Open PHACTS
BioExcel use case #6
Leveraging integrated pharmacological
datasets for cross-domain queries



This work is licensed under a
Creative Commons Attribution 4.0 International License
.
BioExcel All Hands meeting, Schiphol, 2016-04-21

This work has been done as part of the BioExcel CoE ( www.bioexcel.eu),
a project funded by the EC H2020 program, EINFRA-5-2015 contract number 675728

Bringing together pharmacological data resources
in an integrated, interoperable infrastructure
Data sources integrated and linked together so that you can easily see the relationships between compounds, targets, pathways, diseases and tissues.
ChEBI, ChEMBL, ChemSpider, ConceptWiki, DisGeNET, DrugBank, FAERS, Gene Ontology, neXtProt, SureChEMBL, UniProt, WikiPathways

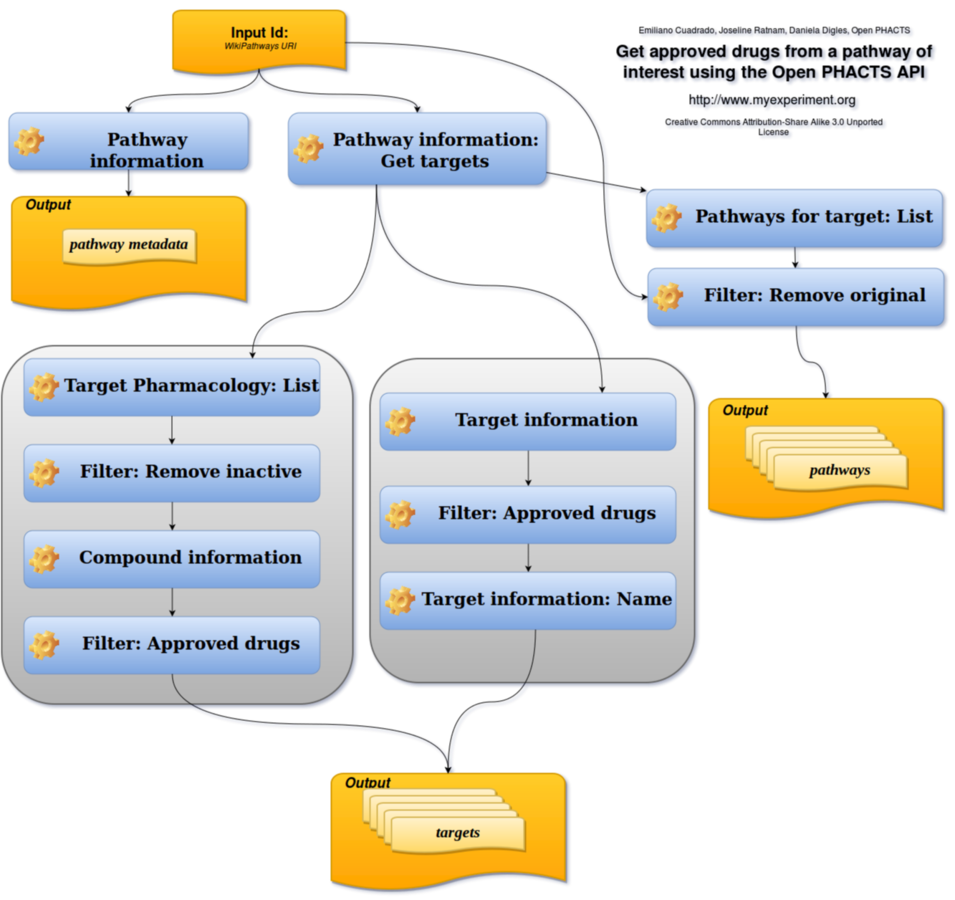
Workflow gap analysis
Combining private and public data ➟ Tutorials
Local install of Open PHACT ➟ Docker
Testing and comparing different APIs
Integrate third-party tools ➟ Identifier mappings
API changes ➟ Semantic versioning
Lack of flexibility for:
Workflow language, API URLs, Data sources
Common format for bioinformatics tool execution
Community based standards effort
Choose your own workflow engine
Designed for cluster & cloud environments
Designed for containers (e.g. Docker)
Main focus: command line tools
and ongoing discussions for super computer support
Adapted from Peter Amstutz' Broad Institute CWL meetup 2015-11-13
Implementors:
cwltool
Rabix
Arvados
Galaxy
Parallel Recipes
Toil
CancerCollaboratory
Airflow (SciDAP)
cwl2script
Apache Taverna

Apache Taverna



Tools and
Data Services Registry

Work so far
Apache Taverna:
Initiated CWL and Docker support
Open PHACTS:
Docker install
Data distribution of sources
Sure CHEMBL (patent data)
Future work
Docker tool for local/remote Open PHACTS API
CWL tool descriptions (workflow building blocks)
Use case 6 workflow as CWL
Configuration of data sources / API
Open PHACTS platform as EGI VM appliance
