Building pharmacological workflow blocks for virtual screening
Part 2: Workflows and deployment
Stian Soiland-Reyes, The University of Manchester


This work is licensed under a
Creative Commons Attribution 4.0 International License
.
BioExcel Webinar, 2017-07-17

This work has been done as part of the BioExcel CoE (www.bioexcel.eu),
a project funded by the EC H2020 program, EINFRA-5-2015 contract number 675728


BioExcel & Open PHACTS
{
"format": "linked-data-api",
"version": "1.5",
"result": {
"_about": "https://beta.openphacts.org/1.5/compound?uri=http%3A%2F%2Fwww.conceptwiki.org%2Fconcept%2F38932552-111f-4a4e-a46a-4ed1d7bdf9d5&app_id=161aeb7d&app_key=bbcba81896020f0b95e3dd35b55e3345&_format=json",
"definition": "https://beta.openphacts.org/api-config",
"extendedMetadataVersion": "https://beta.openphacts.org/1.5/compound?uri=http%3A%2F%2Fwww.conceptwiki.org%2Fconcept%2F38932552-111f-4a4e-a46a-4ed1d7bdf9d5&app_id=161aeb7d&app_key=bbcba81896020f0b95e3dd35b55e3345&_format=json&_metadata=all%2Cviews%2Cformats%2Cexecution%2Cbindings%2Csite",
"linkPredicate": "http://www.w3.org/2004/02/skos/core#exactMatch",
"activeLens": "Default",
"primaryTopic": {
"_about": "http://www.conceptwiki.org/concept/38932552-111f-4a4e-a46a-4ed1d7bdf9d5",
"inDataset": "http://www.conceptwiki.org",
"exactMatch": [
{
"_about": "http://bio2rdf.org/drugbank:DB00398",
"description_en": "Sorafenib (rINN), marketed as Nexavar by Bayer, is a drug approved for the treatment of advanced renal cell carcinoma (primary kidney cancer). It has also received \"Fast Track\" designation by the FDA for the treatment of advanced hepatocellular carcinoma (primary liver cancer), and has since performed well in Phase III trials.\nSorafenib is a small molecular inhibitor of Raf kinase, PDGF (platelet-derived growth factor), VEGF receptor 2 & 3 kinases and c Kit the receptor for Stem cell factor. A growing number of drugs target most of these pathways. The originality of Sorafenib lays in its simultaneous targeting of the Raf/Mek/Erk pathway.",
"description": "Sorafenib (rINN), marketed as Nexavar by Bayer, is a drug approved for the treatment of advanced renal cell carcinoma (primary kidney cancer). It has also received \"Fast Track\" designation by the FDA for the treatment of advanced hepatocellular carcinoma (primary liver cancer), and has since performed well in Phase III trials.\nSorafenib is a small molecular inhibitor of Raf kinase, PDGF (platelet-derived growth factor), VEGF receptor 2 & 3 kinases and c Kit the receptor for Stem cell factor. A growing number of drugs target most of these pathways. The originality of Sorafenib lays in its simultaneous targeting of the Raf/Mek/Erk pathway.",
"drugType_en": [
"investigational",
"approved"
],
"drugType": [
"investigational",
"approved"
],
"genericName_en": "Sorafenib",
"genericName": "Sorafenib",
"metabolism_en": "Sorafenib is metabolized primarily in the liver, undergoing oxidative metabolism, mediated by CYP3A4, as well as glucuronidation mediated by UGT1A9. Sorafenib accounts for approximately 70-85% of the circulating analytes in plasma at steady- state. Eight metabolites of sorafenib have been identified, of which five have been detected in plasma. The main circulating metabolite of sorafenib in plasma, the pyridine N-oxide, shows in vitro potency similar to that of sorafenib. This metabolite comprises approximately 9-16% of circulating analytes at steady-state.",
"metabolism": "Sorafenib is metabolized primarily in the liver, undergoing oxidative metabolism, mediated by CYP3A4, as well as glucuronidation mediated by UGT1A9. Sorafenib accounts for approximately 70-85% of the circulating analytes in plasma at steady- state. Eight metabolites of sorafenib have been identified, of which five have been detected in plasma. The main circulating metabolite of sorafenib in plasma, the pyridine N-oxide, shows in vitro potency similar to that of sorafenib. This metabolite comprises approximately 9-16% of circulating analytes at steady-state.",
"proteinBinding_en": "99.5% bound to plasma proteins.",
"proteinBinding": "99.5% bound to plasma proteins.",
"toxicity_en": "The highest dose of sorafenib studied clinically is 800 mg twice daily. The adverse reactions observed at this dose were primarily diarrhea and dermatologic events. No information is available on symptoms of acute overdose in animals because of the saturation of absorption in oral acute toxicity studies conducted in animals.",
"toxicity": "The highest dose of sorafenib studied clinically is 800 mg twice daily. The adverse reactions observed at this dose were primarily diarrhea and dermatologic events. No information is available on symptoms of acute overdose in animals because of the saturation of absorption in oral acute toxicity studies conducted in animals.",
"inDataset": "http://www.openphacts.org/bio2rdf/drugbank",
"drugInteraction": [
{
"_about": "http://bio2rdf.org/drugbank_resource:DB00398_DB00755",
"text_en": "DDI between Sorafenib and Tretinoin - The strong CYP2C8 inhibitor, Sorafenib, may decrease the metabolism and clearance of oral Tretinoin. Consider alternate therapy or monitor for changes in Tretinoin effectiveness and adverse/toxic effects if Sorafenib is initiated, discontinued to dose changed.",
"text": "DDI between Sorafenib and Tretinoin - The strong CYP2C8 inhibitor, Sorafenib, may decrease the metabolism and clearance of oral Tretinoin. Consider alternate therapy or monitor for changes in Tretinoin effectiveness and adverse/toxic effects if Sorafenib is initiated, discontinued to dose changed.",
"inDataset": "http://www.openphacts.org/bio2rdf/drugbank",
"interactingDrug": "http://bio2rdf.org/drugbank:DB00755"
},
{
"_about": "http://bio2rdf.org/drugbank_resource:DB00398_DB00958",
"text_en": "DDI between Sorafenib and Carboplatin - Sorafenib may enhance the adverse/toxic effect of carboplatin. Concurrent use of sorafenib with carboplatin and placlitaxel in patients with squamous cell lung cancer is contraindicated. The use of this combination in other settings is not specifically contraindicated, but any such use should be approached with added caution.",
"text": "DDI between Sorafenib and Carboplatin - Sorafenib may enhance the adverse/toxic effect of carboplatin. Concurrent use of sorafenib with carboplatin and placlitaxel in patients with squamous cell lung cancer is contraindicated. The use of this combination in other settings is not specifically contraindicated, but any such use should be approached with added caution.",
"inDataset": "http://www.openphacts.org/bio2rdf/drugbank",
"interactingDrug": "http://bio2rdf.org/drugbank:DB00958"
},
{
"_about": "http://bio2rdf.org/drugbank_resource:DB00398_DB06414",
"text_en": "DDI between Sorafenib and Etravirine - Sorafebib, when used concomitantly with etravirine, may experience a decrease in serum concentration. It is recommended to avoid concurrent therapy.",
"text": "DDI between Sorafenib and Etravirine - Sorafebib, when used concomitantly with etravirine, may experience a decrease in serum concentration. It is recommended to avoid concurrent therapy.",
"inDataset": "http://www.openphacts.org/bio2rdf/drugbank",
"interactingDrug": "http://bio2rdf.org/drugbank:DB06414"
},
{
"_about": "http://bio2rdf.org/drugbank_resource:DB00072_DB00398",
"text_en": "DDI between Trastuzumab and Sorafenib - Trastuzumab may increase the risk of neutropenia and anemia. Monitor closely for signs and symptoms of adverse events.",
"text": "DDI between Trastuzumab and Sorafenib - Trastuzumab may increase the risk of neutropenia and anemia. Monitor closely for signs and symptoms of adverse events.",
"inDataset": "http://www.openphacts.org/bio2rdf/drugbank",
"interactingDrug": "http://bio2rdf.org/drugbank:DB00072"
},
{
"_about": "http://bio2rdf.org/drugbank_resource:DB00112_DB00398",
"text_en": "DDI between Bevacizumab and Sorafenib - Monitor therapy due to increased adverse effects of sorafenib, especially hand-foot skin reaction.",
"text": "DDI between Bevacizumab and Sorafenib - Monitor therapy due to increased adverse effects of sorafenib, especially hand-foot skin reaction.",
"inDataset": "http://www.openphacts.org/bio2rdf/drugbank",
"interactingDrug": "http://bio2rdf.org/drugbank:DB00112"
}
]
},
{
"_about": "http://aers.data2semantics.org/resource/drug/SORAFENIB",
"inDataset": "http://aers.data2semantics.org/",
"reportedAdverseEvent": [
{
"_about": "http://aers.data2semantics.org/resource/diagnosis/CARDIAC_FAILURE_ACUTE",
"inDataset": "http://aers.data2semantics.org/",
"prefLabel": "CARDIAC FAILURE ACUTE"
},
{
"_about": "http://aers.data2semantics.org/resource/diagnosis/RENAL_IMPAIRMENT",
"inDataset": "http://aers.data2semantics.org/",
"prefLabel": "RENAL IMPAIRMENT"
},
{
"_about": "http://aers.data2semantics.org/resource/diagnosis/HYPERURICAEMIA",
"inDataset": "http://aers.data2semantics.org/",
"prefLabel": "HYPERURICAEMIA"
},
{
"_about": "http://aers.data2semantics.org/resource/diagnosis/TUMOUR_LYSIS_SYNDROME",
"inDataset": "http://aers.data2semantics.org/",
"prefLabel": "TUMOUR LYSIS SYNDROME"
},
{
"_about": "http://aers.data2semantics.org/resource/diagnosis/LEFT_VENTRICULAR_DYSFUNCTION",
"inDataset": "http://aers.data2semantics.org/",
"prefLabel": "LEFT VENTRICULAR DYSFUNCTION"
},
{
"_about": "http://aers.data2semantics.org/resource/diagnosis/METABOLIC_ACIDOSIS",
"inDataset": "http://aers.data2semantics.org/",
"prefLabel": "METABOLIC ACIDOSIS"
}
],
},
{
"_about": "http://rdf.ebi.ac.uk/resource/chembl/molecule/CHEMBL1336",
"mw_freebase": 464.82,
"inDataset": "http://www.ebi.ac.uk/chembl",
"type": "http://rdf.ebi.ac.uk/terms/chembl#SmallMolecule"
},
{
"_about": "http://ops.rsc.org/OPS379634",
"inDataset": "http://ops.rsc.org",
"hba": 7,
"hbd": 3,
"inchi": "InChI=1S/C21H16ClF3N4O3/c1-26-19(30)18-11-15(8-9-27-18)32-14-5-2-12(3-6-14)28-20(31)29-13-4-7-17(22)16(10-13)21(23,24)25/h2-11H,1H3,(H,26,30)(H2,28,29,31)",
"inchikey": "MLDQJTXFUGDVEO-UHFFFAOYSA-N",
"logp": 5.158,
"molformula": "C21H16ClF3N4O3",
"molweight": 464.825,
"psa": 92.35,
"ro5_violations": 1,
"rtb": 5,
"smiles": "CNC(=O)C1=NC=CC(=C1)OC2=CC=C(C=C2)NC(=O)NC3=CC(=C(C=C3)Cl)C(F)(F)F"
}
],
"prefLabel_en": "Sorafenib",
"prefLabel": "Sorafenib",
"isPrimaryTopicOf": "https://beta.openphacts.org/1.5/compound?uri=http%3A%2F%2Fwww.conceptwiki.org%2Fconcept%2F38932552-111f-4a4e-a46a-4ed1d7bdf9d5&app_id=161aeb7d&app_key=bbcba81896020f0b95e3dd35b55e3345&_format=json"
}
}
}<?xml version="1.0" encoding="utf-8"?>
<result format="linked-data-api" version="1.5" href="https://beta.openphacts.org/1.5/compound?uri=http%3A%2F%2Fwww.conceptwiki.org%2Fconcept%2F38932552-111f-4a4e-a46a-4ed1d7bdf9d5&app_id=161aeb7d&app_key=bbcba81896020f0b95e3dd35b55e3345&_format=xml">
<primaryTopic href="http://www.conceptwiki.org/concept/38932552-111f-4a4e-a46a-4ed1d7bdf9d5">
<prefLabel xml:lang="en">Sorafenib</prefLabel>
<exactMatch>
<item href="http://rdf.ebi.ac.uk/resource/chembl/molecule/CHEMBL1336">
<type href="http://rdf.ebi.ac.uk/terms/chembl#SmallMolecule"/>
<inDataset href="http://www.ebi.ac.uk/chembl"/>
<mw_freebase datatype="double">464.82</mw_freebase>
</item>
<item href="http://ops.rsc.org/OPS379634">
<smiles>CNC(=O)C1=NC=CC(=C1)OC2=CC=C(C=C2)NC(=O)NC3=CC(=C(C=C3)Cl)C(F)(F)F</smiles>
<rtb datatype="double">5.0</rtb>
<ro5_violations datatype="double">1.0</ro5_violations>
<psa datatype="double">92.35</psa>
<molweight datatype="double">464.825</molweight>
<molformula>C21H16ClF3N4O3</molformula>
<logp datatype="double">5.158</logp>
<inchikey>MLDQJTXFUGDVEO-UHFFFAOYSA-N</inchikey>
<inchi>InChI=1S/C21H16ClF3N4O3/c1-26-19(30)18-11-15(8-9-27-18)32-14-5-2-12(3-6-14)28-20(31)29-13-4-7-17(22)16(10-13)21(23,24)25/h2-11H,1H3,(H,26,30)(H2,28,29,31)</inchi>
<hbd datatype="double">3.0</hbd>
<hba datatype="double">7.0</hba>
<inDataset href="http://ops.rsc.org"/>
</item>
<item href="http://aers.data2semantics.org/resource/drug/NEXAVAR">
<prefLabel>NEXAVAR</prefLabel>
<reportedAdverseEvent>
<item href="http://aers.data2semantics.org/resource/diagnosis/HEAD_INJURY">
<prefLabel>HEAD INJURY</prefLabel>
<inDataset href="http://aers.data2semantics.org/"/>
</item>
<item href="http://aers.data2semantics.org/resource/diagnosis/SUPRAVENTRICULAR_TACHYCARDIA">
<prefLabel>SUPRAVENTRICULAR TACHYCARDIA</prefLabel>
<inDataset href="http://aers.data2semantics.org/"/>
</item>
<!-- .. -->
</reportedAdverseEvent>
<inDataset href="http://aers.data2semantics.org/"/>
</item>
<item href="http://www.conceptwiki.org/concept/38932552-111f-4a4e-a46a-4ed1d7bdf9d5"/>
<item href="http://bio2rdf.org/drugbank:DB00398">
<drugInteraction>
<item href="http://bio2rdf.org/drugbank_resource:DB00398_DB00755">
<interactingDrug href="http://bio2rdf.org/drugbank:DB00755"/>
<inDataset href="http://www.openphacts.org/bio2rdf/drugbank"/>
<text xml:lang="en">DDI between Sorafenib and Tretinoin - The strong CYP2C8 inhibitor, Sorafenib, may decrease the metabolism and clearance of oral Tretinoin. Consider alternate therapy or monitor for changes in Tretinoin effectiveness and adverse/toxic effects if Sorafenib is initiated, discontinued to dose changed.</text>
</item>
<item href="http://bio2rdf.org/drugbank_resource:DB00398_DB00958">
<interactingDrug href="http://bio2rdf.org/drugbank:DB00958"/>
<inDataset href="http://www.openphacts.org/bio2rdf/drugbank"/>
<text xml:lang="en">DDI between Sorafenib and Carboplatin - Sorafenib may enhance the adverse/toxic effect of carboplatin. Concurrent use of sorafenib with carboplatin and placlitaxel in patients with squamous cell lung cancer is contraindicated. The use of this combination in other settings is not specifically contraindicated, but any such use should be approached with added caution.</text>
</item>
<!-- .. -->
</drugInteraction>
<inDataset href="http://www.openphacts.org/bio2rdf/drugbank"/>
<toxicity xml:lang="en">The highest dose of sorafenib studied clinically is 800 mg twice daily. The adverse reactions observed at this dose were primarily diarrhea and dermatologic events. No information is available on symptoms of acute overdose in animals because of the saturation of absorption in oral acute toxicity studies conducted in animals.</toxicity>
<proteinBinding xml:lang="en">99.5% bound to plasma proteins.</proteinBinding>
<metabolism xml:lang="en">Sorafenib is metabolized primarily in the liver, undergoing oxidative metabolism, mediated by CYP3A4, as well as glucuronidation mediated by UGT1A9. Sorafenib accounts for approximately 70-85% of the circulating analytes in plasma at steady- state. Eight metabolites of sorafenib have been identified, of which five have been detected in plasma. The main circulating metabolite of sorafenib in plasma, the pyridine N-oxide, shows <i>in vitro</i> potency similar to that of sorafenib. This metabolite comprises approximately 9-16% of circulating analytes at steady-state.</metabolism>
<genericName xml:lang="en">Sorafenib</genericName>
<drugType>
<item xml:lang="en">investigational</item>
<item xml:lang="en">approved</item>
</drugType>
<description xml:lang="en">Sorafenib (rINN), marketed as Nexavar by Bayer, is a drug approved for the treatment of advanced renal cell carcinoma (primary kidney cancer). It has also received "Fast Track" designation by the FDA for the treatment of advanced hepatocellular carcinoma (primary liver cancer), and has since performed well in Phase III trials.
Sorafenib is a small molecular inhibitor of Raf kinase, PDGF (platelet-derived growth factor), VEGF receptor 2 & 3 kinases and c Kit the receptor for Stem cell factor. A growing number of drugs target most of these pathways. The originality of Sorafenib lays in its simultaneous targeting of the Raf/Mek/Erk pathway.</description>
</item>
</exactMatch>
<inDataset href="http://www.conceptwiki.org"/>
</primaryTopic>
<activeLens>Default</activeLens>
<linkPredicate href="http://www.w3.org/2004/02/skos/core#exactMatch"/>
<extendedMetadataVersion href="https://beta.openphacts.org/1.5/compound?uri=http%3A%2F%2Fwww.conceptwiki.org%2Fconcept%2F38932552-111f-4a4e-a46a-4ed1d7bdf9d5&app_id=161aeb7d&app_key=bbcba81896020f0b95e3dd35b55e3345&_format=xml&_metadata=all%2Cviews%2Cformats%2Cexecution%2Cbindings%2Csite"/>
<definition href="https://beta.openphacts.org/api-config"/>
</result>
Linux Container technology
..light-weight "virtual" virtual machine
A container is started from a image
Images downloaded from Docker Hub
Dockerfile: Layer-based recipe
Philosophy: One service, one image → microservices
Cloud's best friend: scalable, reproducible, customizable


ops-docker
Docker Compose

Which images to download
Which data volumes to use
Which network ports are exposed
How are containers linked
How to start/stop the containers
# Open PHACTS platform
# Docker Compose configuration
explorer:
image: openphacts/explorer2
ports:
- "3001:3000"
links:
- api
environment:
- API_URL=http://localhost:3002
#restart: always
api:
image: openphacts/ops-linkeddataapi
ports:
- "3002:80"
links:
- ims
- memcached
- virtuoso:sparql
# SPARQL server
virtuoso:
build: virtuoso-ops
ports:
- "3003:8890"
volumes_from:
- virtuosodata
virtuosodata:
image: busybox
volumes:
- /virtuoso
mysqldata:
image: busybox
volumes:
- /var/lib/mysql
mysql:
image: mysql
volumes_from:
- mysqldata
environment:
- MYSQL_ROOT_PASSWORD=uCie0ahgah
- MYSQL_DATABASE=ims
- MYSQL_USER=ims
- MYSQL_PASSWORD=ims
ims:
image: openphacts/identitymappingservice
ports:
- "3004:8080"
links:
- mysql
memcached:
image: memcached
mysqlstaging:
container_name: ops-mysqlstaging
image: openphacts/identitymappingservice-staging
links:
- mysql
# Populate RDF from virtuoso backup download
virtuosostaging:
build: virtuosodata-frombackup
volumes_from:
- virtuosodata
# To customize RDF dataloading, comment OUT the above 'virtuosostaging' block,
# uncomment the below block, and then run
# docker-compose up -d virtuosostagingrdf
#
## BEGIN custom loading
### Download from data.openphacts.org
#openphactsrdf:
# build: openphacts-rdf
# volumes:
# # To specify alternative data folder, use instad:
# # - /media/big-SSD/download:/download
# # - /media/big-SSD/staging:/staging
# - /download
# - /staging
# # /download
#
### Load into virtuoso
#virtuosostagingrdf:
# build: virtuosodata-fromrdf
# volumes_from:
# - virtuosodata
# - openphactsrdf
### END custom loading
## Future services
#elasticsearch:
# container_name: ops-elasticsearch
# image: elasticsearch
## TODO: Data loading
#ops-search:
# container_name: ops-search
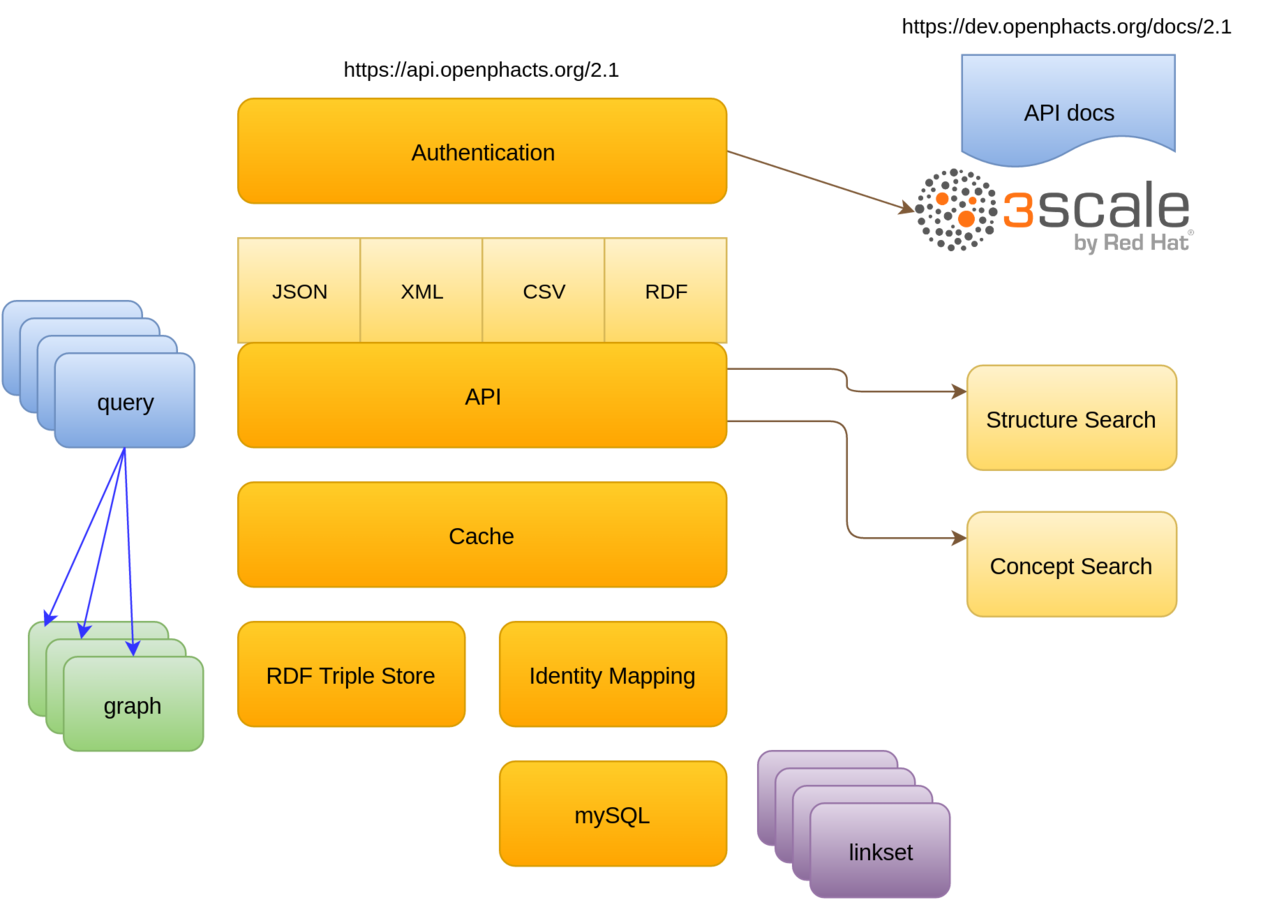
# image: openphacts/ops-searchAPI architecture

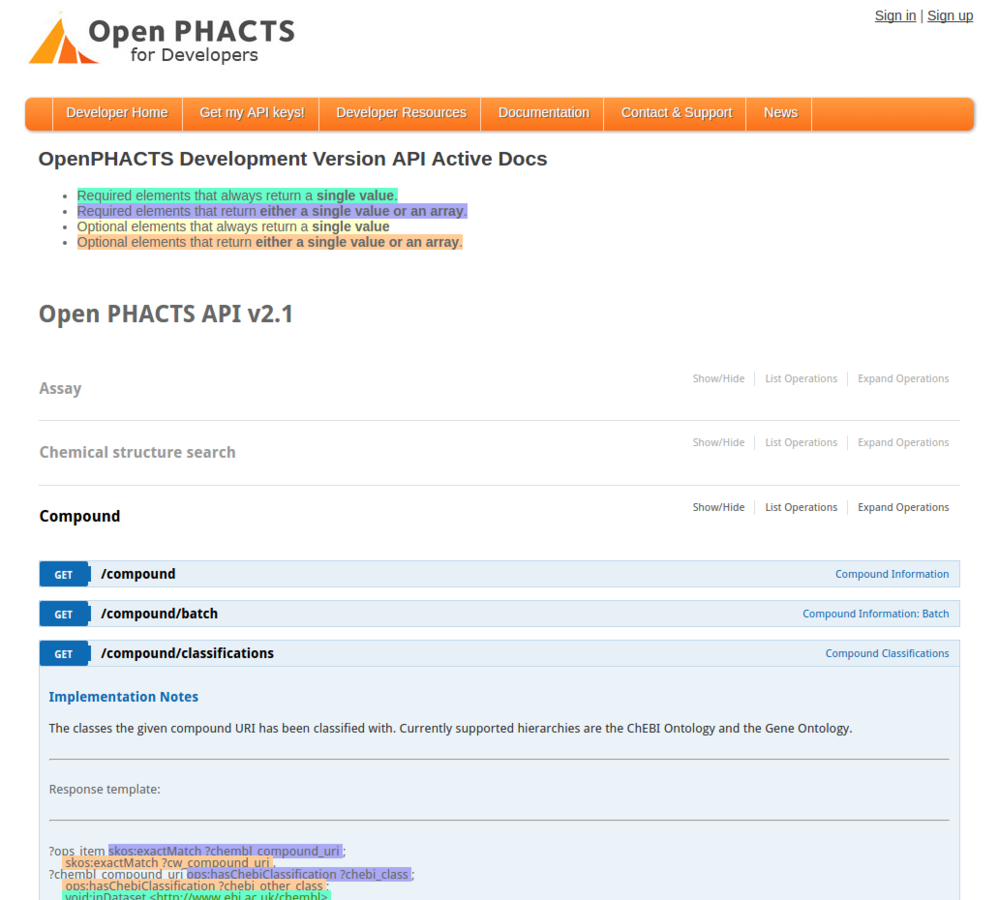
Get the software
$ curl -L https://github.com/openphacts/ops-docker/archive/master.tar.gz | tar xzv
$ cd ops-docker-master
$ sudo docker-compose pull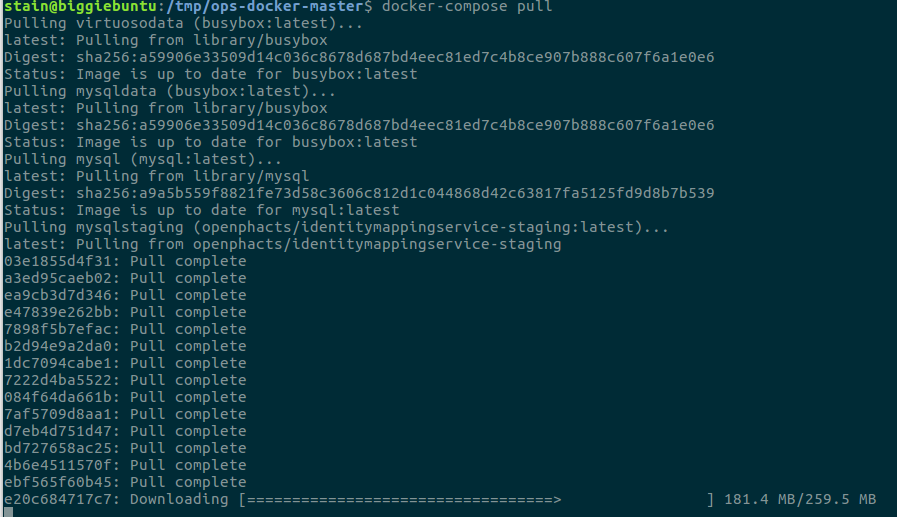
Get the data
$ sudo docker-compose up --no-recreate -d mysqlstaging virtuosostaging
$ sudo docker-compose logs mysqlstaging virtuosostaging
ops-mysqlstaging | mySQL staging finished
ops-mysqlstaging exited with code 0
ops-virtuosostaging | 09:13:35 --> Backup file # 675 [0x3F02-0x74-0x8A]
ops-virtuosostaging | 09:13:36 --> Backup file # 676 [0x3F02-0x74-0x8A]
ops-virtuosostaging | 09:13:37 End of restoring from backup, 6751701 pages
ops-virtuosostaging | 09:13:37 Server exiting
ops-virtuosostaging | Loading completed
ops-virtuosostaging exited with code 0
Start the services
$ sudo docker-compose up --no-recreate -d
$ sudo docker-compose logs --tail=5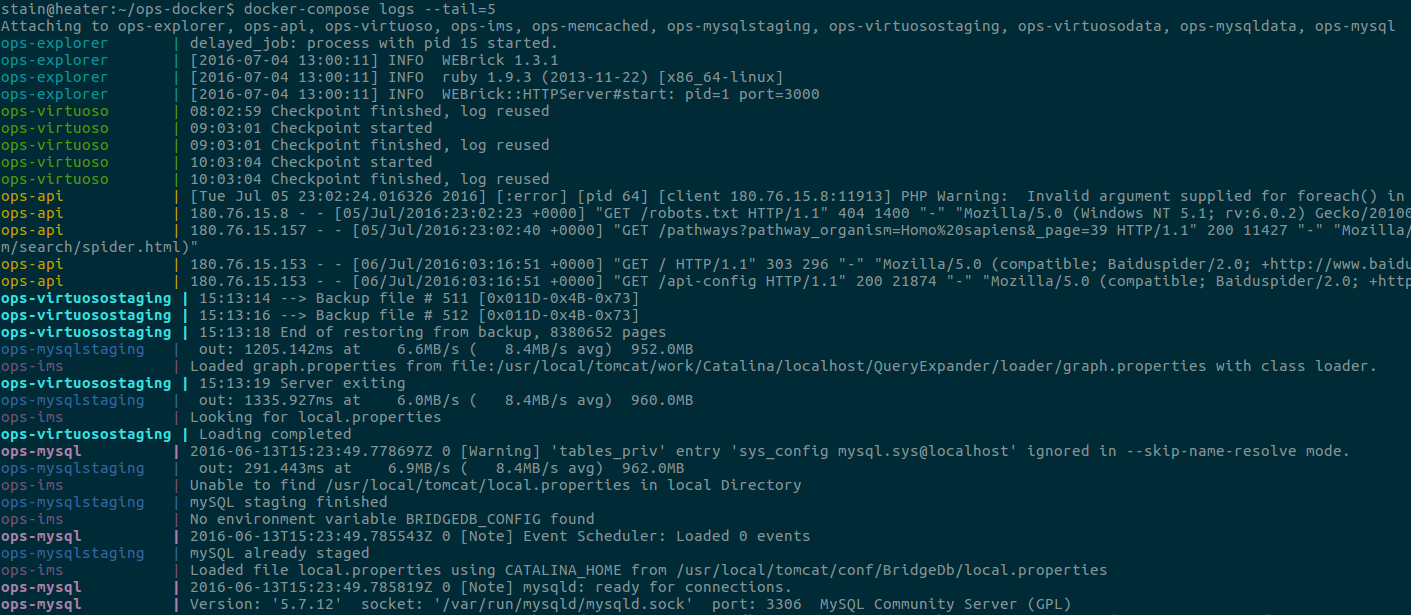
http://localhost:3001/ Explorer
http://localhost:3002/ REST API
http://localhost:3003/ SPARQL queries
http://localhost:3004/QueryExpander Identity Mapping
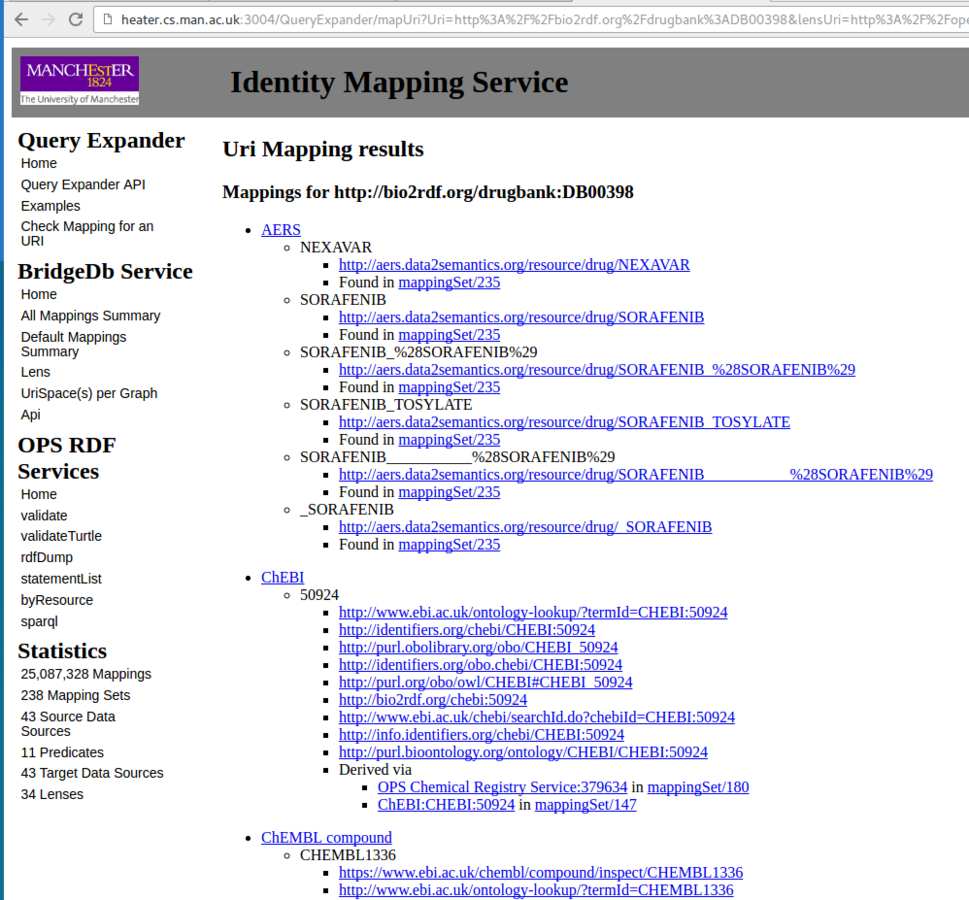
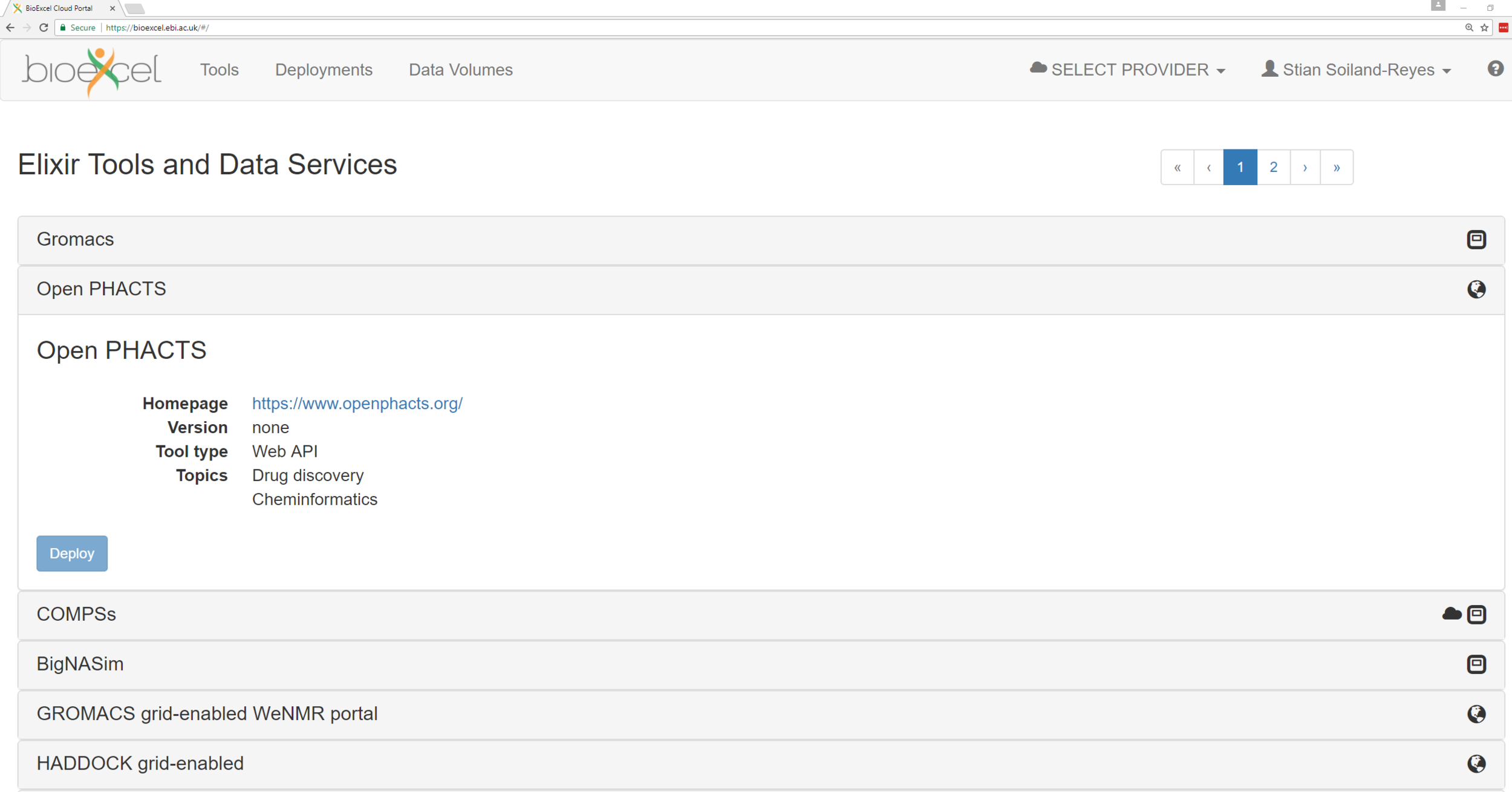
Deploy
Virtual Machine Images


Coming soon to a cloud near you
Using Open PHACTS from Workflows
BIOVIA Pipeline Pilot
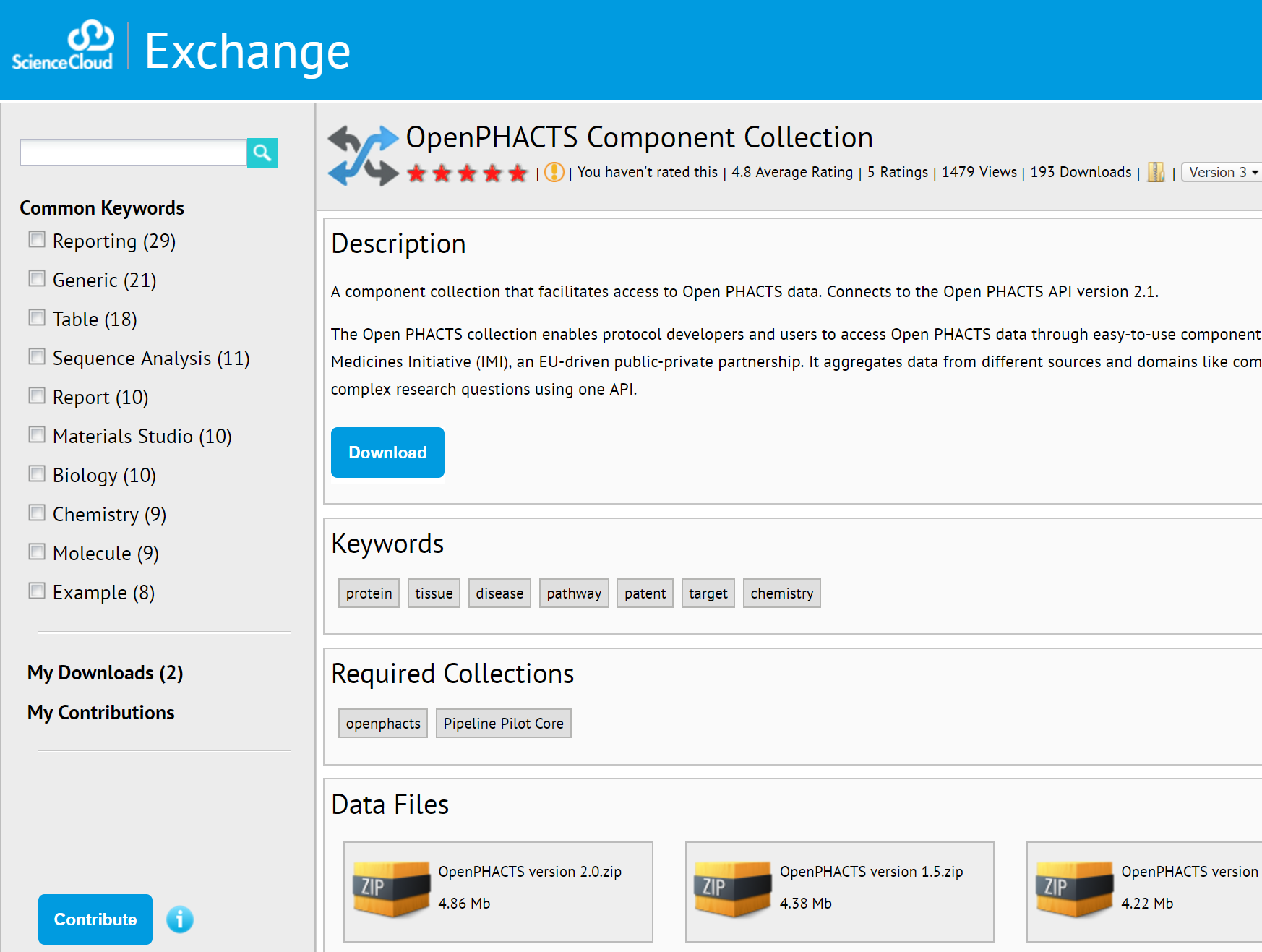
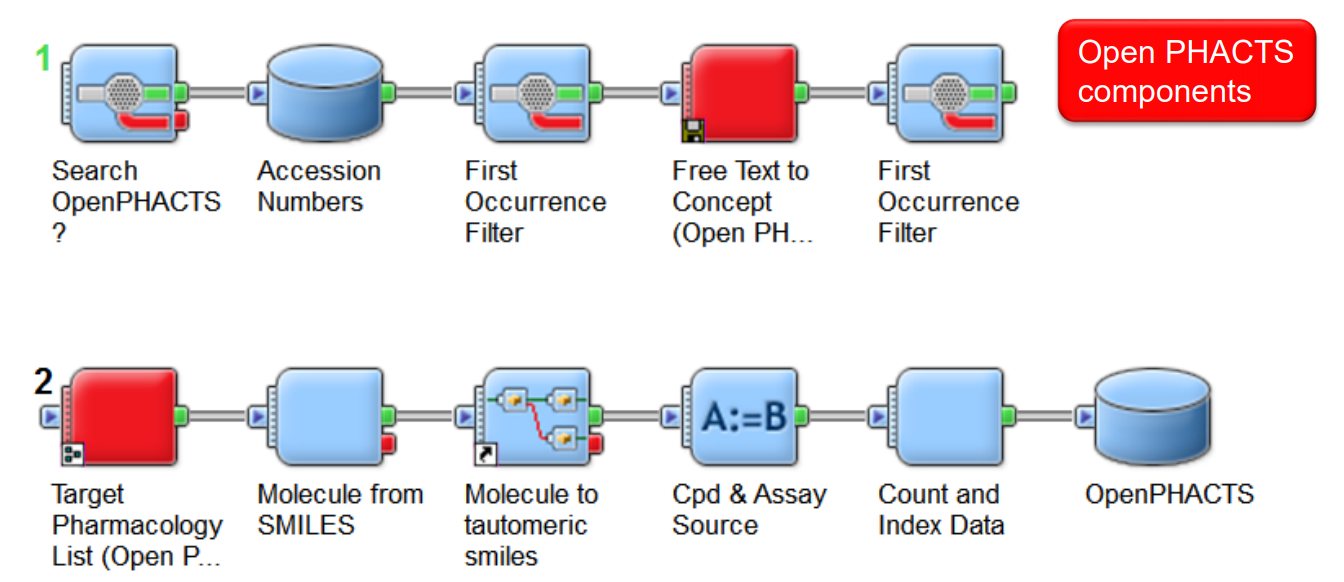
Jean-Marc Neefs, Janssen

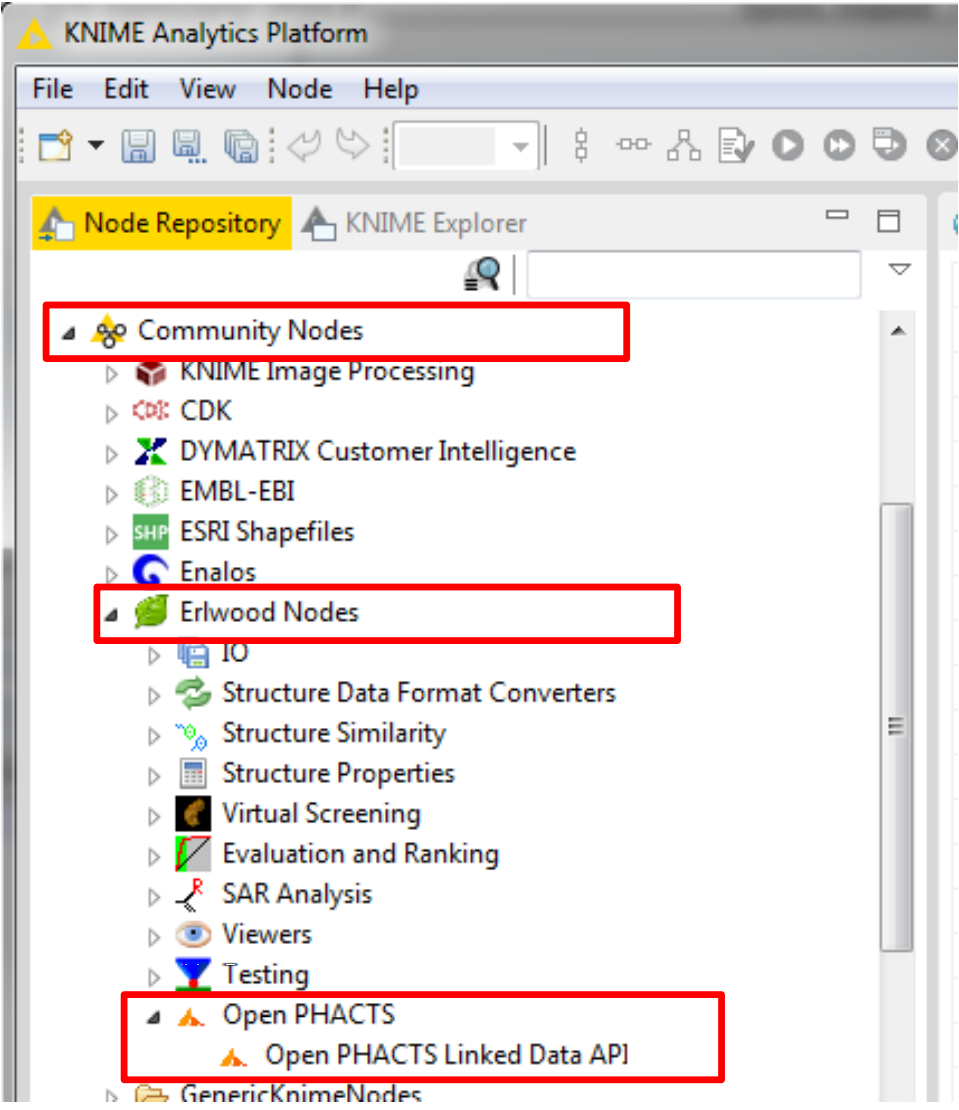
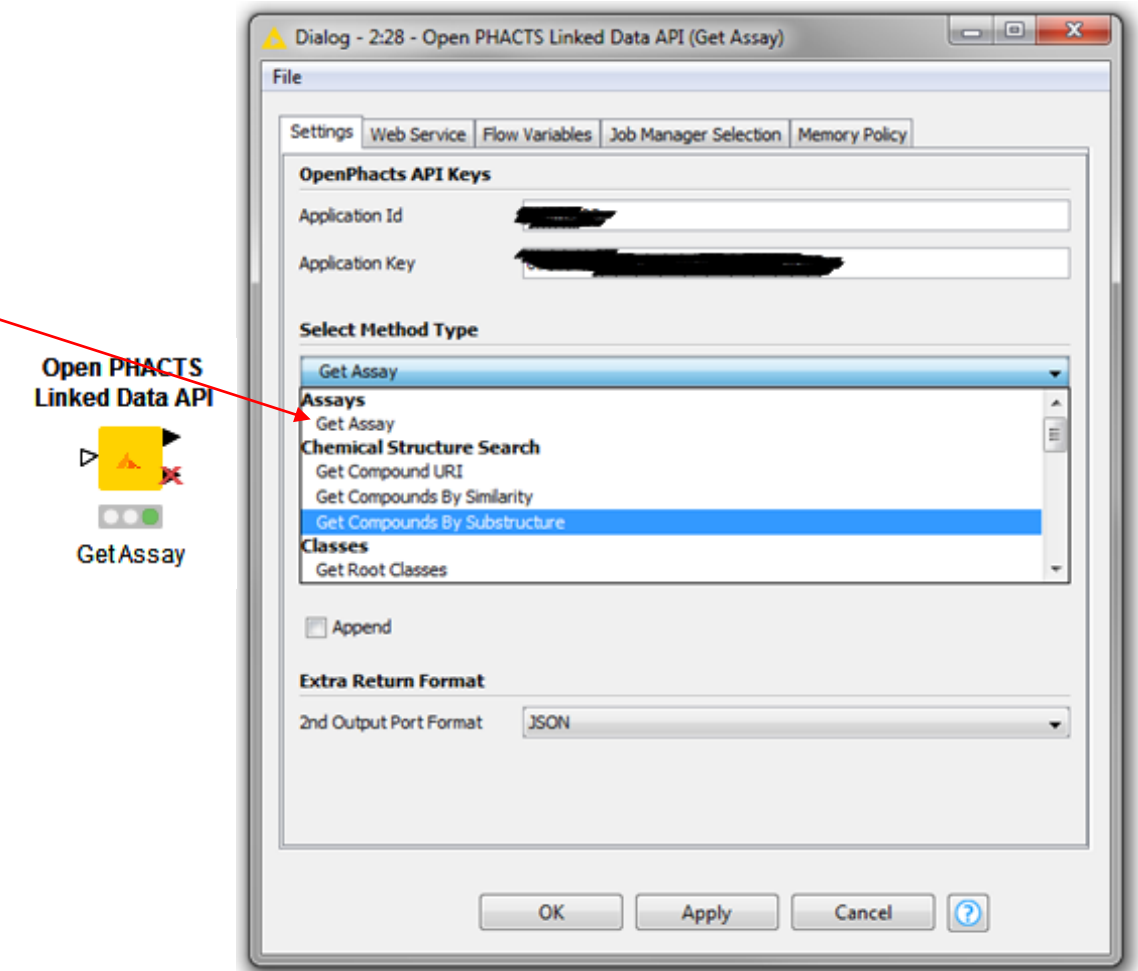

Coming soon to a KNIME node near you
James A. Lumley, Eli Lilly

Common format for pipeline execution
Community-based standards effort
Implemented by multiple workflow engines
Defined with a schema, specification, tests
Designed for shared nothing cluster/cloud
Designed for containers (e.g. Docker)
Main focus: command line tools
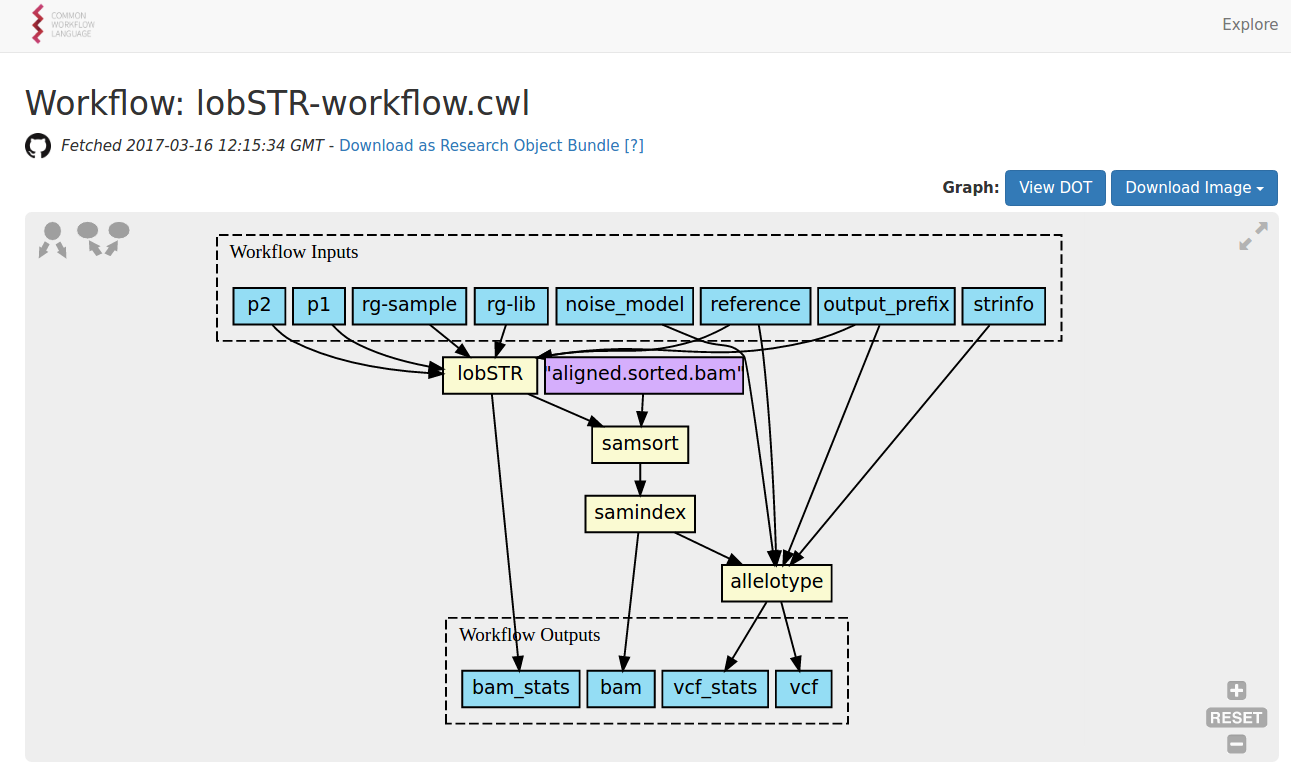
#!/usr/bin/env cwl-runner
class: Workflow
inputs:
- id: pairedEnds
type:
- type: array
items: File
description: list of files containing the first end of paired end reads in fasta or fastq format
outputs:
- id: bam
type: File
source: "#samindex/bam_with_bai"
steps:
- id: lobSTR
run: lobSTR-tool.cwl
inputs:
- { id: p1, source: "#pairedEnds" }
- { id: p2, source: "#p2" }
- { id: output_prefix, source: "#output_prefix" }
- { id: reference, source: "#reference" }
- { id: rg-sample, source: "#rg-sample" }
- { id: rg-lib, source: "#rg-lib" }
outputs:
- { id: bam }
- { id: bam_stats }
- id: samsort
run: samtools-sort.cwl
inputs:
- { id: input, source: "#lobSTR/bam" }
- { id: output_name, default: "aligned.sorted.bam" }
outputs:
- { id: output_file }
- id: samindex
run: samtools-index.cwl
inputs:
- { id: input, source: "#samsort/output_file" }
outputs:
- { id: bam_with_bai }
- id: allelotype
run: allelotype.cwl
inputs:
- { id: bam, source: "#samindex/bam_with_bai" }
- { id: reference, source: "#reference" }
- { id: output_prefix, source: "#output_prefix" }
- { id: noise_model, source: "#noise_model" }
- { id: strinfo, source: "#strinfo" }
outputs:
- { id: vcf }
- { id: vcf_stats }