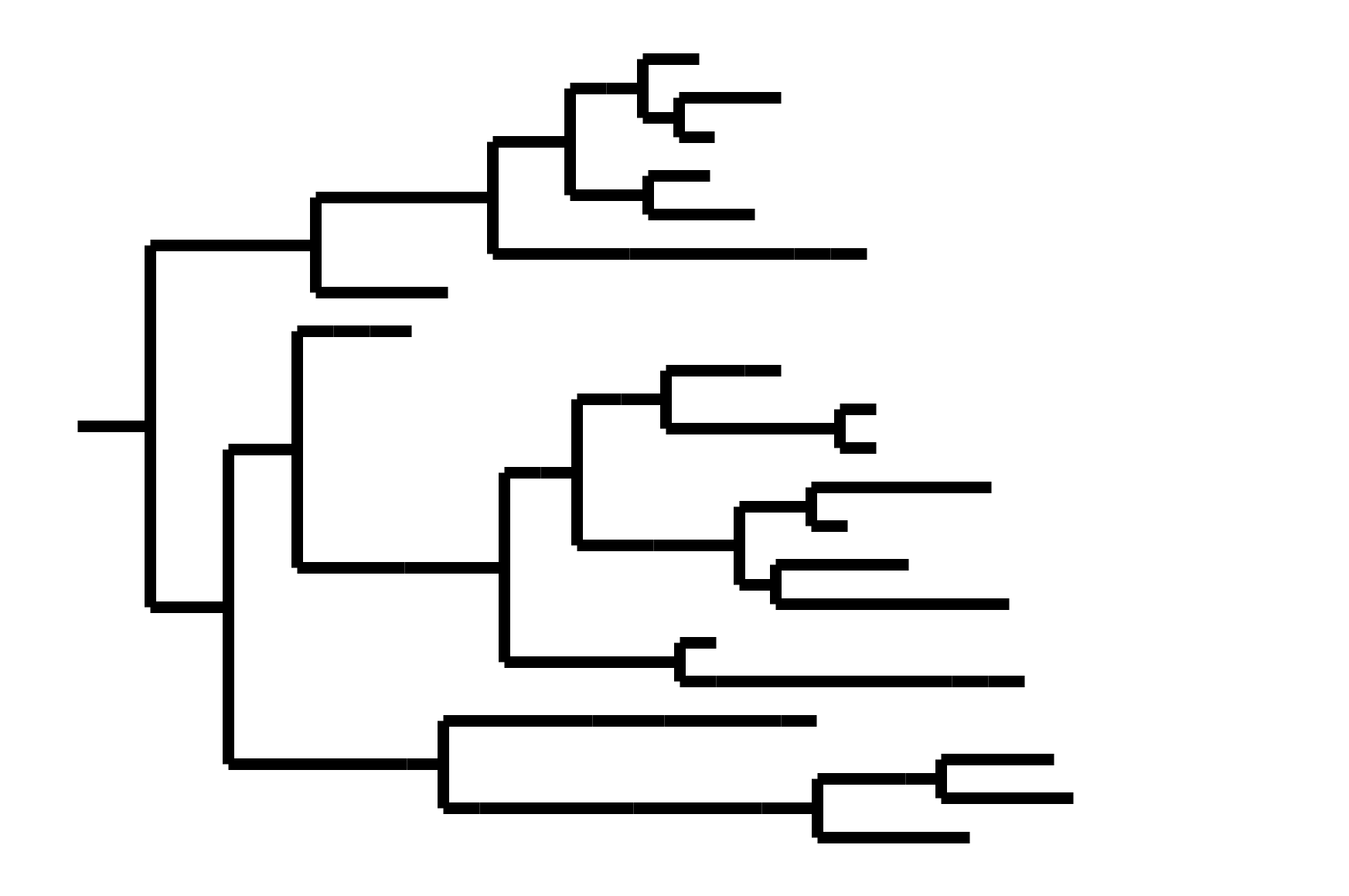
CoV-OC43 split into two ladder-like lineages. Influenza B evolves this way too. It's theoretically possible to pick well-matched bivalent vaccine.
In non-ladder-like tree, next variant not descended from recent successful one. Makes picking vaccine strains difficult.
So far, SARS-CoV-2 not entirely ladder-like
Omicron emerged on a branch with excess amino-acid mutations only in spike

For each Nextstrain clade, number of mutations plotted versus date of clade origin, with trend line fit only to non-Omicron clades. Omicron clades are circled in red.
Data extracted from Neher (2022) and re-plotted.
This pattern is consistent with antibody selection in chronic human infection

See here for more details on why Omicron probably evolved in a chronic human infection.
However, the first 2.5 years of SARS-CoV-2 evolution might be somewhat non-typical
A substantial amount of the evolution so far has been driven by increases in inherent transmissibility.
However, we are increasingly seen evolution driven by immune escape.
Human endemic viruses probably eventually plateau inherent transmissibility and evolve mostly to escape immunity. So the increased-transmissibility aspects of SARS-CoV-2 evolution may be transient features of the first few years.
Outline
- Conceptual principles of viral antigenic evolution
- SARS-CoV-2 receptor-binding domain (RBD)
- Interpreting evolution of the SARS-CoV-2 RBD
Strongest evolutionary selection is in RBD
Sites of evolutionary change in the spike of CoV-229E over the last four decades
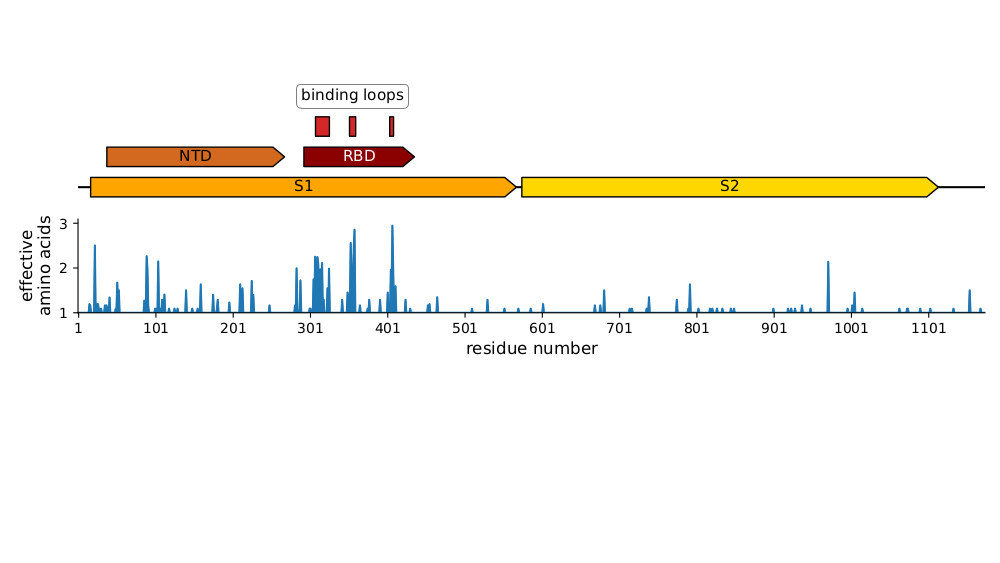
Strongest evolutionary selection is in RBD
Sites of evolutionary change in the spike of CoV-229E over the last four decades

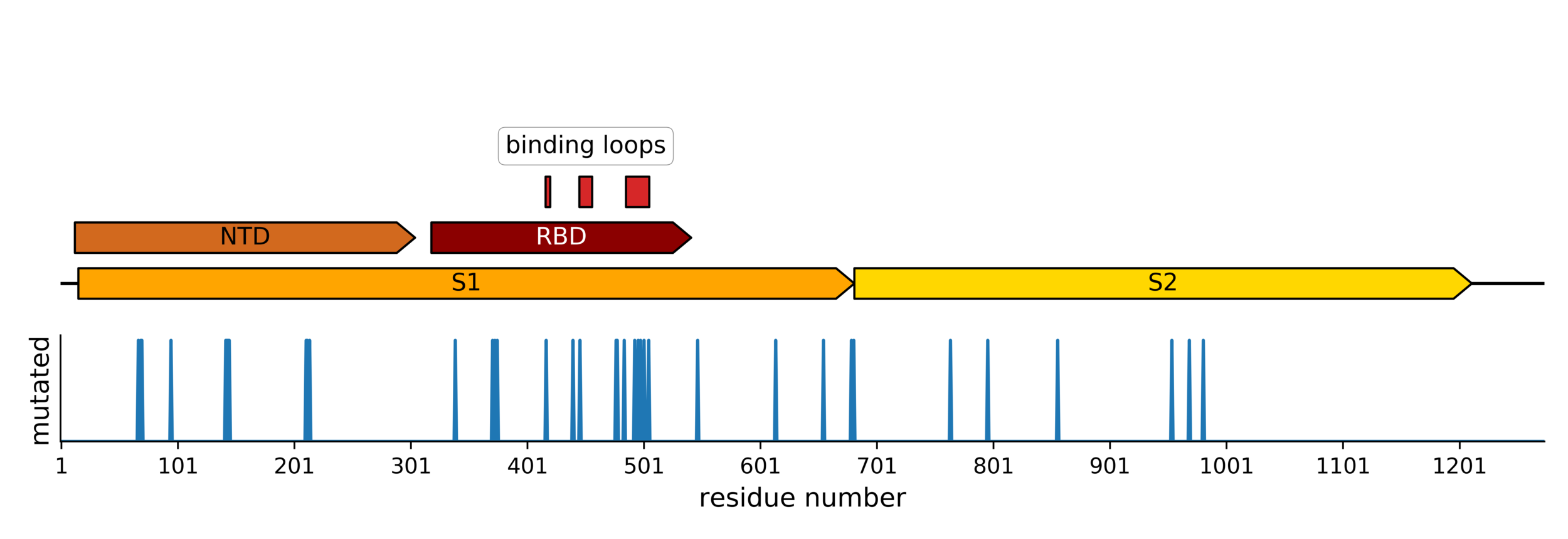
Sites of mutations in SARS-CoV-2 Omicron (BA.1) spike relative to Wuhan-Hu-1
Main difference is SARS-CoV-2 also fixing transmissibility-enhancing spike mutations that affect proteolytic processing and stabilize defects cause by furin-cleavage site
Majority of neutralizing antibody response in vaccinated/infected humans targets RBD
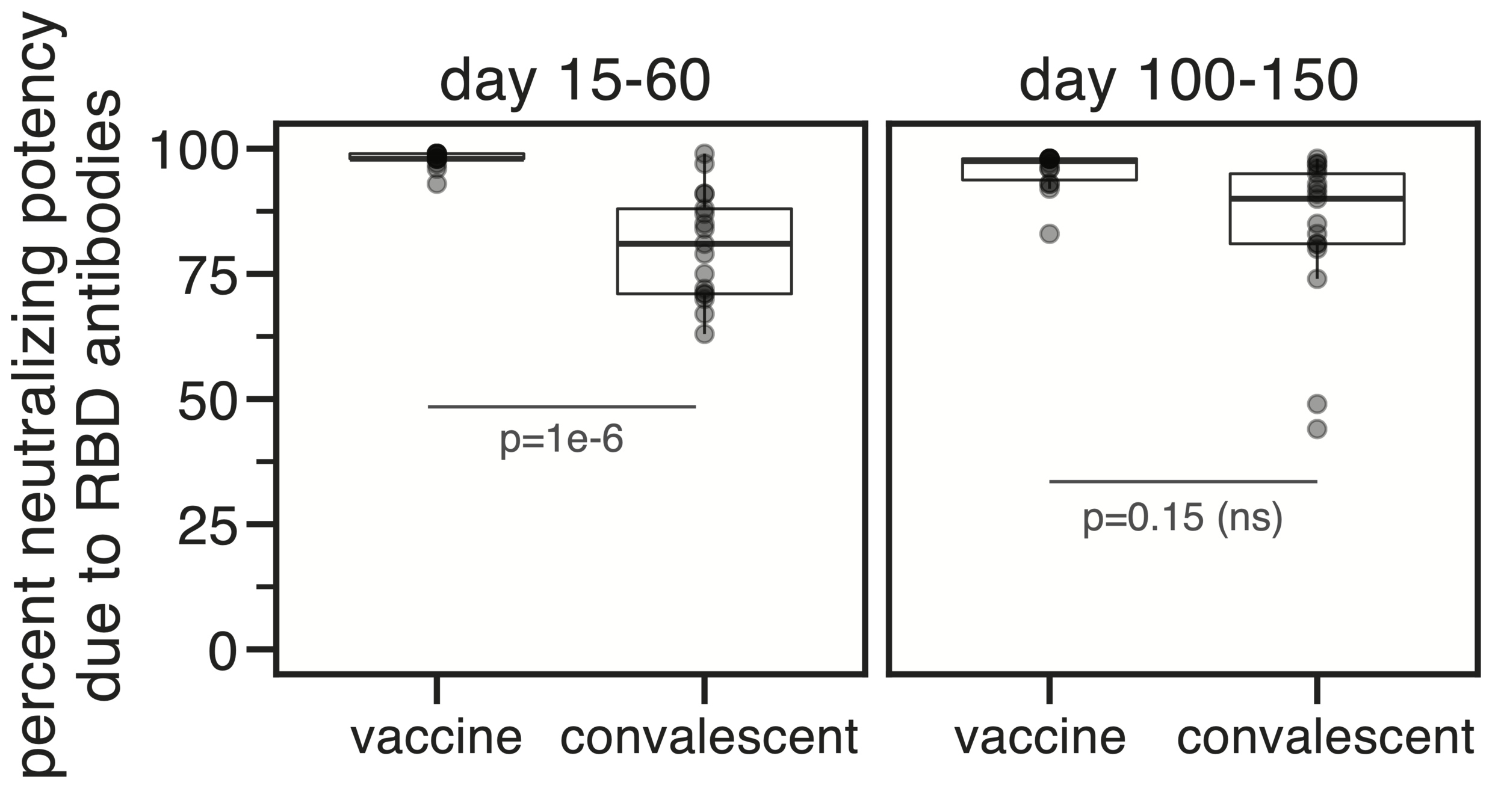
Importance of RBD
Human CoVs, which evolve to escape transmission-blocking immunity, show strongest selection in RBD.
So virus is telling us RBD antibodies matter most for blocking transmission. But other antibodies and T-cells still matter, and may reduce disease severity while putting less selection on virus.
Most neutralizing activity from RBD antibodies (although antibodies to other domains can also be neutralizing).
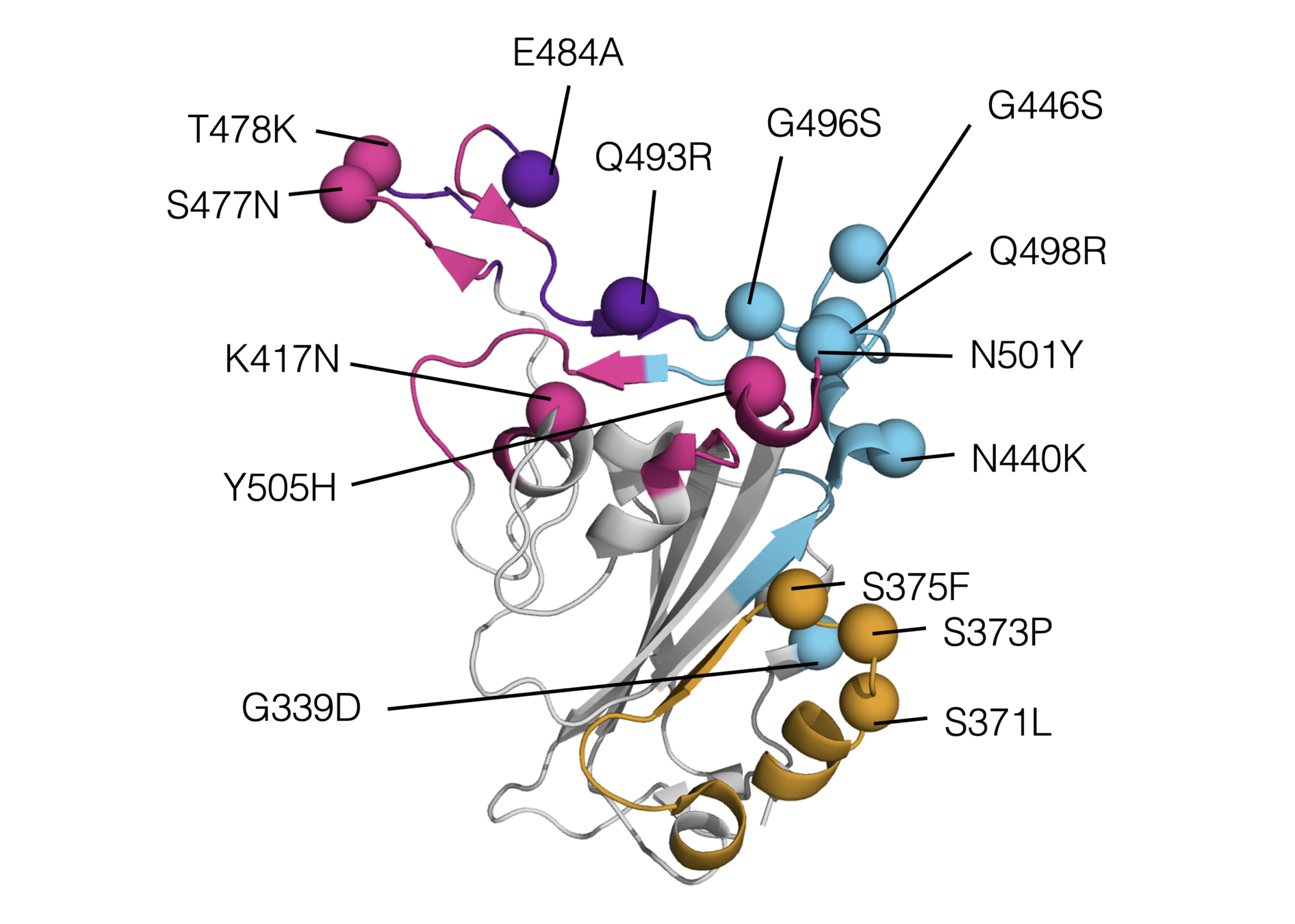
SARS-CoV-2 variants like Omicron have a lot of mutations in RBD
RBD mutations in Omicron BA.1
How does Omicron even bind ACE2 with so many mutations?
To answer this question, we measure how RBD mutations affect ACE2 affinity

RBD
fluorescent ACE2
yeast
fluorescent tag on RBD
Importantly, we use ACE2 titrations to measure true affinities, not just relative FACS binding signal; see here for details.
Deep mutational scanning: measurements for a library of all RBD mutations

Library of yeast each expressing a different RBD mutant. Click here for details on how library is made.
Interactive heatmaps are available here, and are from Starr et al, 2022 and unpublished work.
How does Omicron bind ACE2 with so many mutations?
Mutations in Omicron have net negative effect on ACE2 binding if summed as single mutants
How does Omicron bind ACE2 with so many mutations?


Mutations in Omicron have net negative effect on ACE2 binding if summed as single mutants
However, two mutations (Q498R & N501Y) work together so net effect ~zero when both present
ACE2 affinity-enhancing mutations buffer antibody-escape mutations
RBD will not run out of evolutionary space
25 of 31 residues in CoV-229E RBD that contact receptor varied during virus's evolution in humans over last ~50 years (Li et al, 2019)
There are lots of mutations to SARS-CoV-2 RBD that retain (and sometimes even enhance) ACE2 affinity (Starr et al, 2020; Starr et al, 2022)
Outline
- Principles of viral antigenic evolution and emergence of Omicron
- Importance of the SARS-CoV-2 receptor-binding domain (RBD)
- Mapping antibody escape to interpret viral evolution
We use deep mutational scanning to map all RBD mutations that escape antibody binding
RBD
fluorescently labeled antibody
yeast
fluorescent tag on RBD
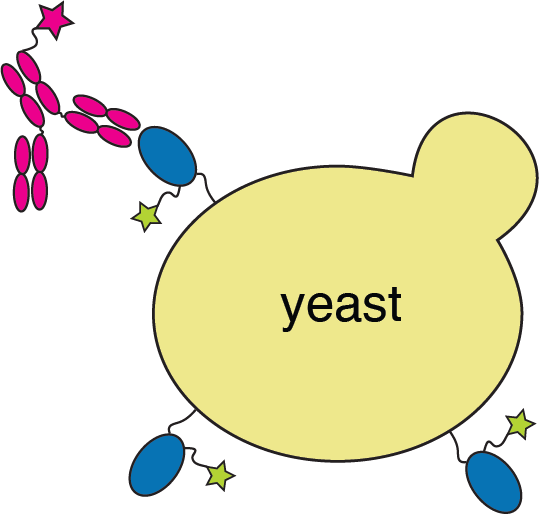
Experiments combine flow cytometry and deep sequencing of a library of yeast expressing all RBD mutants
For interactive escape map, see: https://jbloomlab.github.io/SARS-CoV-2-RBD_MAP_LY-CoV555/
Escape map from one antibody (LY-CoV555, ie bamlanivimab). Peaks indicate sites where mutations escape binding.
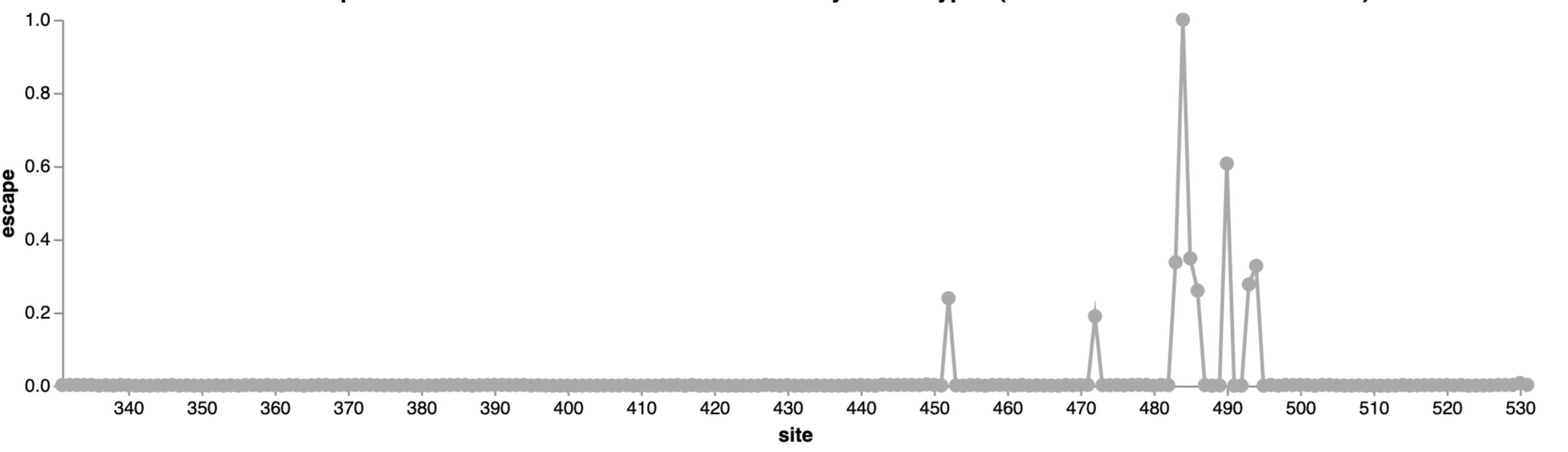
484
452
490
Infection / vaccination elicit polyclonal antibodies
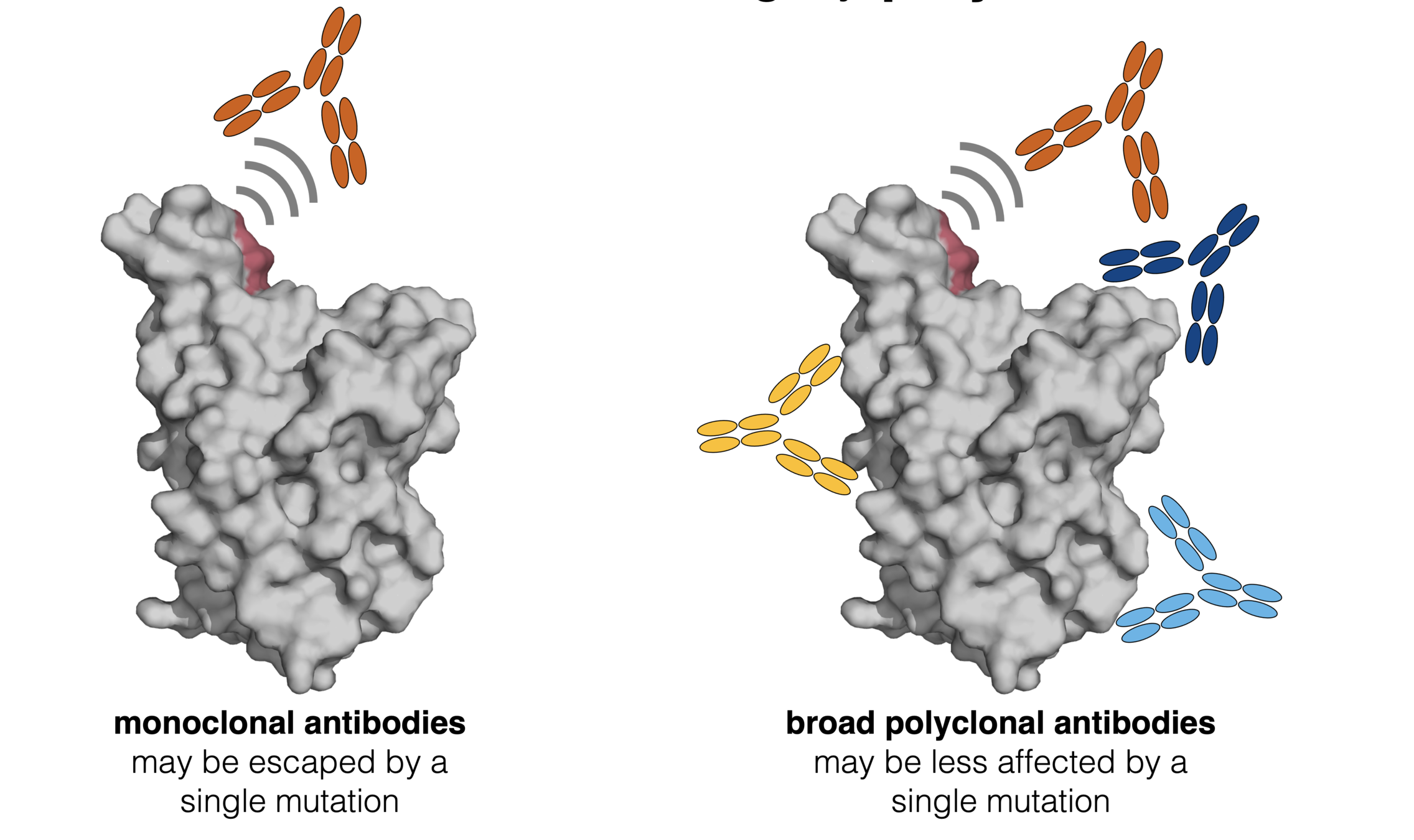
How do mutations affect polyclonal antibodies? First, consider an equal mix of three monoclonal antibodies.

Interactive version of this mini example is at https://jbloomlab.github.io/SARS2_RBD_Ab_escape_maps/mini-example-escape-calc/
LY-CoV555 is escaped at both sites 484 and 490, so mutating either site has same overall effect
Average escape across all antibodies

Mutating site 484 or 490 eliminates neutralization by antibody LY-CoV555, as reflected in thick black line showing average
Interactive version of this mini example is at https://jbloomlab.github.io/SARS2_RBD_Ab_escape_maps/mini-example-escape-calc/
Antibody-escape calculator extends this principle to deep mutational scanning data for ~1,500 different human antibodies
Escape calculator is described in Greaney et al (2022), and is available at https://jbloomlab.github.io/SARS2_RBD_Ab_escape_maps/escape-calc/
36 antibodies mapped by Tyler Starr & Allie Greaney in Bloom lab, from early SARS-CoV-2 strains
Antibody-escape calculator extends this principle to deep mutational scanning data for ~1,500 different human antibodies
Escape calculator is described in Greaney et al (2022), and is available at https://jbloomlab.github.io/SARS2_RBD_Ab_escape_maps/escape-calc/
36 antibodies mapped by Tyler Starr & Allie Greaney in Bloom lab, from early SARS-CoV-2 strains
1,522 (!) antibodies mapped by Sunney Xie, Richard Cao, Fanchong Jian, et al at Peking University. From early strains, BA.1, & patients with prior SARS-CoV-1 infection. See here.
What does escape calculator tell us about evolution that has already happened?
Antibodies elicited by early SARS-CoV-2 that neutralize Wuhan-Hu-1 heavily focus on sites like 484, 417, 446
417
446
484

417
446
484
Omicron BA.1 is mutated at many of these sites, which is why it was neutralized substantially less well by current vaccines

417
446
484
Omicron BA.2 has some different RBD mutations than BA.1, but similar overall escape from antibodies from early SARS-CoV-2

Can identify mutations that mediate further escape. In Dec 2021 we predicted 486 as likely site of future evolution--in April 2022, mutation F486V was identified in Omicron BA.4 and BA.5.
486 is largest site of escape for antibodies not already escaped by mutations in BA.2

What are likely next steps in SARS-CoV-2's antigenic evolution?
Sites of escape from antibodies elicited by early (pre-Omicron) infection/vaccination that still neutralize Omicron BA.2
346
444-446
452
450
486
499
mutated in BA.4/BA.5
mutated in BA.2.75

Sites of escape are somewhat different for antibodies from people who had an Omicron BA.1 breakthrough infection
346-348
444-446
356
Differences in antibody-escape mutations between people +/- BA.1 breakthrough infections is consistent with prior studies on other viruses showing that people with different exposure histories have different viral escape mutations.
452
486
mutated in BA.4/BA.5
mutated in BA.2.75

Explore yourself: https://jbloomlab.github.io/SARS2_RBD_Ab_escape_maps/escape-calc/
Conclusions
- A subset of human respiratory viruses evolve to escape immunity
- Unfortunately, coronaviruses are among this subset
- The most important antigenic evolution occurs in the RBD
- We can use deep mutational scanning to identify antigenic mutations
- Sites to watch: 346-348, 356, 444-446, 560, 452, 468, 486, 490, 499
Crowe lab (Vanderbilt)
Chu lab (Univ Wash)
Veesler lab (Univ Wash)
King lab (Univ Wash)
Li lab (Brigham & Women's)
Boeckh lab (Fred Hutch)
Alex Greninger (Univ Wash)
Nussenzweig lab (Rockefeller)
Bjorkman lab (Caltech)
Katie Kistler (Fred Hutch)






Tyler Starr
Allie Greaney
Rachel Eguia
Bloom lab (Fred Hutch)


Sarah Hilton
Kate Crawford


Andrea Loes
These slides at: