
CALYPR: Computation Analysis and Learning Yielding Predictions and Reports
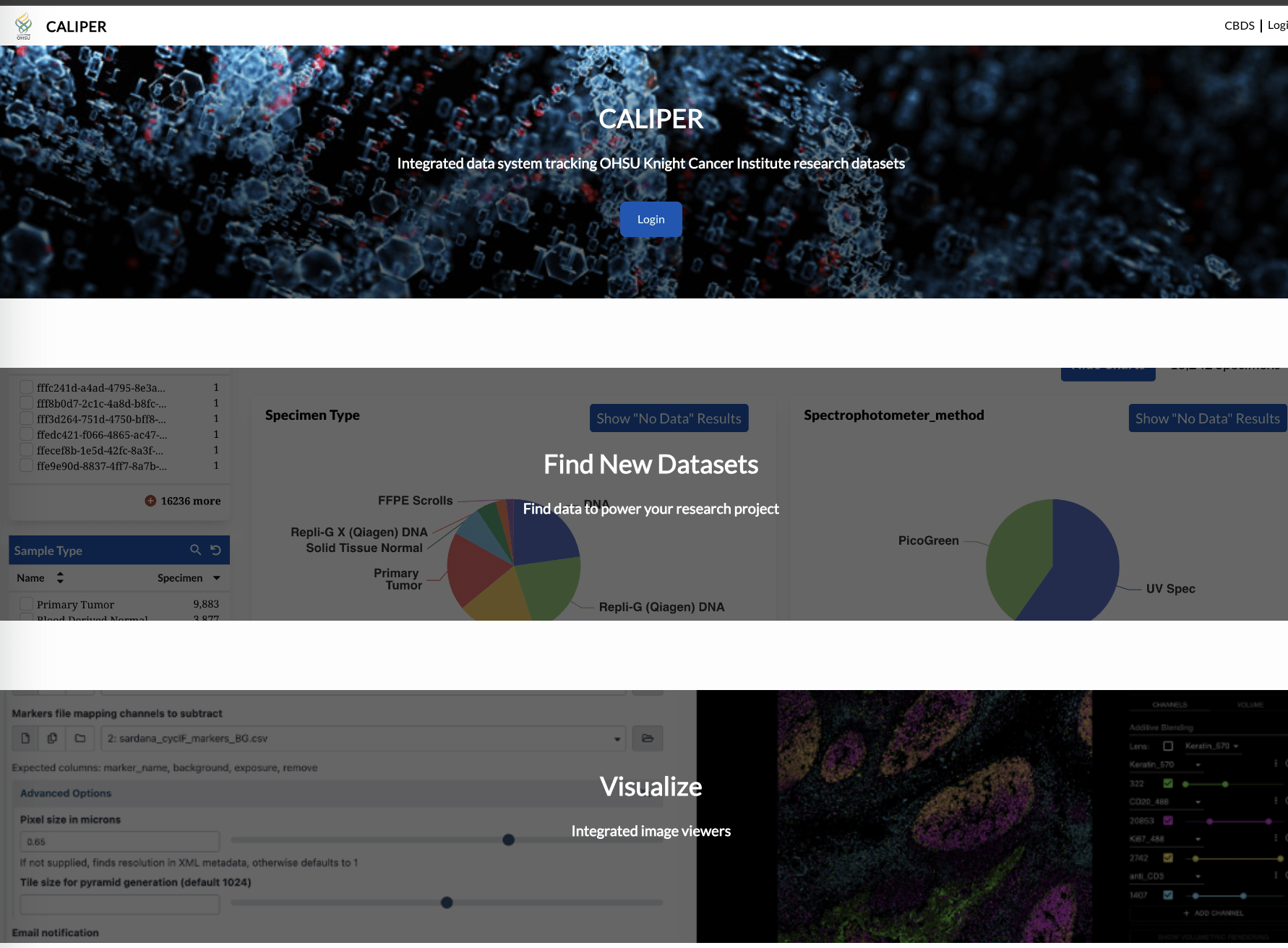
Challenge: Data Silos + Duplications + Interoperability
Data can become fragmented locked in silos, duplicated unnecessarily, and difficult to integrate.

. . .
lab A
lab Z

Data Lake
UI interface
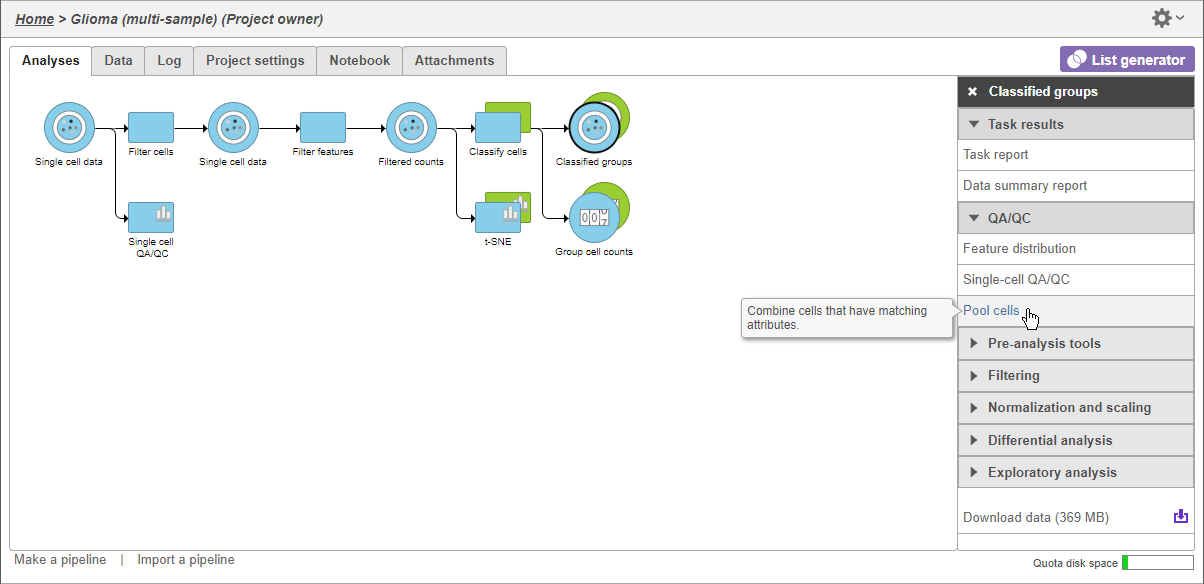
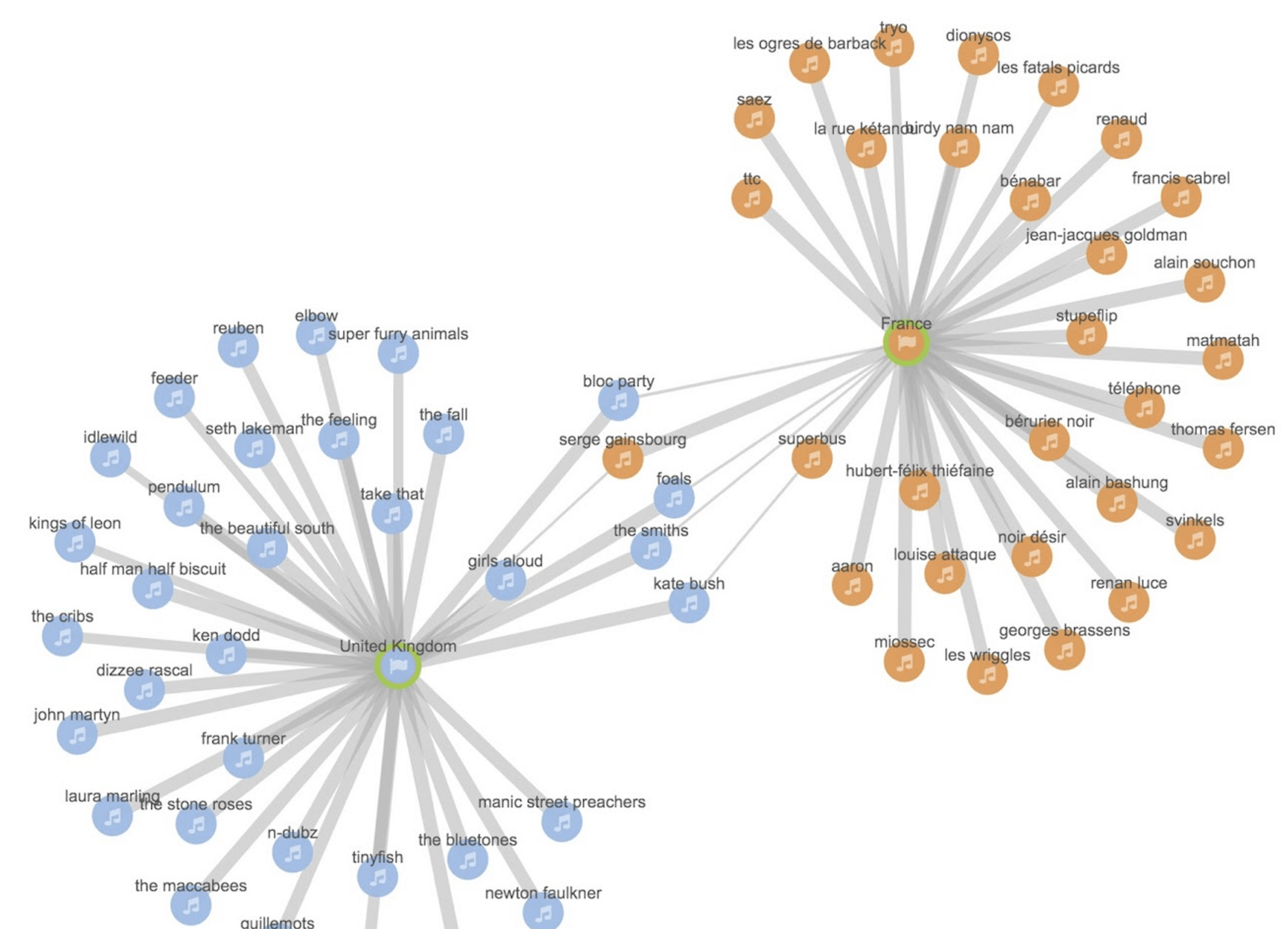


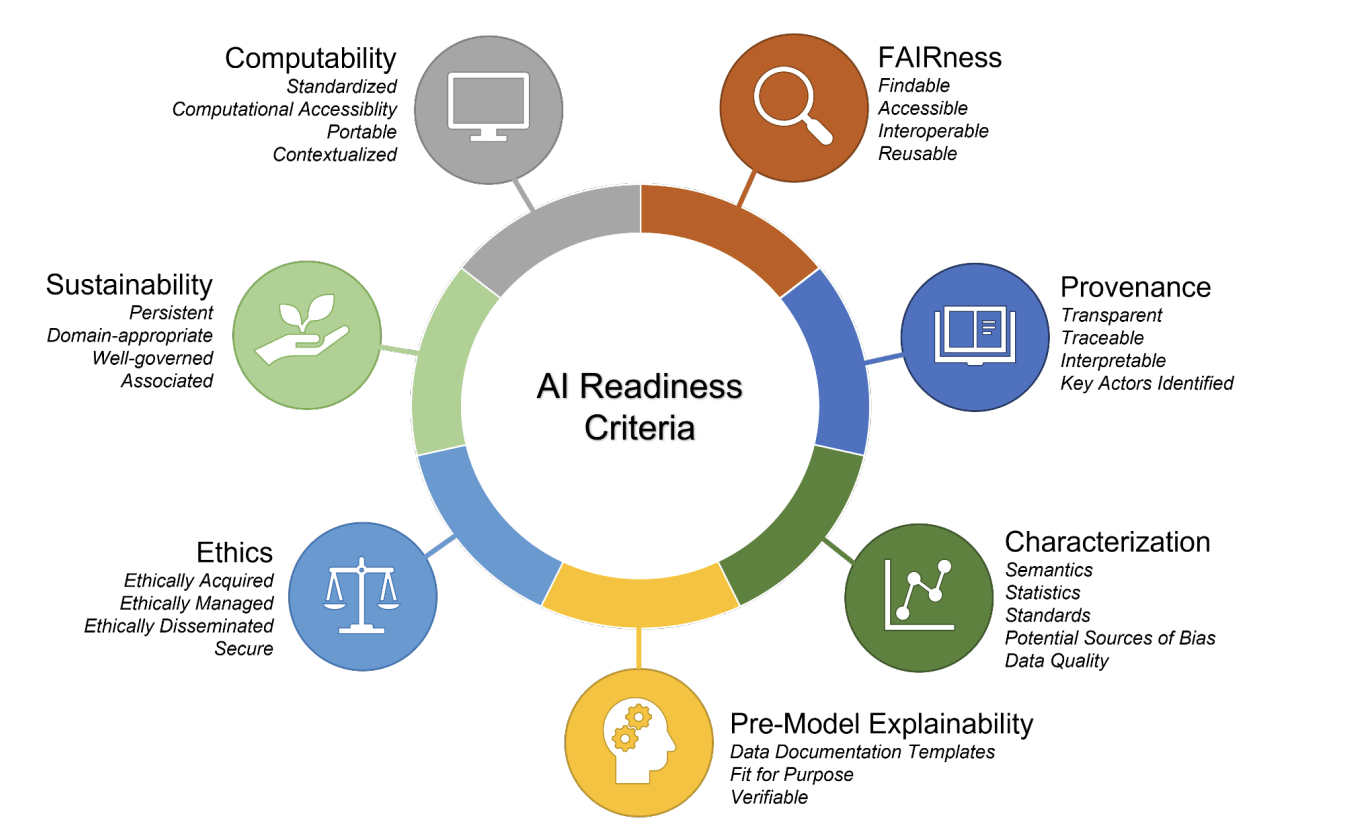
Clark et al. AI-readiness for Biomedical Data Bridge2AI Recommendations
CALYPR: Originally launched as International Data Exchange and Analysis Platform (ACED‑IDP )
A multi‑year, multi‑institution & cross‑department collaboration






Management
ETL
Standards



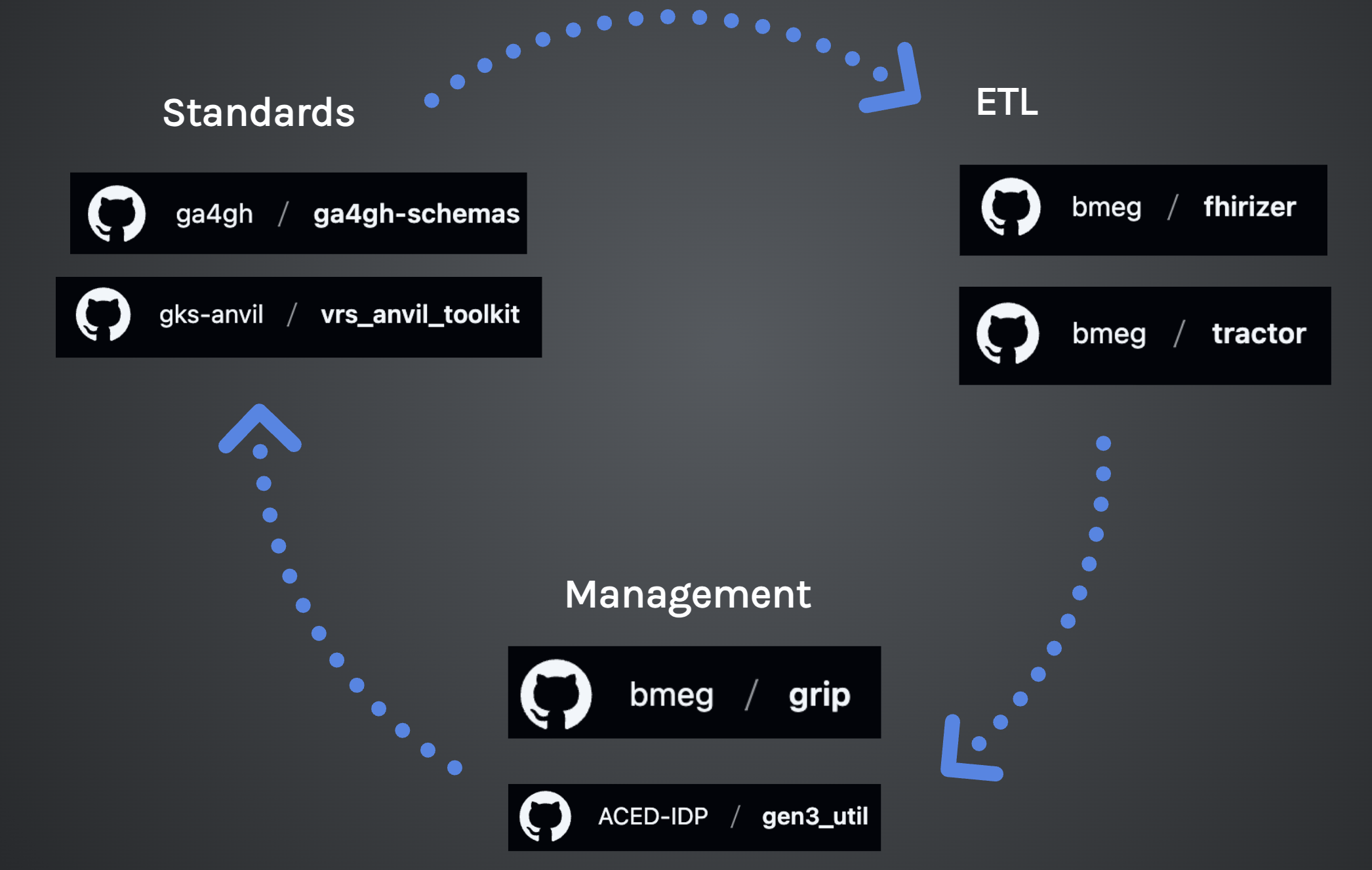
Data Science Garage: Data Standards + ETL + management
CALYPR : Extending Gen3 with Microservices for Bioinformatics Data Storage, Querying & Management



Modular Microservices
- Gen3 User: Build cohort + download
- Gen3 Developer: Build + integrate microservices
- Gen3 operator: Spinup an instance + configure data + customize frontend
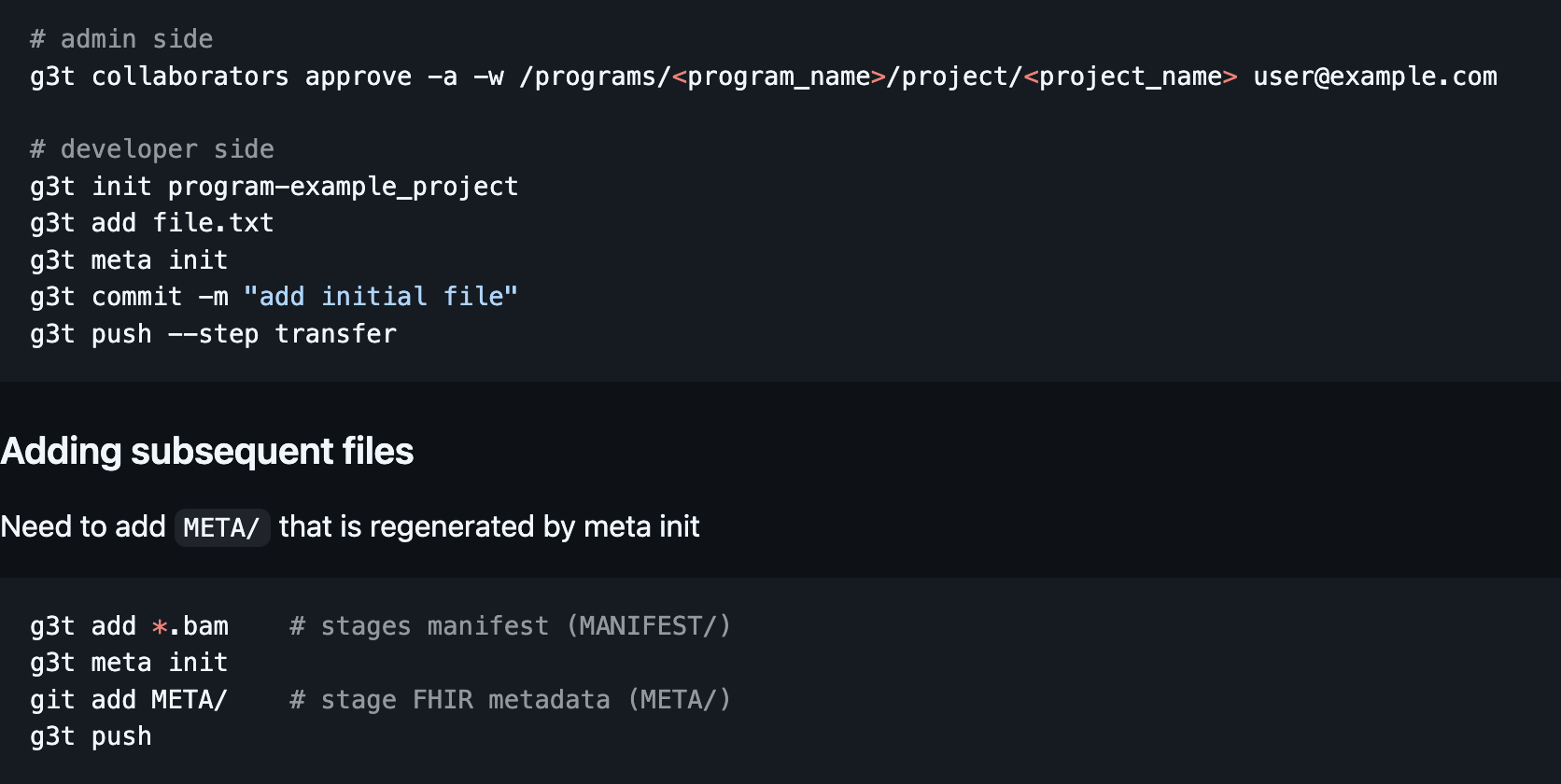
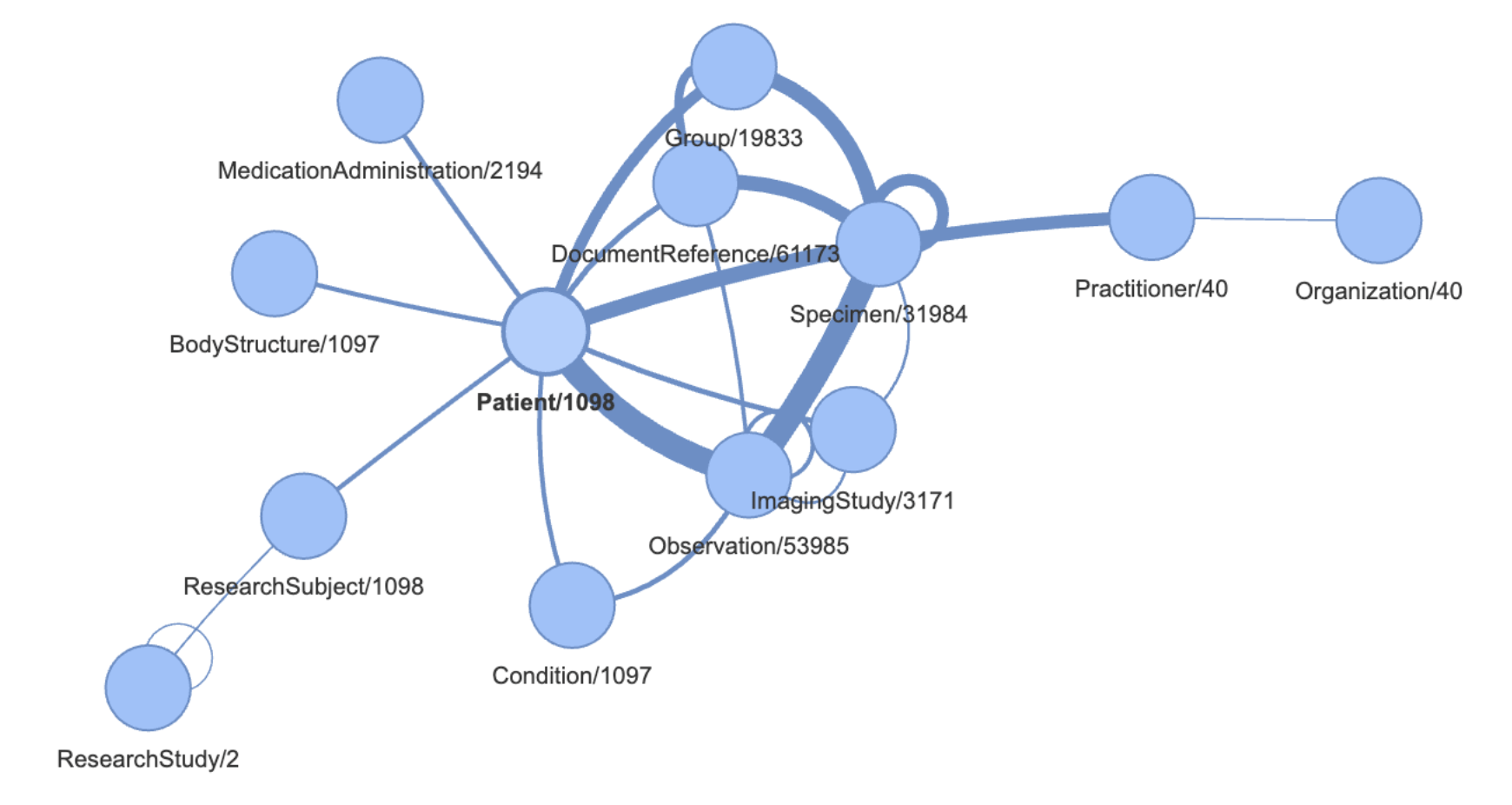
BMEG Iceberg + FHIR Graph Schema - A FAIR framework for capturing study design questions:
Which specimens in a pancreatic cancer cohort have an EGFR mutation status?

Gene
Allele
Protein
Transcript
Pathway


CALYPR: Data Ingestion + Integration
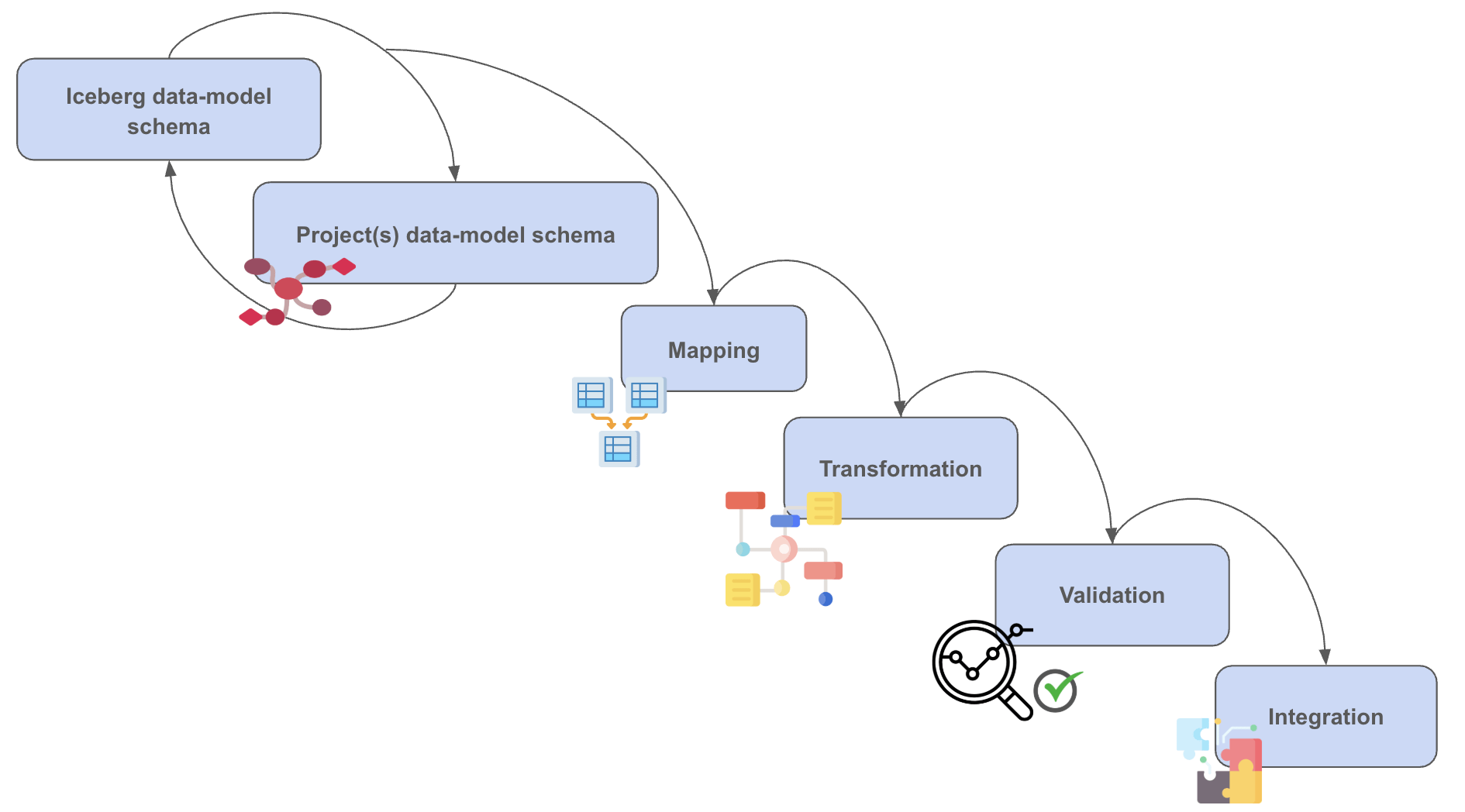






CALYPR: Landing & Main App Page

- Data Explorer
- Storage Manager
- Custom Report Pages
- Data Viewer
- Interactive Computation
- Graph Queries
- Project Management
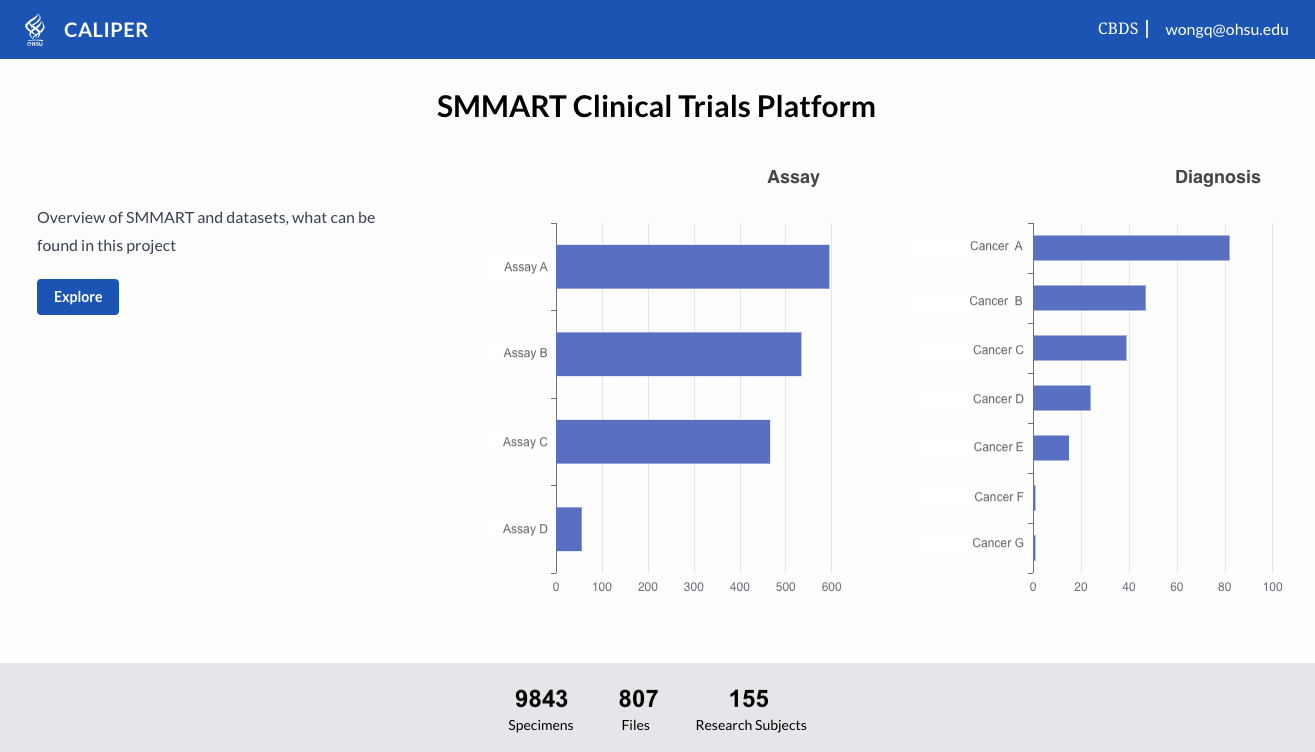
CALYPR: Discovery & Summary Reports

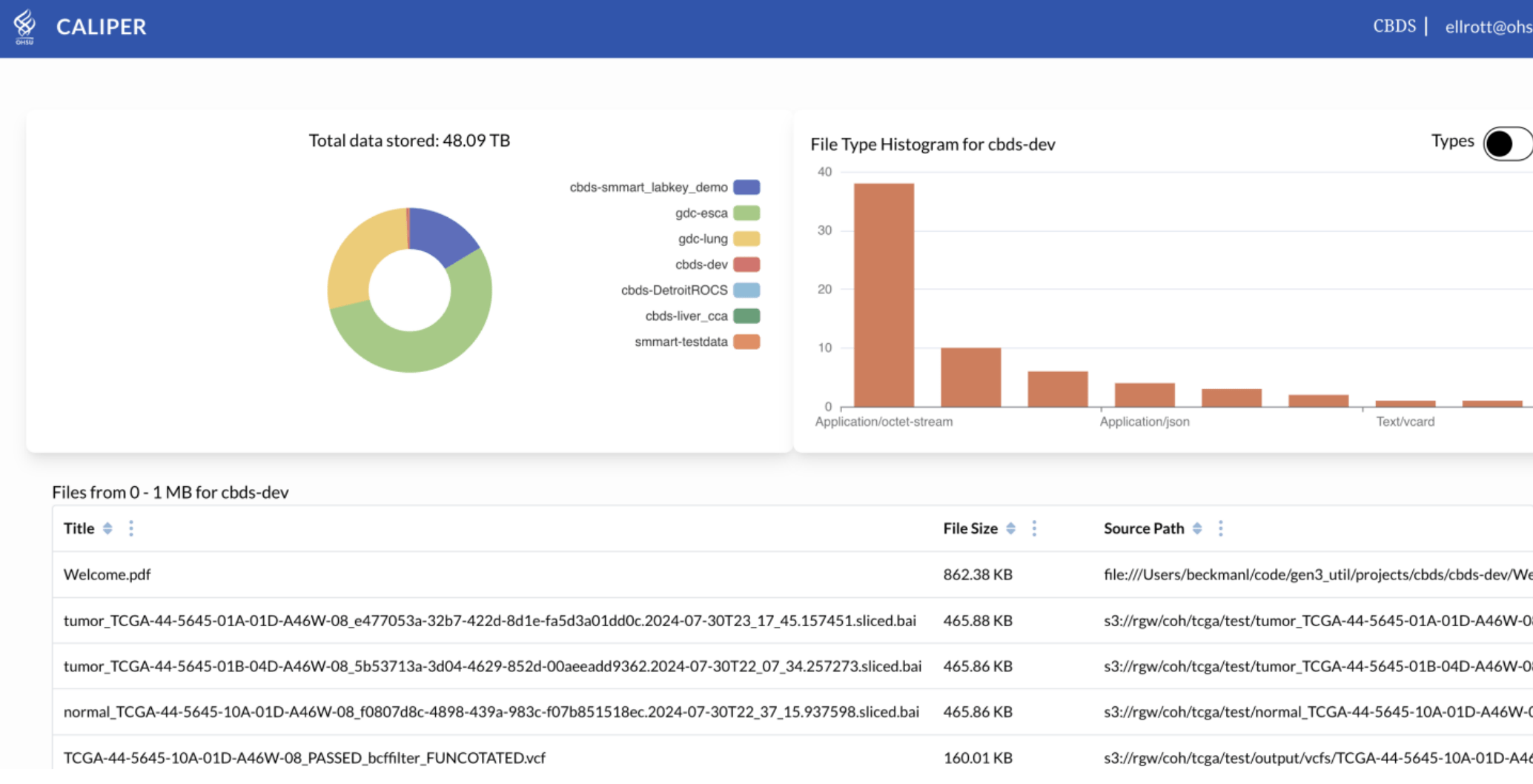
CALYPR: File Viewer

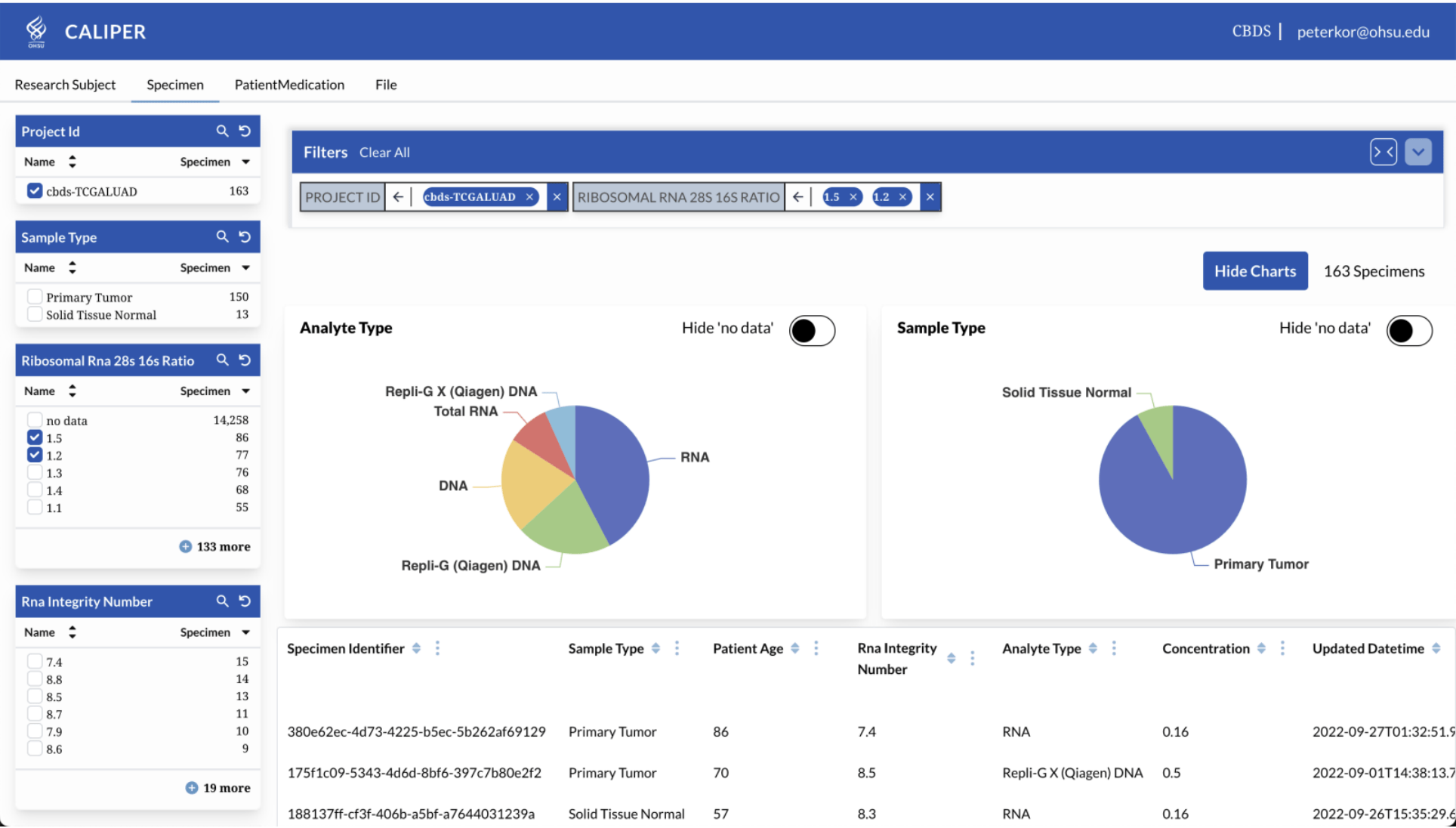
CALYPR: Data Explorer - Cohort Builder

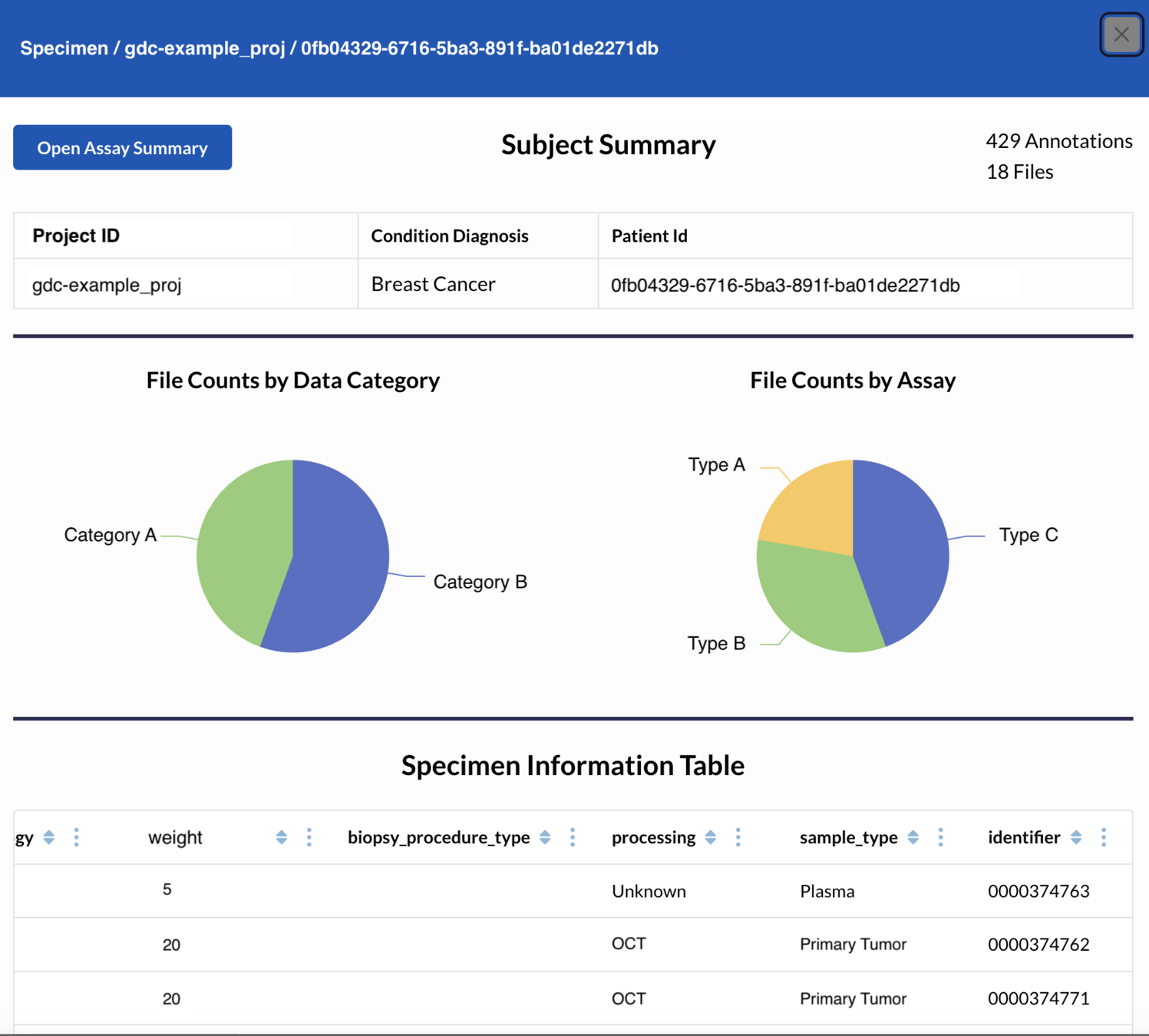
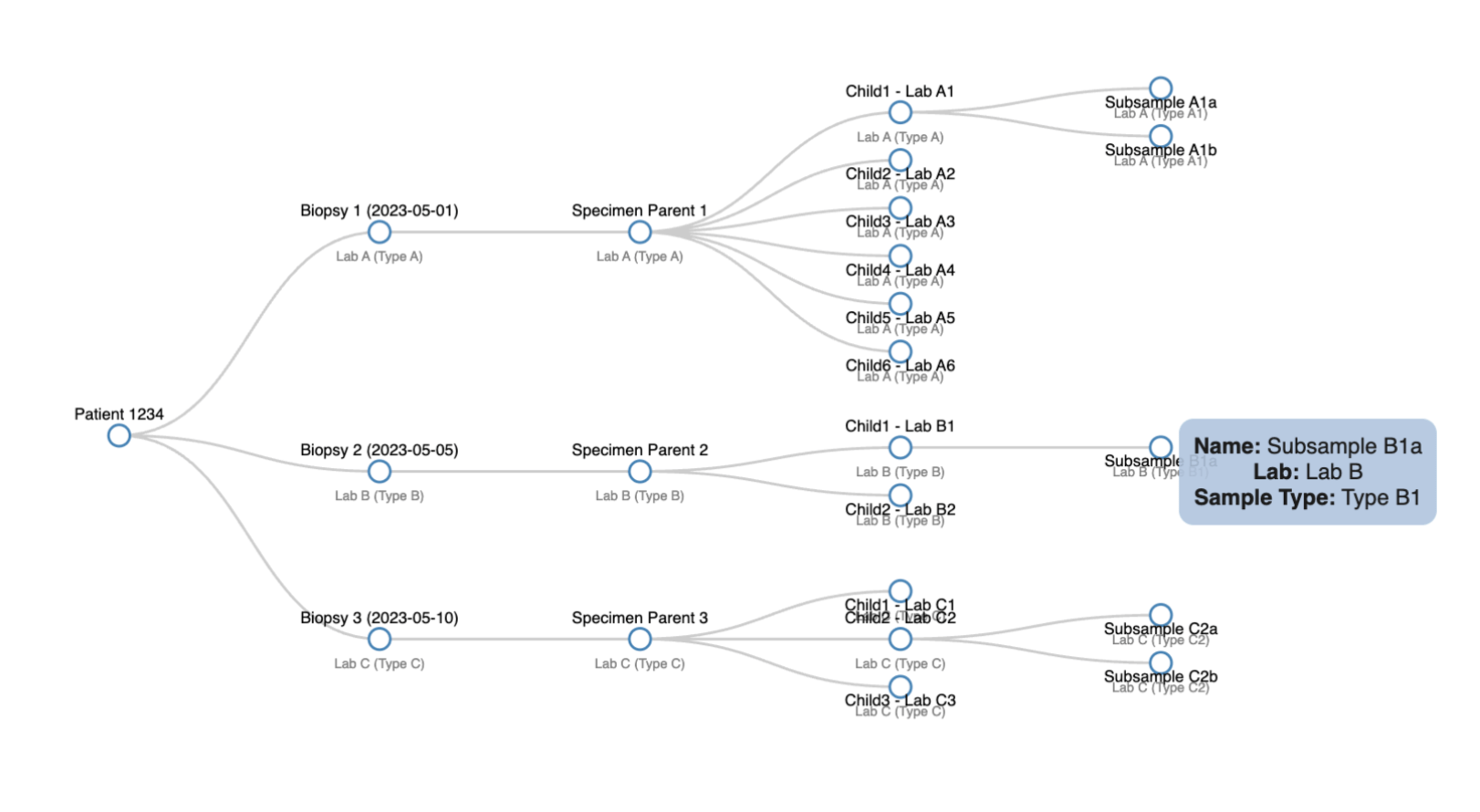
CALYPR: Data Explorer - Temporal Data Tracking
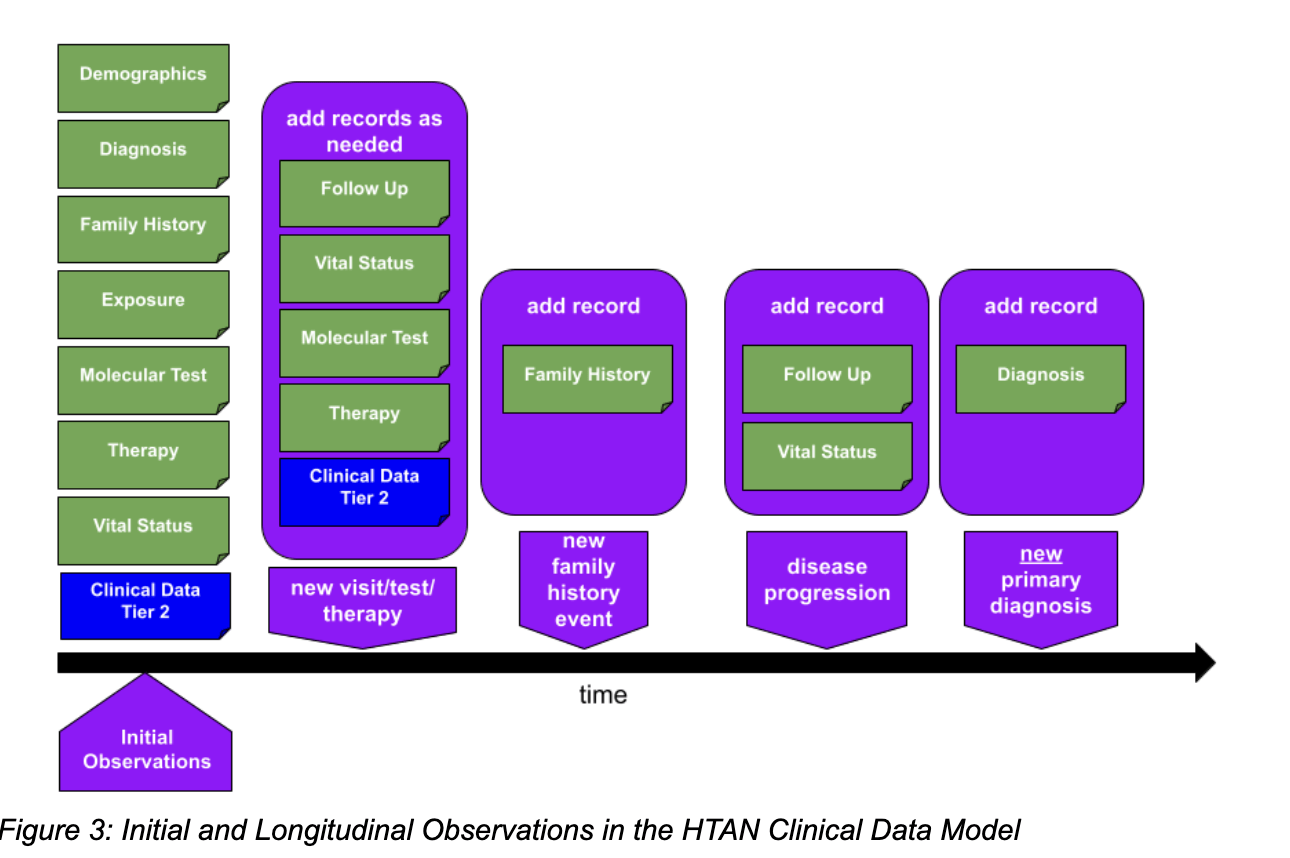
CALYPR: Image Viewer

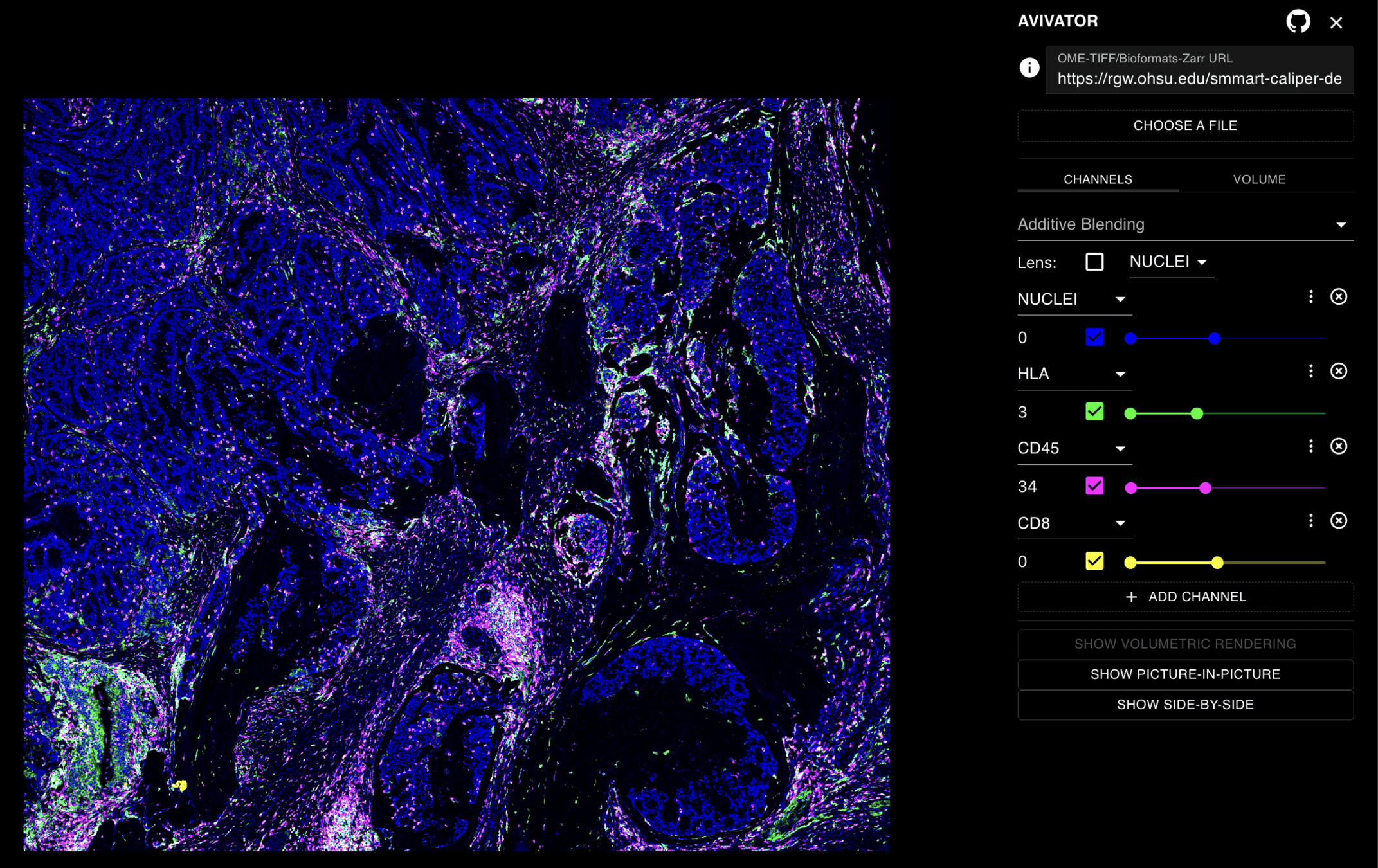
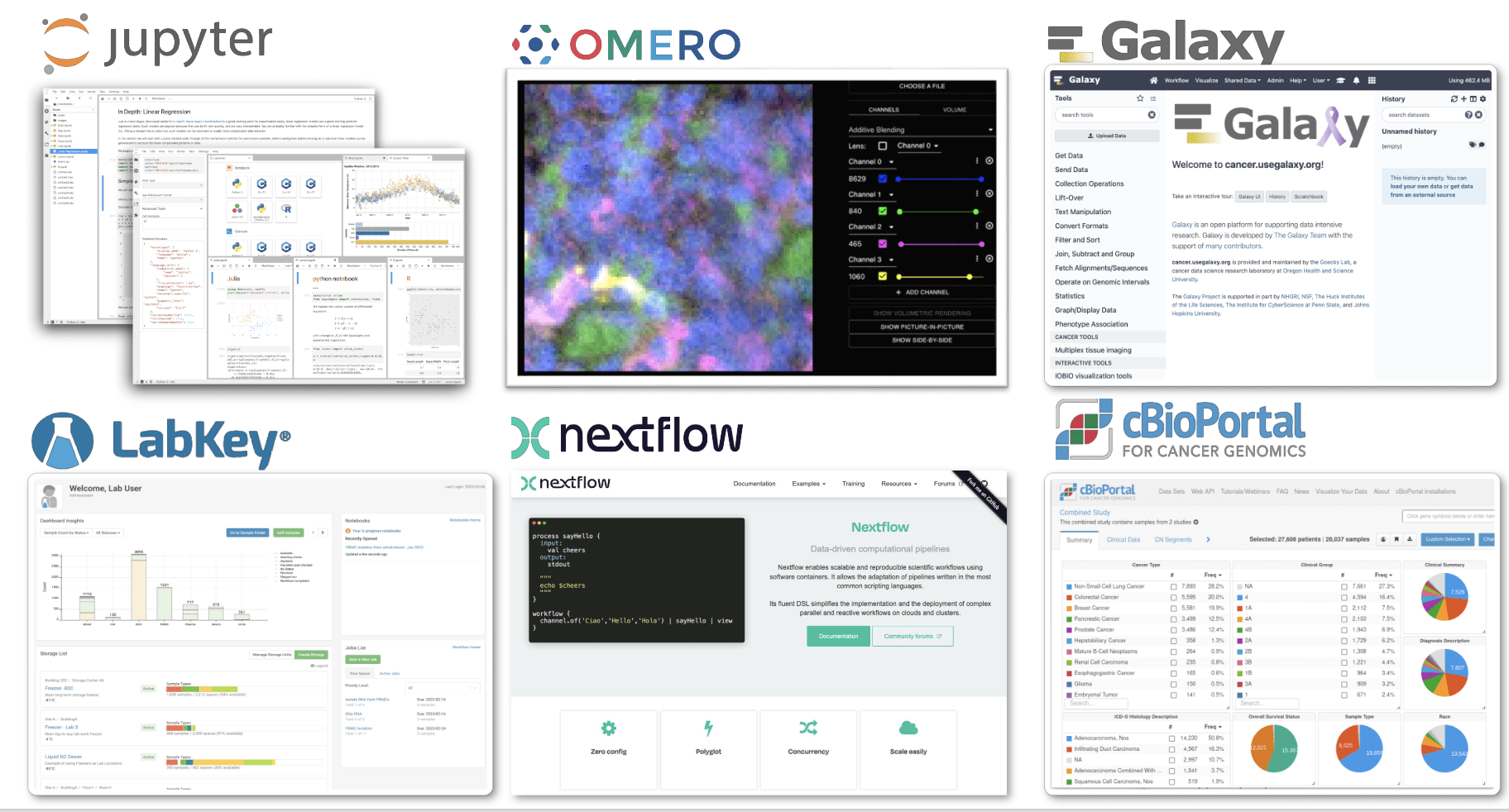
CALYPR: Connecting Applications - Future Direction
Nasim Sanati
Computational Biologist
@teslajoy
Quinn Wai Wong
Research Software Eng.
@quinnwai
Kyle Ellrott
PI + Associate Professor
@kellrott
Liam Beckman
Research Software Eng.
@lbeckman314



Brian Walsh
Sr. Research Software Eng.
@bwalsh

Jordan Tagle
Computational Biologist
@jordan2lee
Matthew Peterkort
Research Software Eng.
@matthewpeterkort

Data Science Garage + Creason's Teams



Allison Creason
Sr. Computational Biologist
Jay Egger
Computational Biologist

Luke Sargent
Sr. Research Software Eng.

Code + Stunning Sunrise + Local Coffee + Great Team


Extra Slides
HTAN: Bidirectional Transformer between FHIR and HTAN schema.
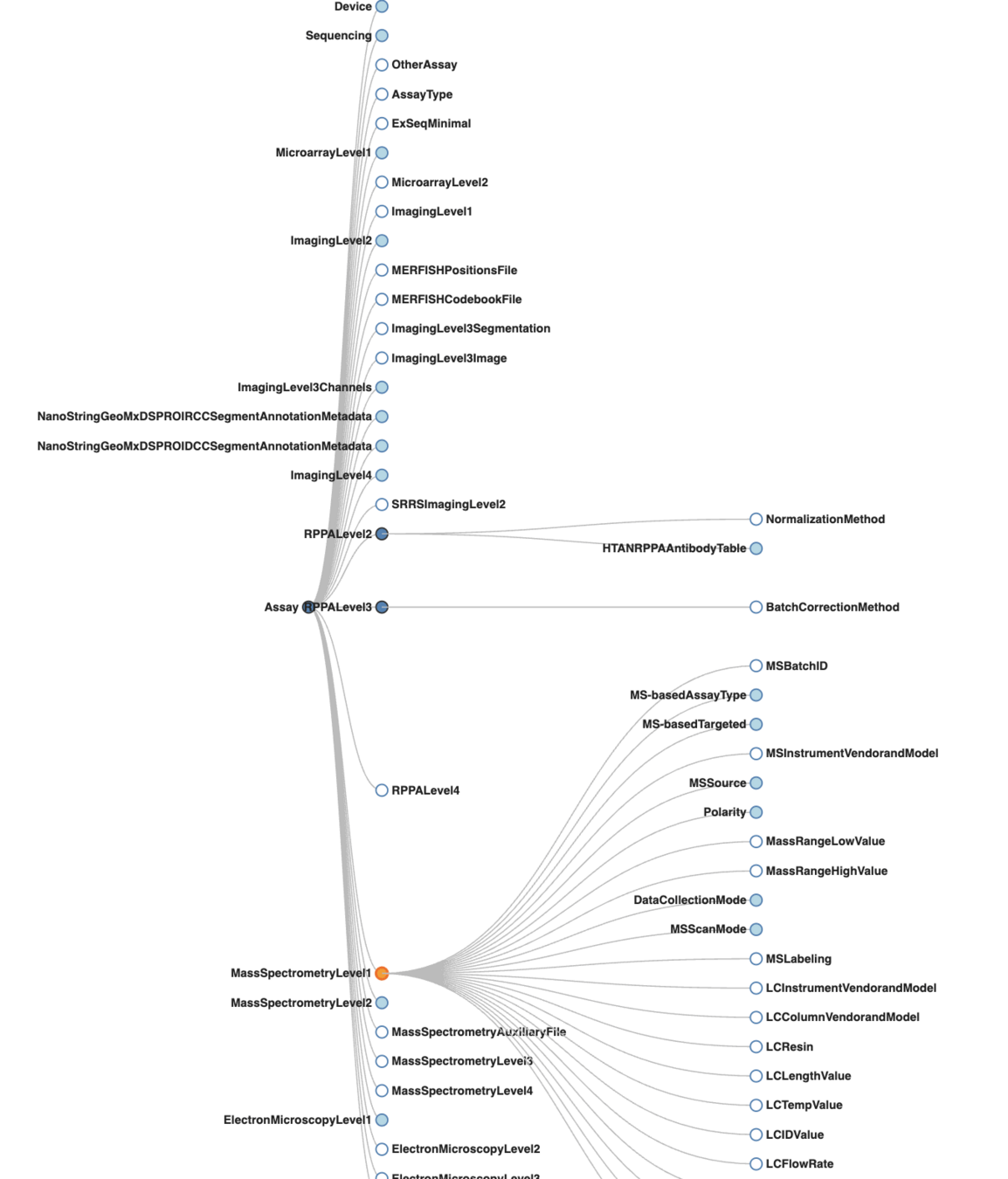
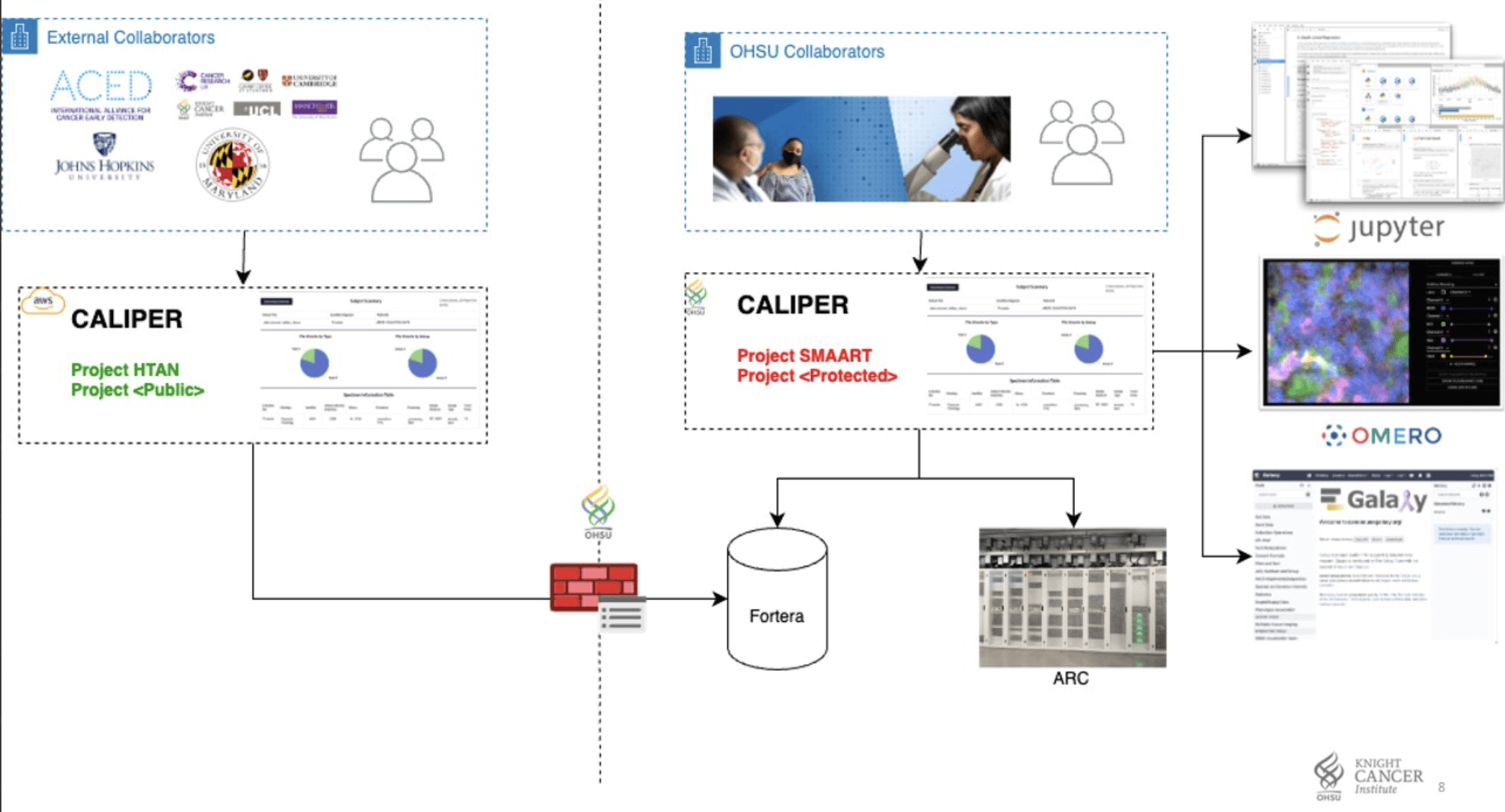




GRIP & FHIR
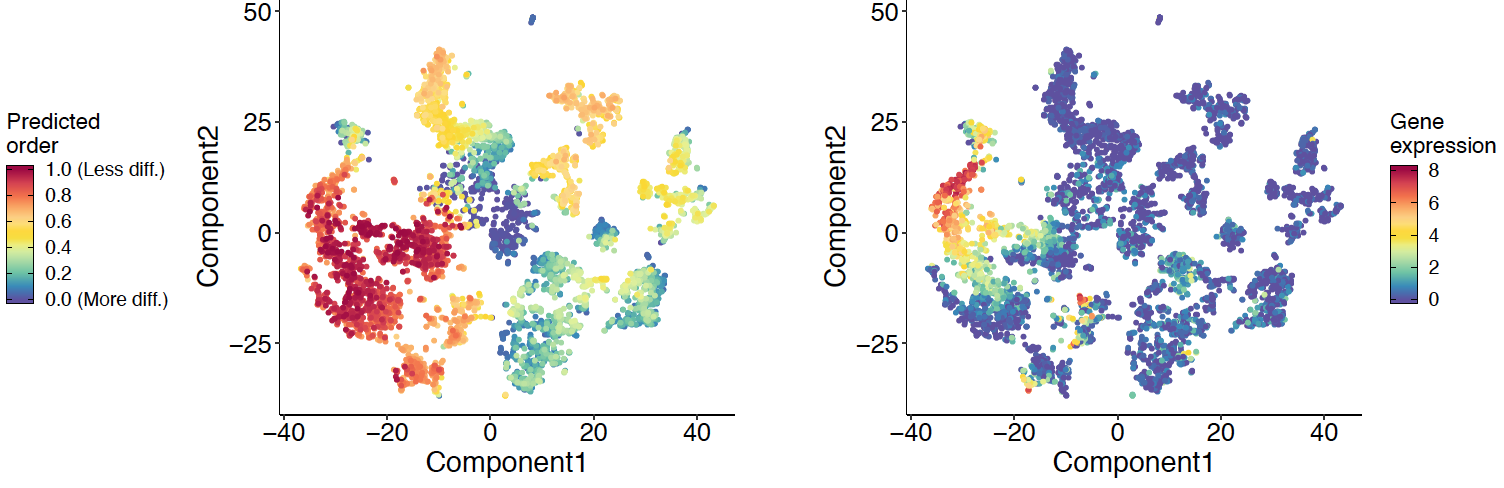
Querying TCGA BRCA Data with subtype labels for classification

Subtype Classification with Neural Networks
ML modeling and Shapley Feature Extraction

Pathway Enrichment Analysis
Extracting Biological Insights from Gene Set Results

Where We Work
Knight Cancer Research Building @ South Waterfront



Across from OMSI and right by Tilikum Crossing

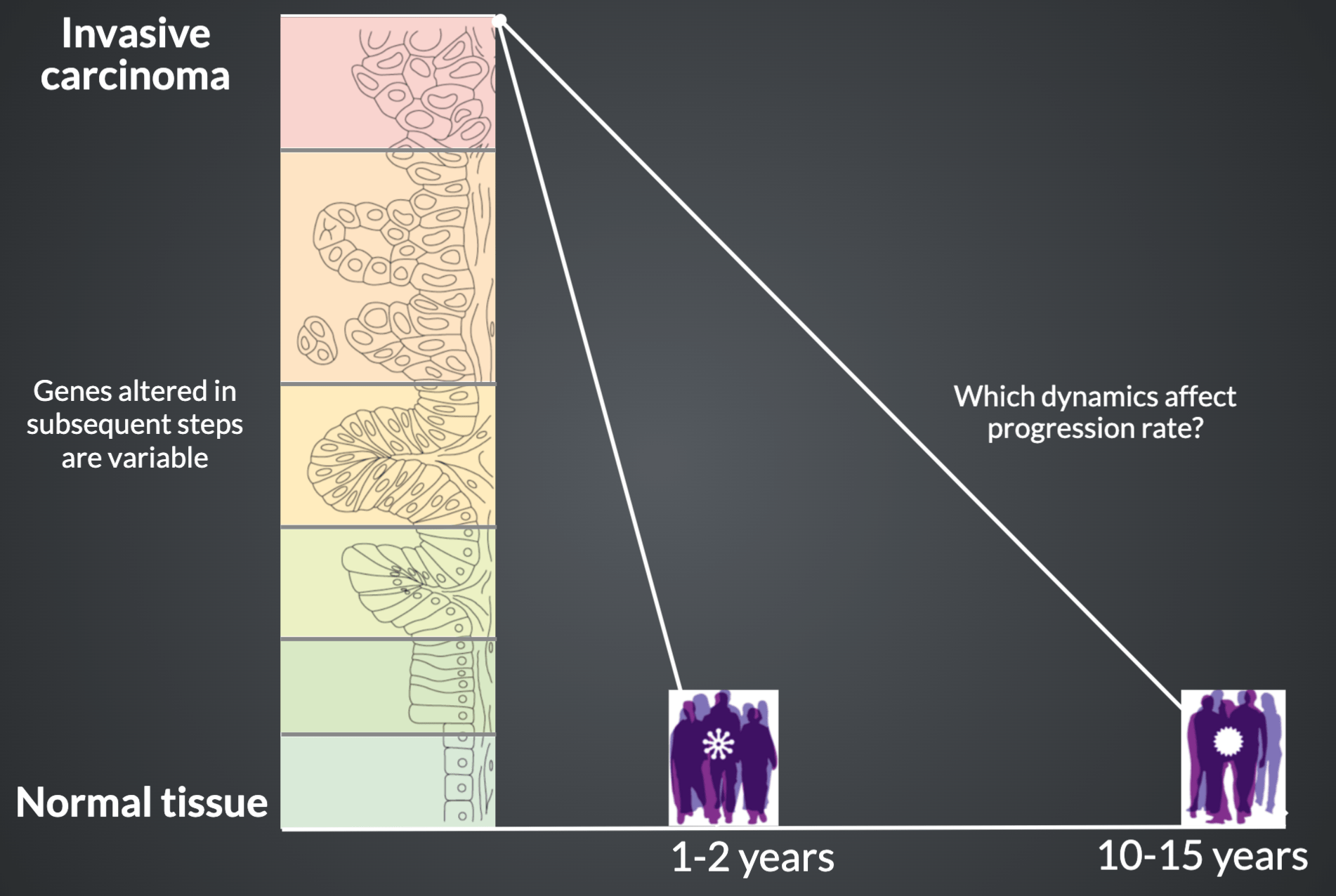
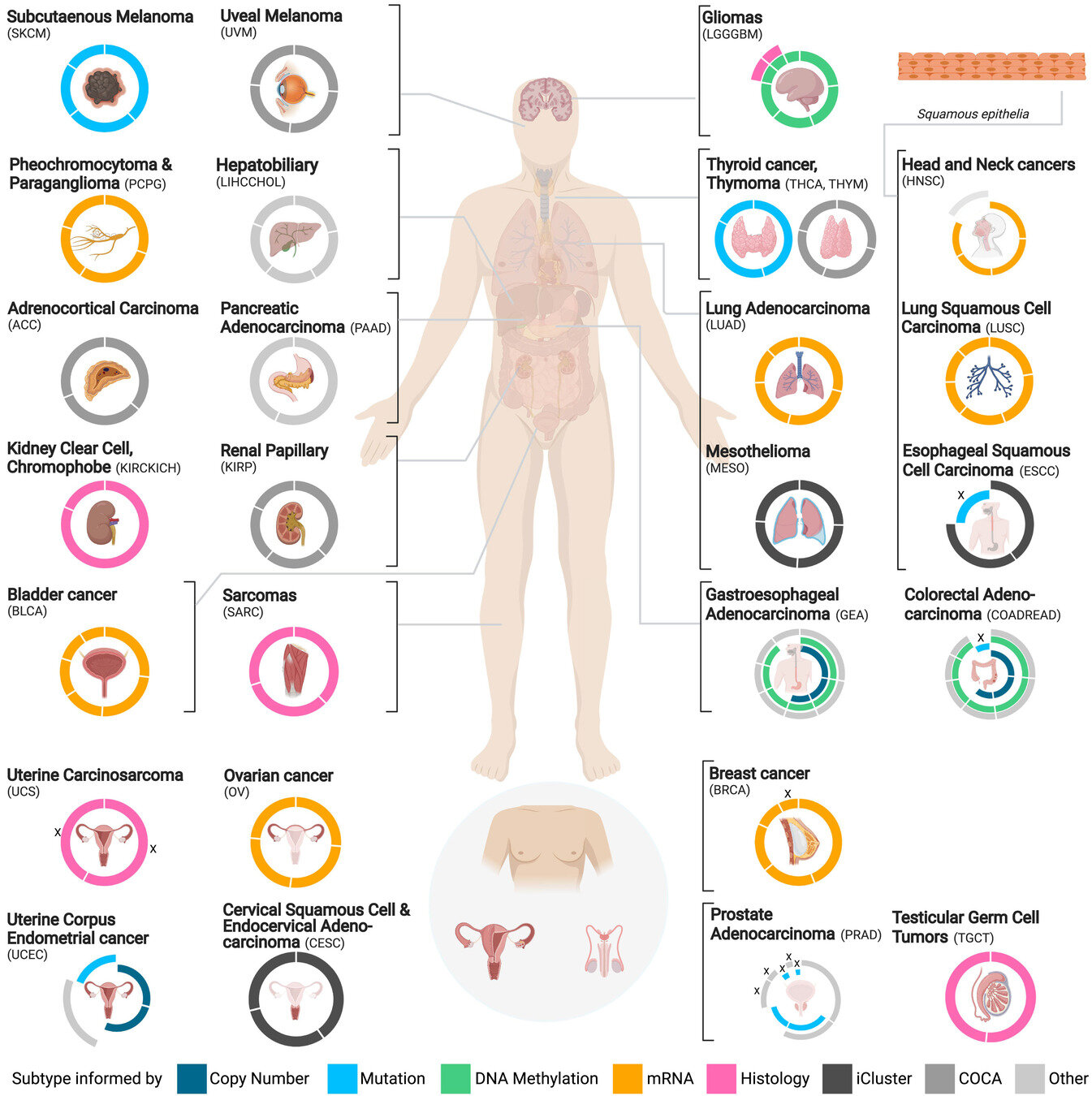
Scientific Problem
How can we find actionable insights in a disease as complex as cancer?


Scientific Problem
How do we integrate data to extract meaningful insights?
-
ETL (Extract, Transform, Load)
-
FHIRizer – Transforms GDC, Cellosaurus, HTAN data into FHIR graph representation.
-
Snakemake & Airflow – Automate and orchestrate ETL workflows.
-
Bringing It All Together with Python
Data Standards, ETL, and Management
-
Management
GRaph Integration Platform (GRIP) - a powerful graph query engine designed to span across multiple database backends.
Gen3 – Data commons for storage, querying, and management.

-
Standards
-
Pydantic – Validates and enforces FHIR schema.
-
GA4GH (Global Alliance for Genomics & Health) – Establishes global standards for genomic data interoperability.
-
Pathway Databases (Pathway Commons, Reactome) – Structure & standardize biological context through molecular interaction graphs.
-




Data Science Garage team
Bringing it all together with data standards + extract transform load (ETL) + management
Management
ETL
Standards



Calypr: Data Explorer - Cohort Builder
Data can become fragmented locked in silos, duplicated unnecessarily, and difficult to integrate.




