Omnibenchmark:
Challenges and opportunities of open and continuous community benchmarking
DMLS seminar,
Almut Lütge - Robinson group,
Zürich, 18.10.2022


benchmark:
Systematic comparison of methods/processes to understand underlying features and/or find the most suitable procedure for a specific task
single cell tools are on the rise
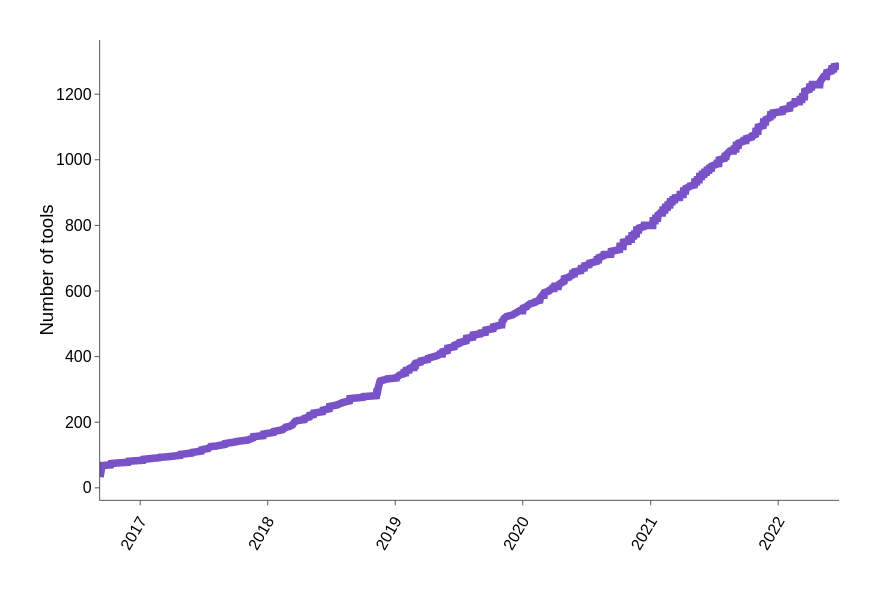
Number of tools per year
https://www.scrna-tools.org/
Comparisons can be ambiguous
Luecken et al., 2020
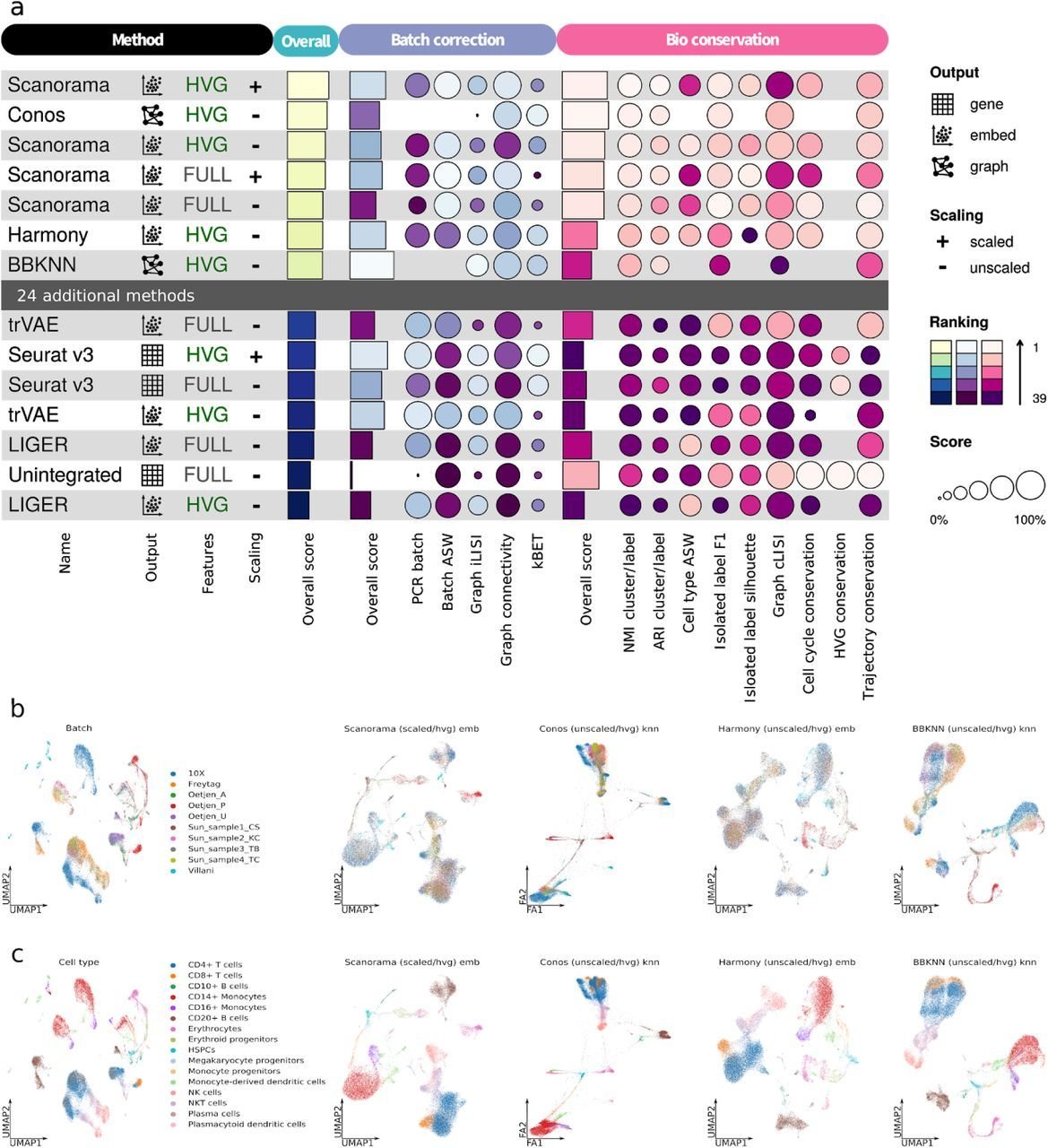
Metrics benchmark:
Systematic comparison of metrics to understand their performance and find the most suitable metric to evaluate batch correction
Task 1: Scaling and detection limits
Aim: Test whether metrics scale with (synthetic) batch strength; Estimate lower limit of batch detection
Spearman correlation of metrics with the batch logFC in simulation series on the same dataset; Minimal batch logFC that is recognized from the metrics as batch effect
Increasing batch log fold changes
Metrics have different ranges to detect and distinguish batch effects
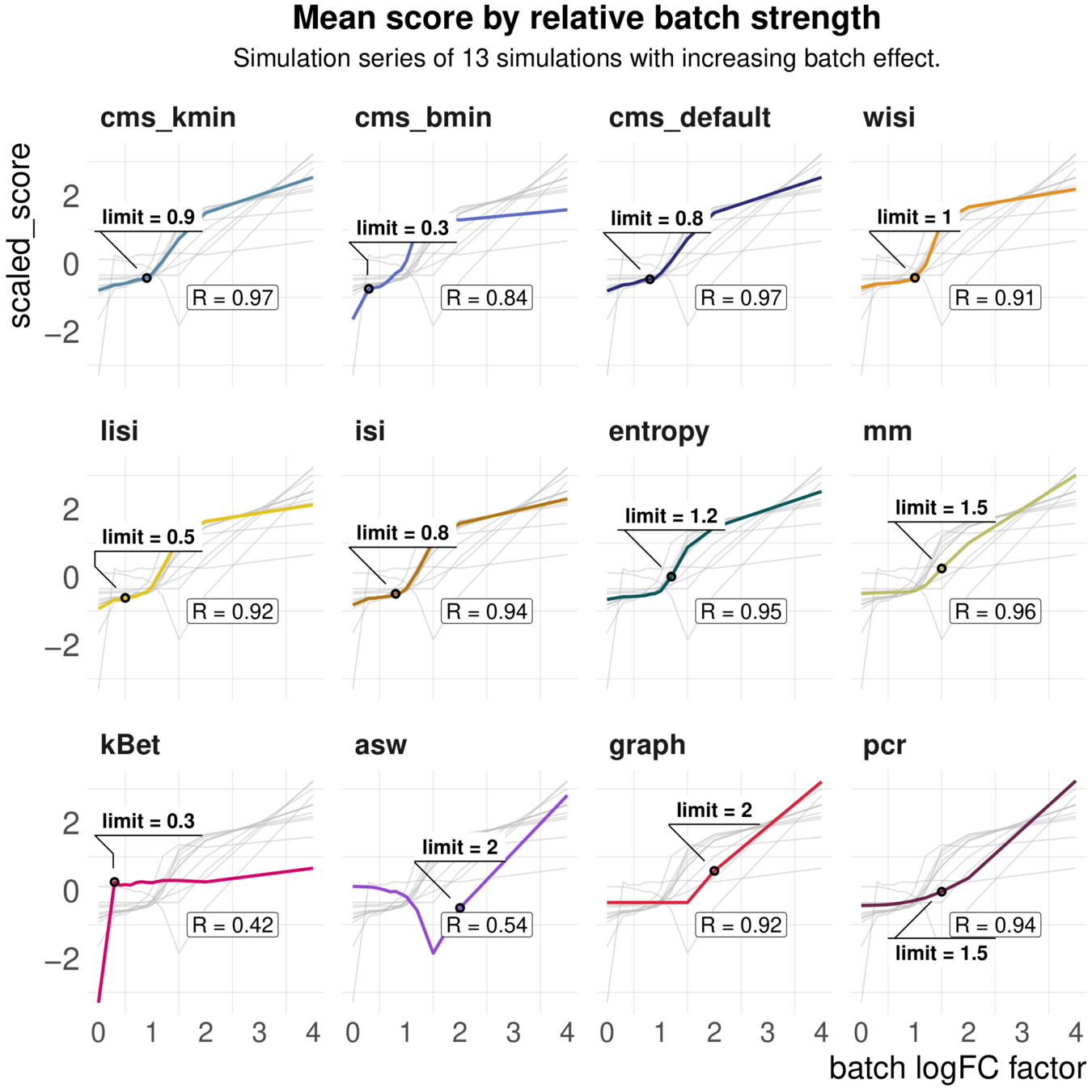
Metrics have different ranges to detect and distinguish batch effects
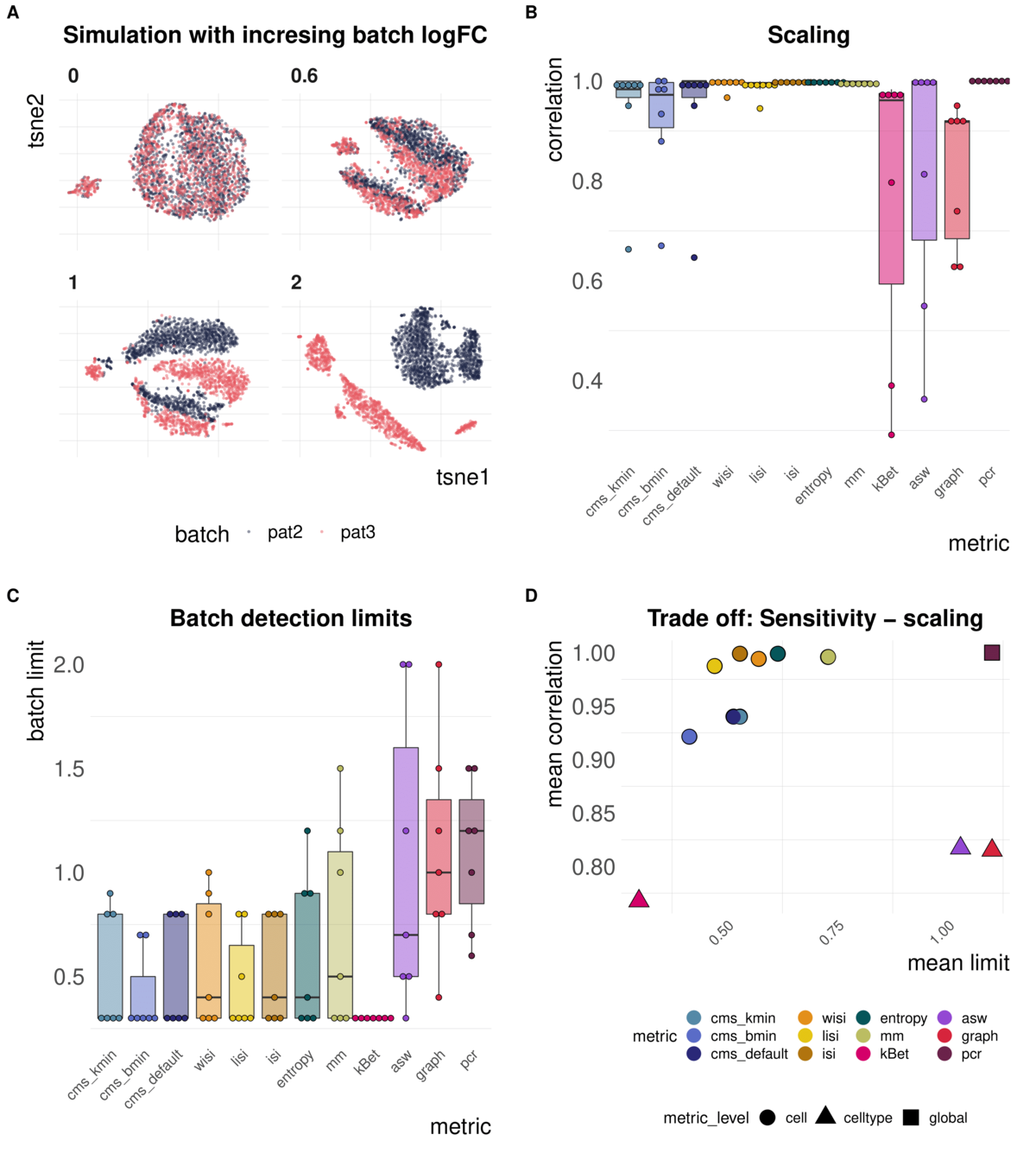

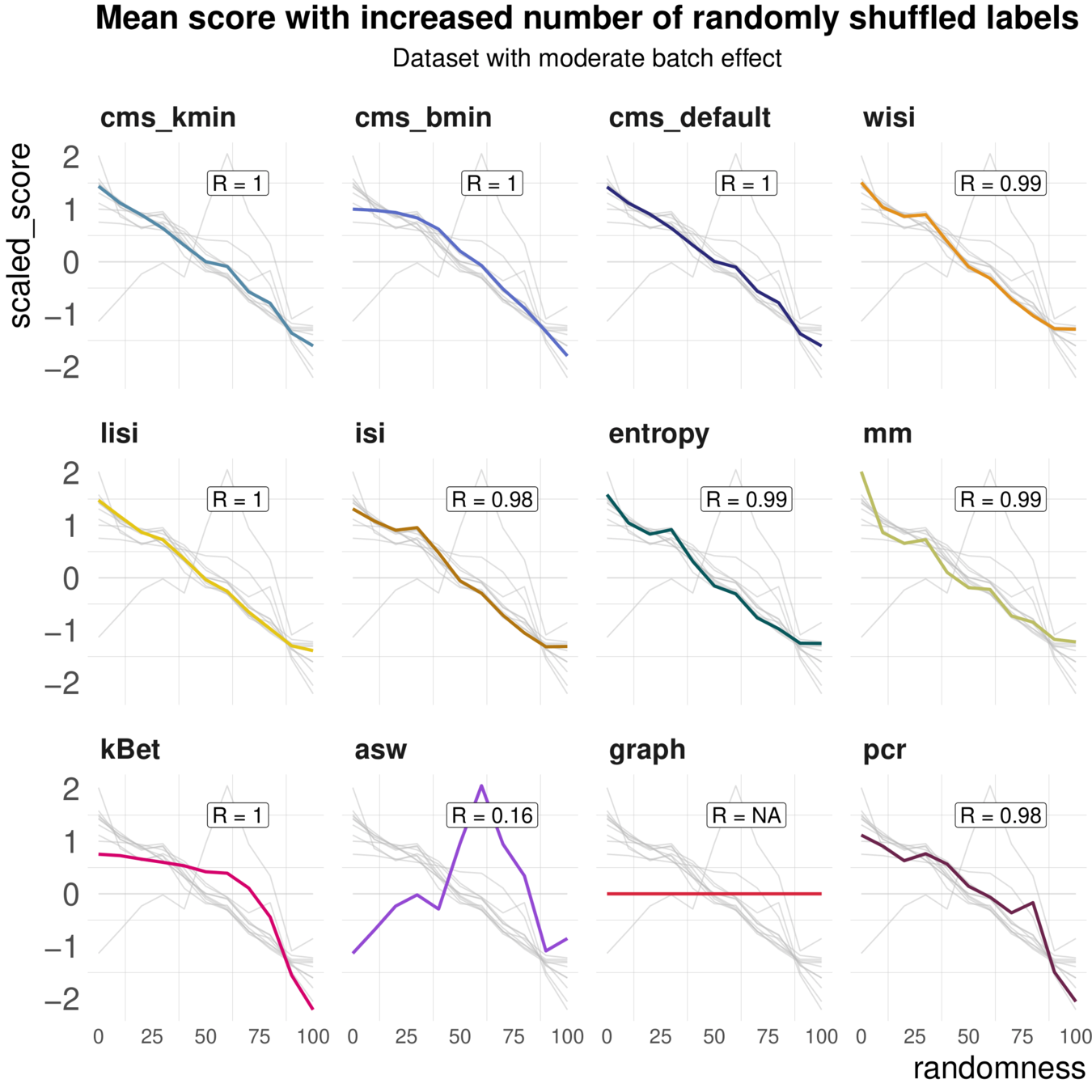
Task 2: Batch label permutation
Aim: Negative control and test whether metrics scale with randomness
Spearman correlation of metrics with the percentage of randomly permuted batch label
Increased percentage of Randomly permuted batch label
Most Metrics scale with label randomness
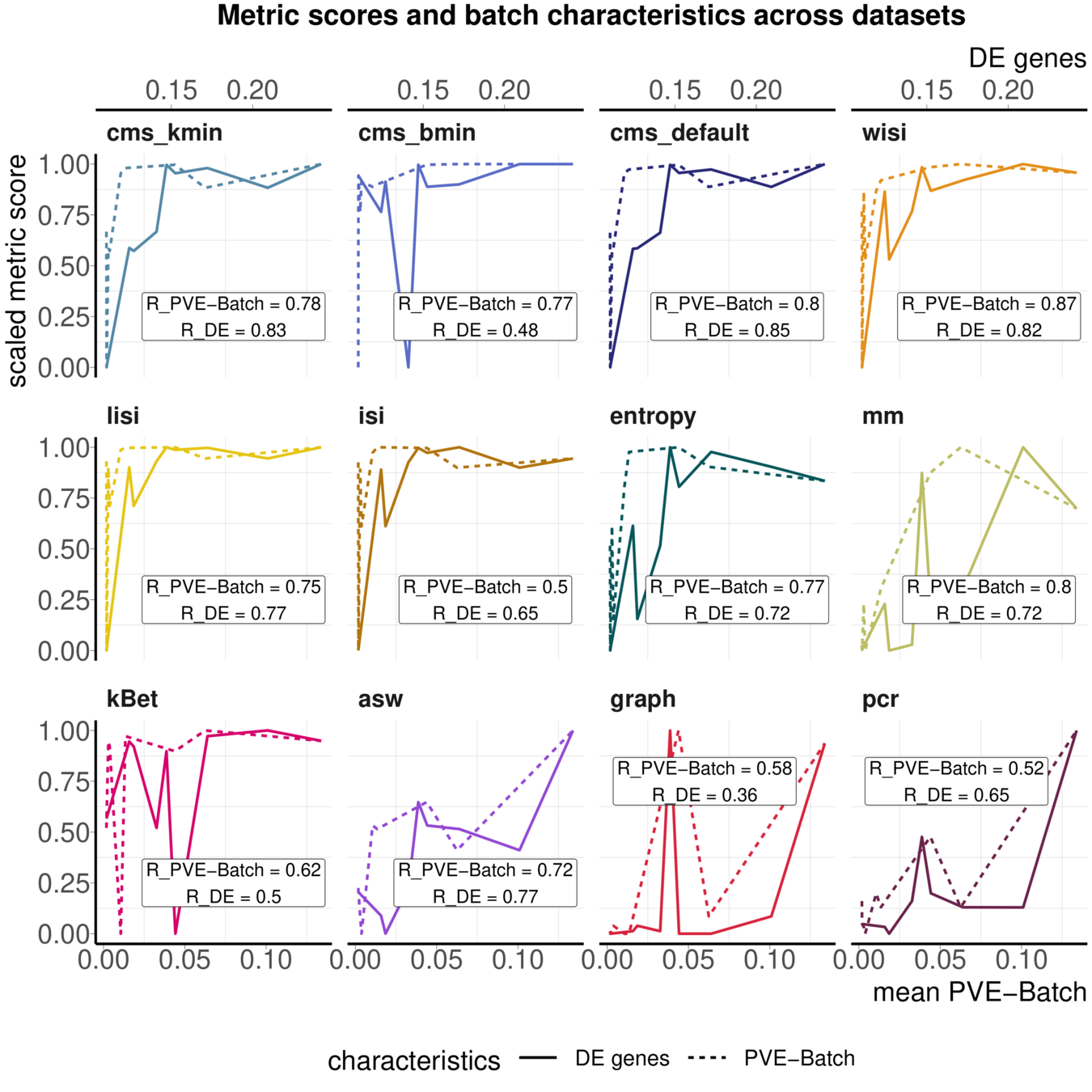
Task 3: Batch characteristics
Aim: Test whether metrics reflect batch strength across datasets
Spearman correlation of metrics with surrogates of batch strength (e.g., percent variance explained by batch (PVE-Batch) and proportion of DE genes between batches) across datasets
Percent variance explained by ..
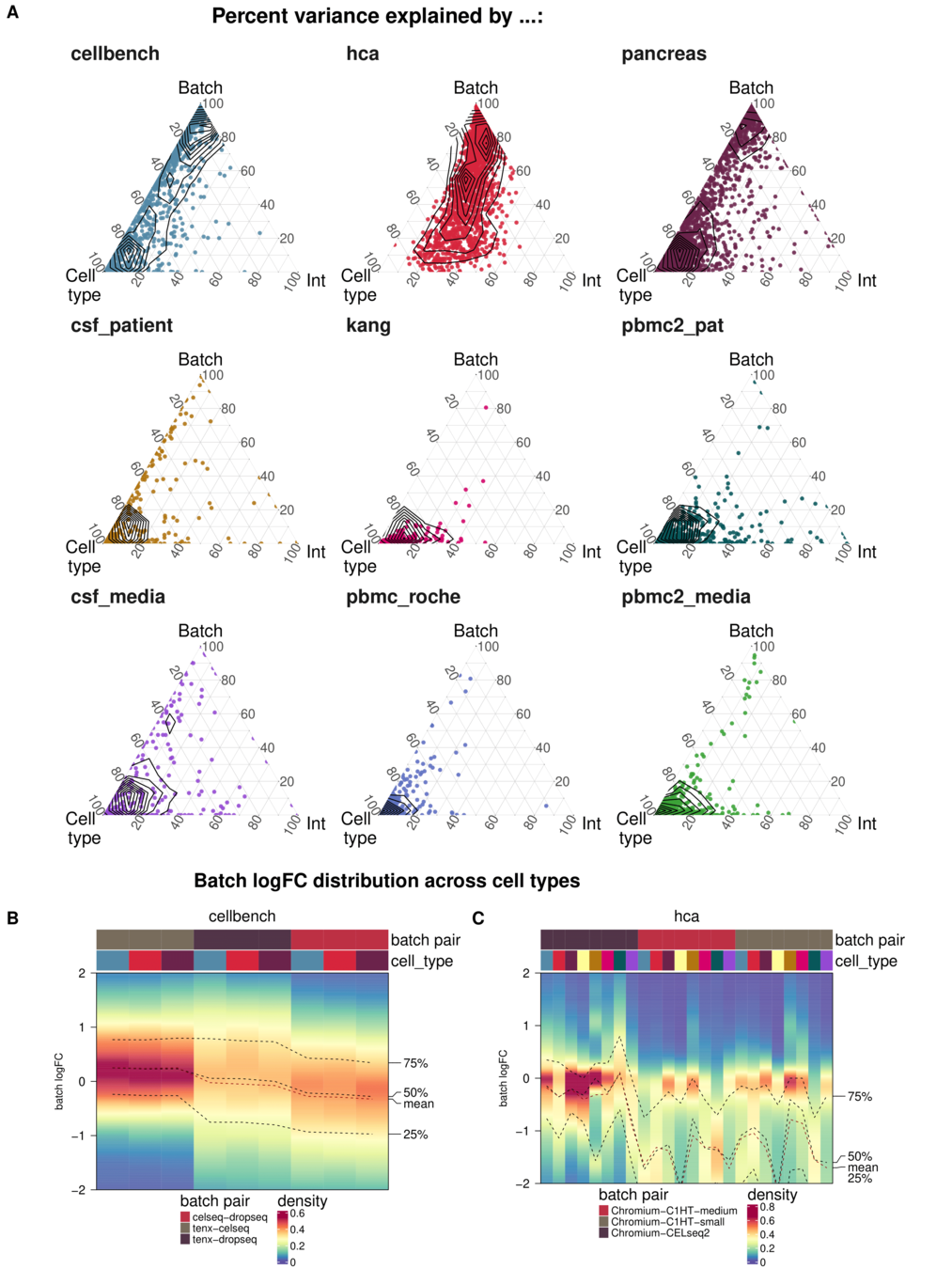

Percent variance explained by ..

Not all Metrics reflect batch characteristics
Task 4: Imbalanced batches
Aim: Reaction of metrics to imbalanced cell type abundance within the same dataset
Test sensitivity towards imbalance of cell type abundance
Increased Imbalance of the batch effect
Imbalanced batch effects affect most local metrics scores
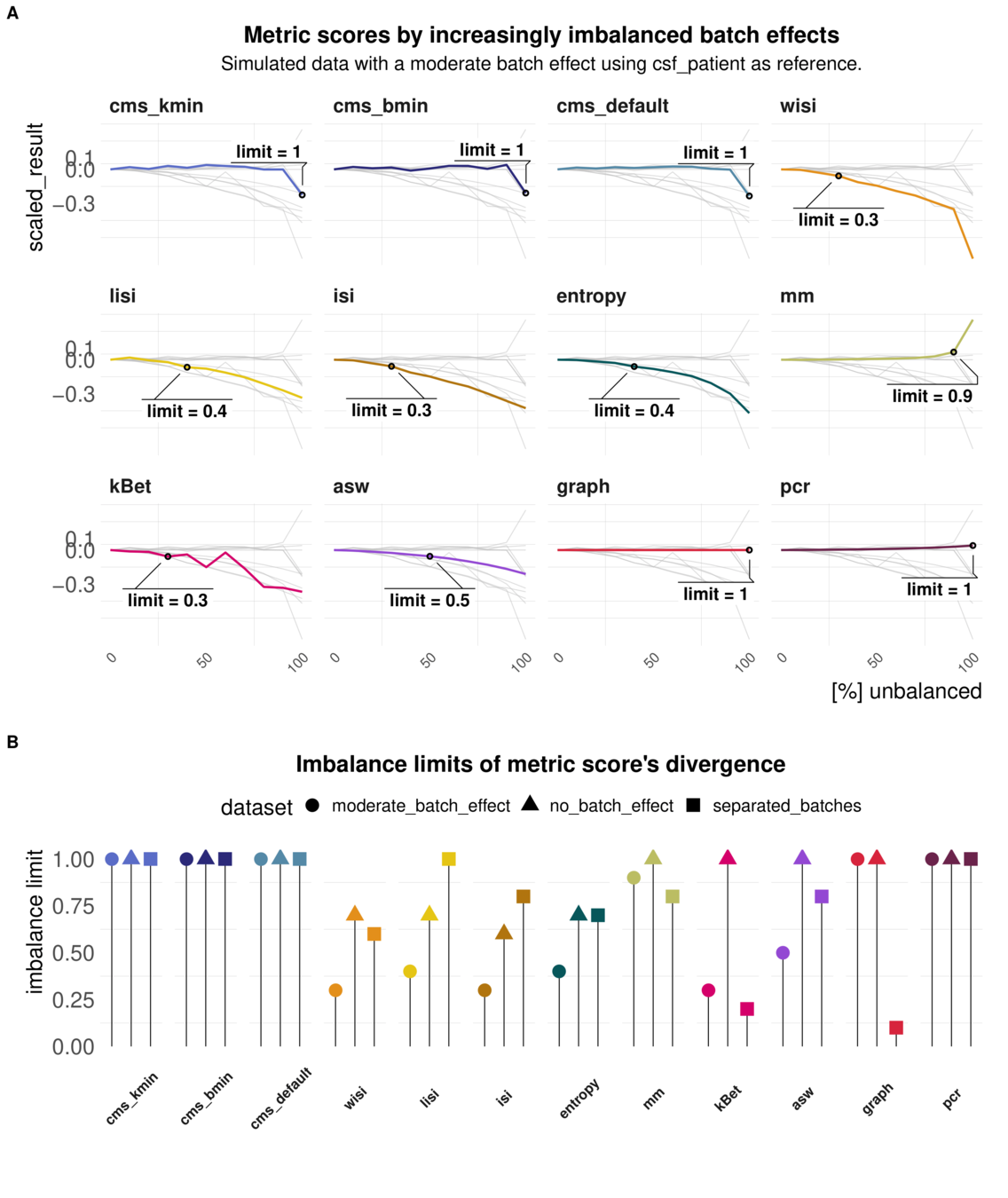
imbalance limits of different metrics

summary Metrics benchmark:
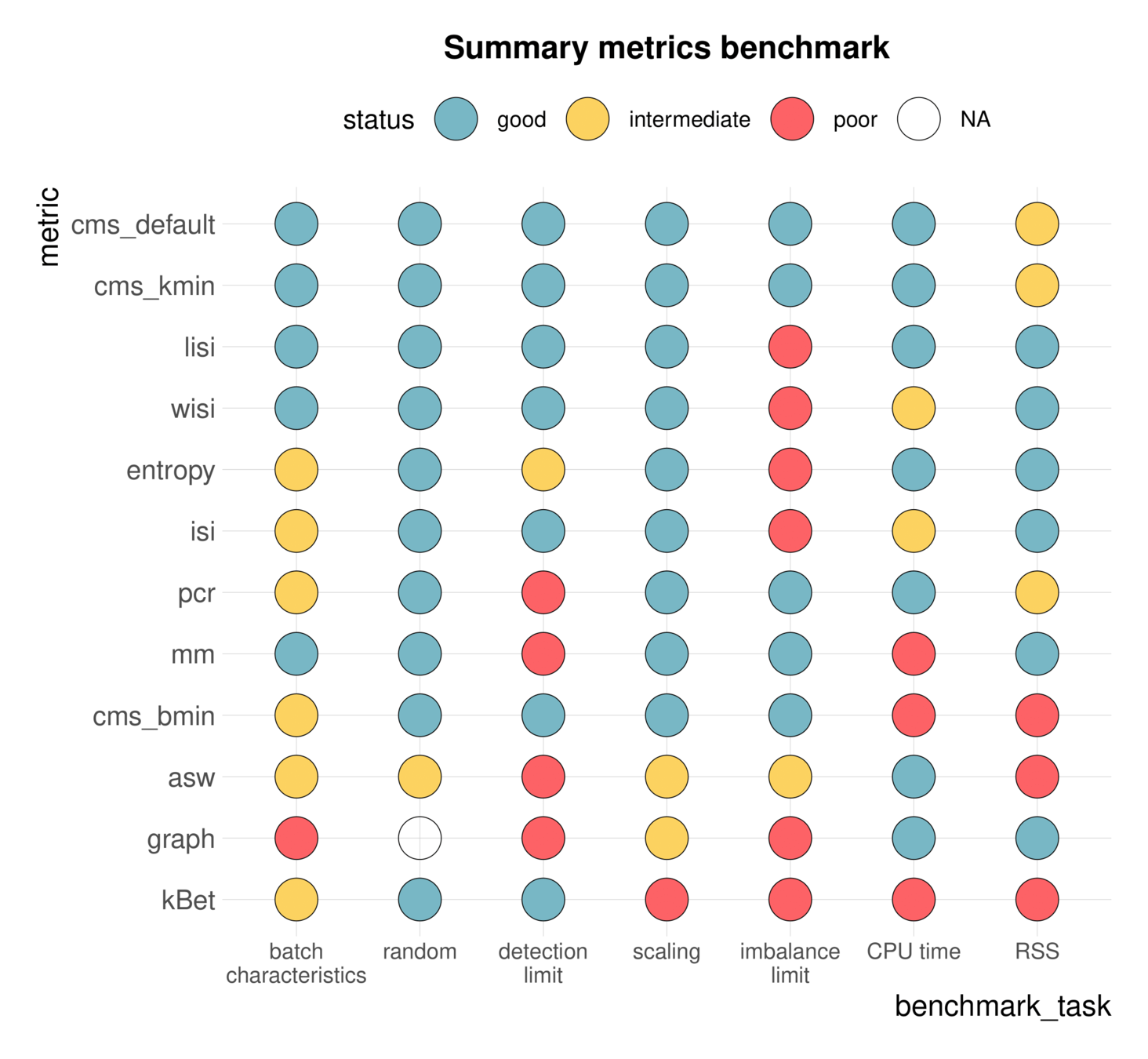
Comparisons can be ambiguous
Luecken et al., 2020

Comparisons can be ambiguous
Luecken et al., 2020
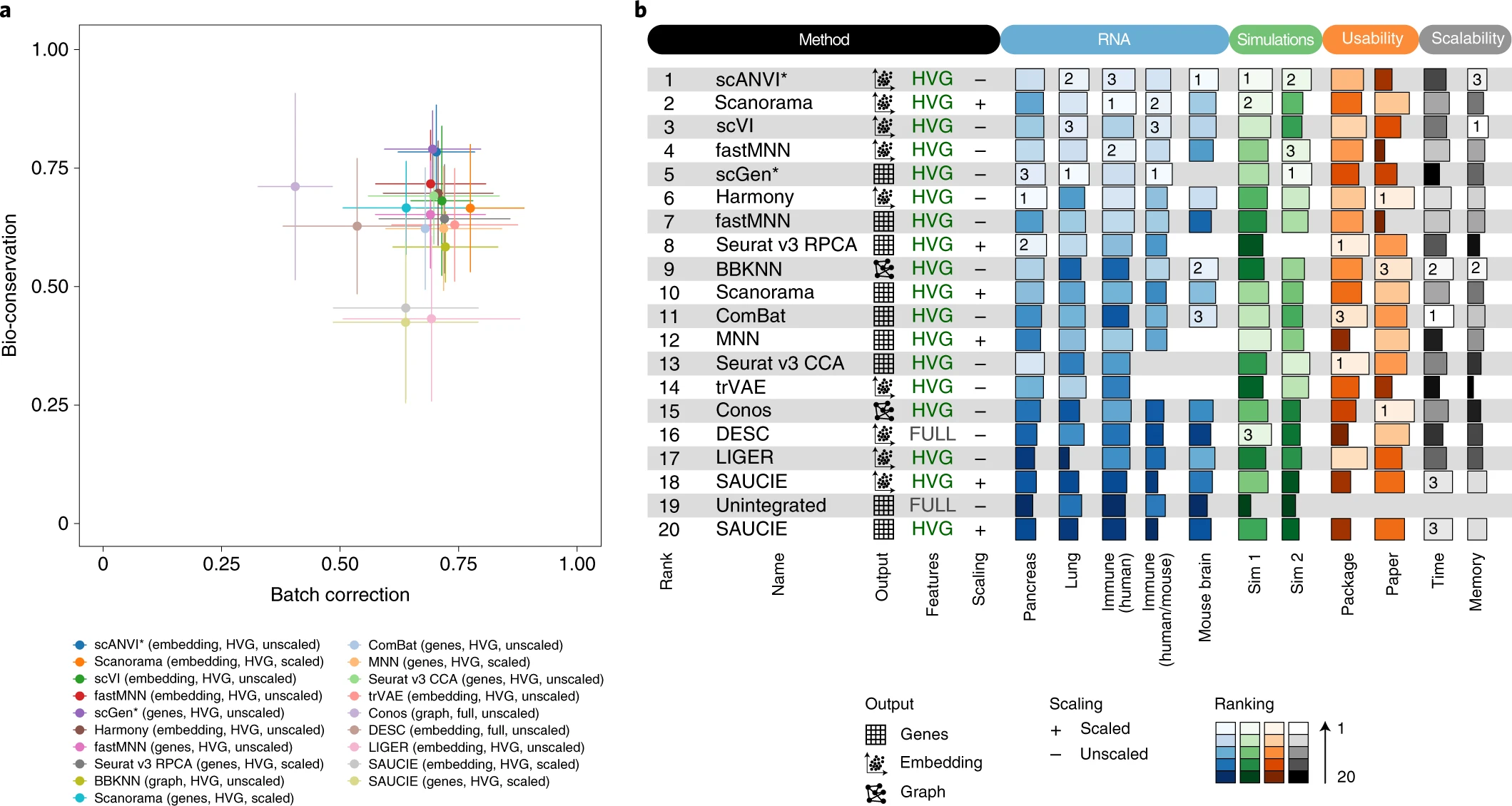
Static benchmarks are limited in scope and interpretability
- There is not one way to benchmark different methods
- There are multiple ways to interpret benchmark results
--> Open extensible community benchmarks
status quo:
Meta-analysis of 62 method benchmarks in the field of single cell omics

62 single cell omics method benchmarks
2 reviewer per benchmark
Meta-analysis:
-
Title
-
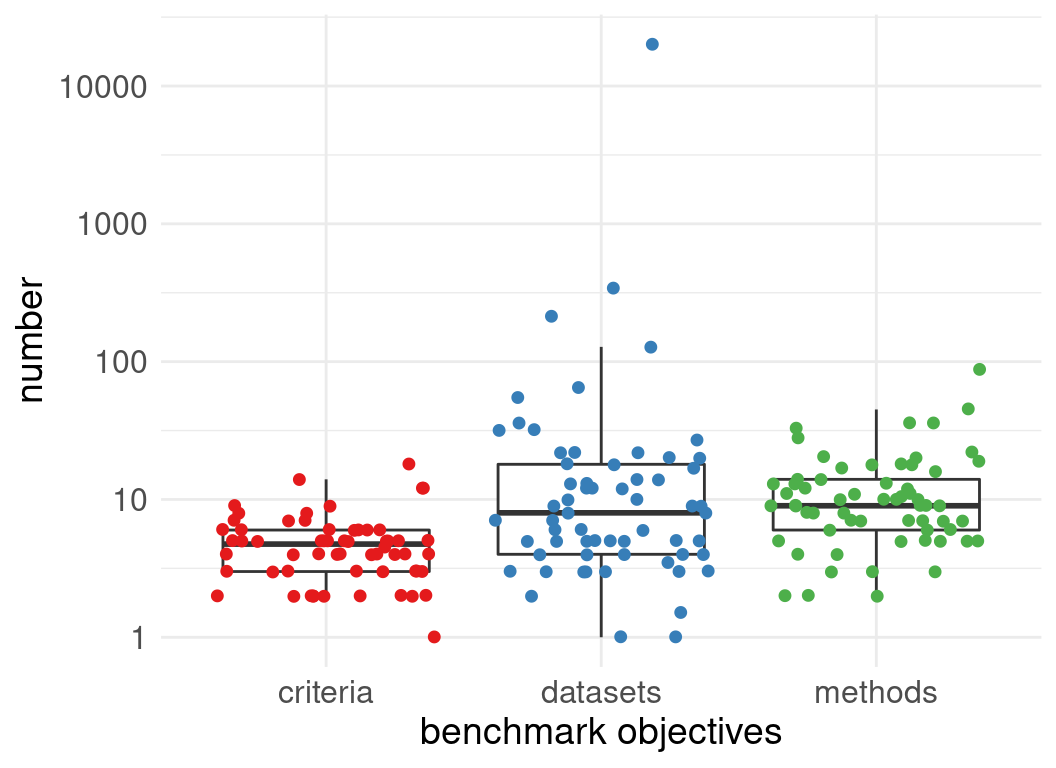
Number of datasets used in evaluations:
-
Number of methods evaluated:
-
Degree to which authors are neutral:
...
22. Type of workflow system used:
independent harmonization of responses

summaries
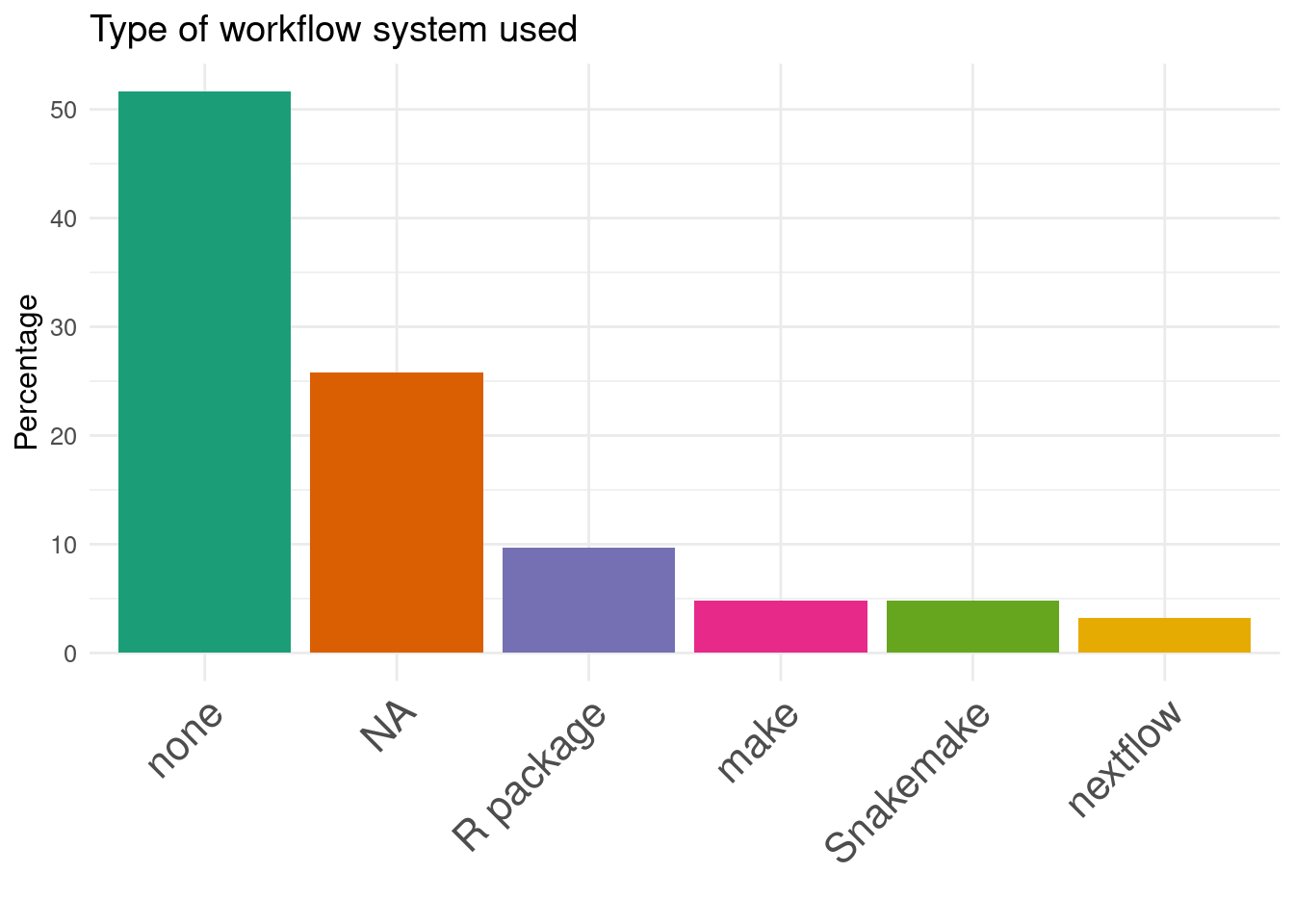
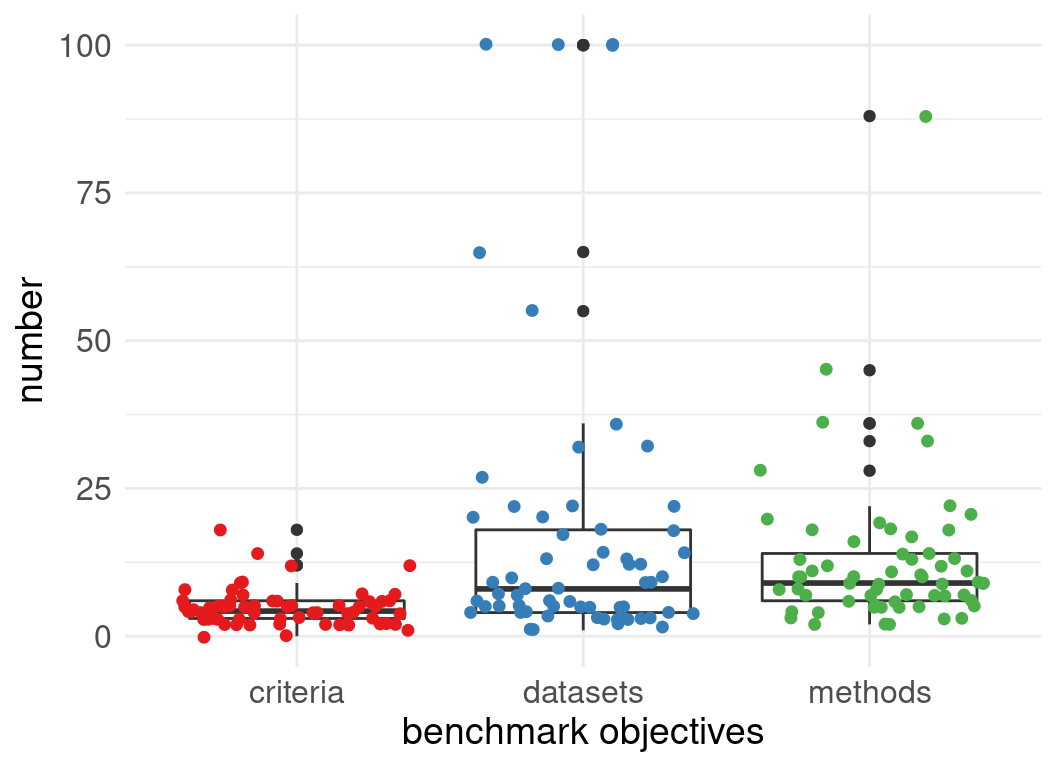
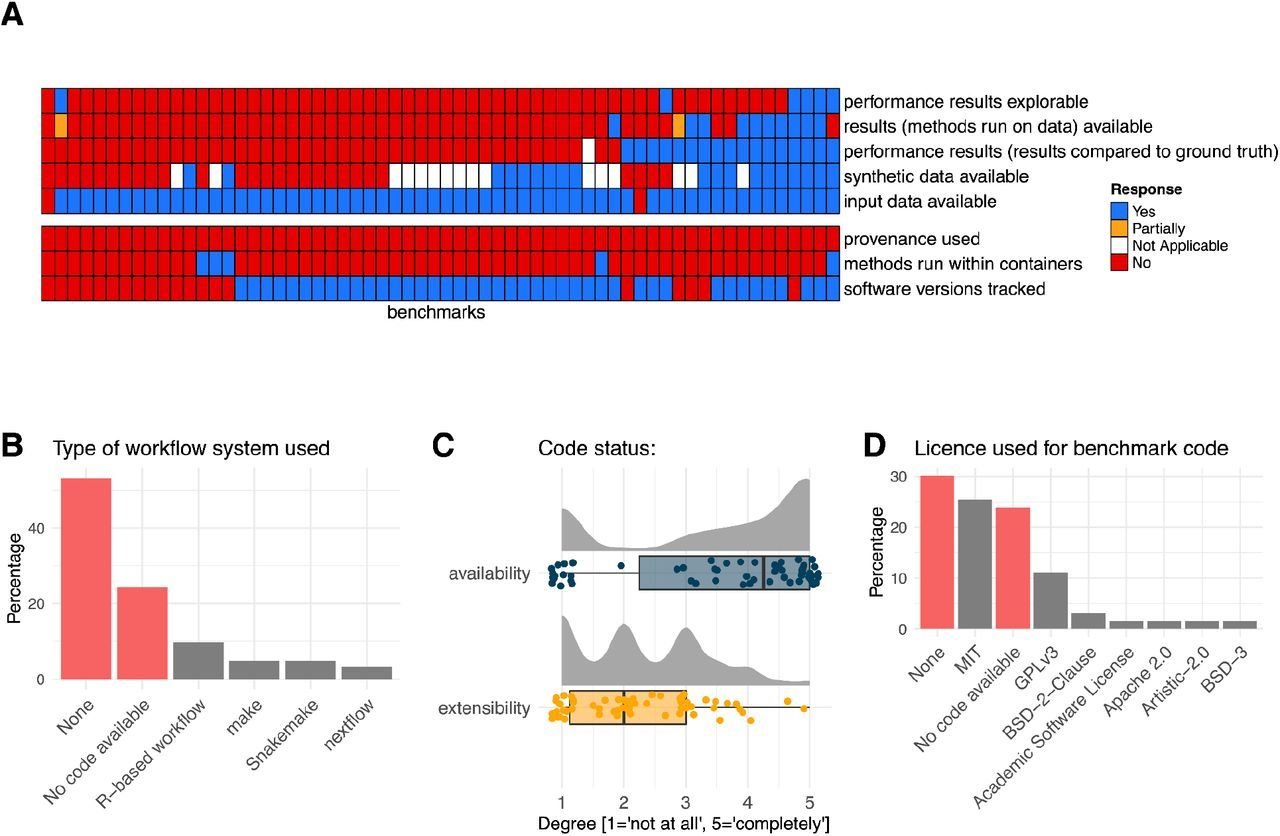
Benchmark designs:
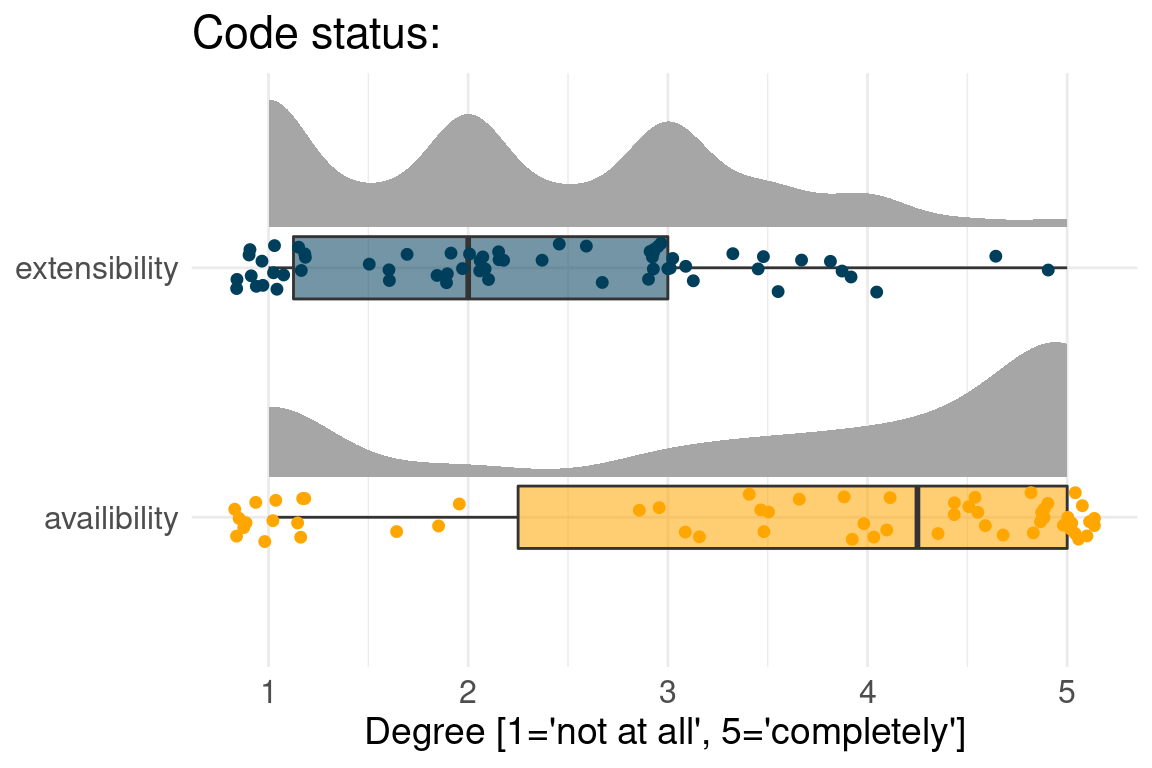
Often Benchmark code is available but not extensible
usually input data are available, but not results
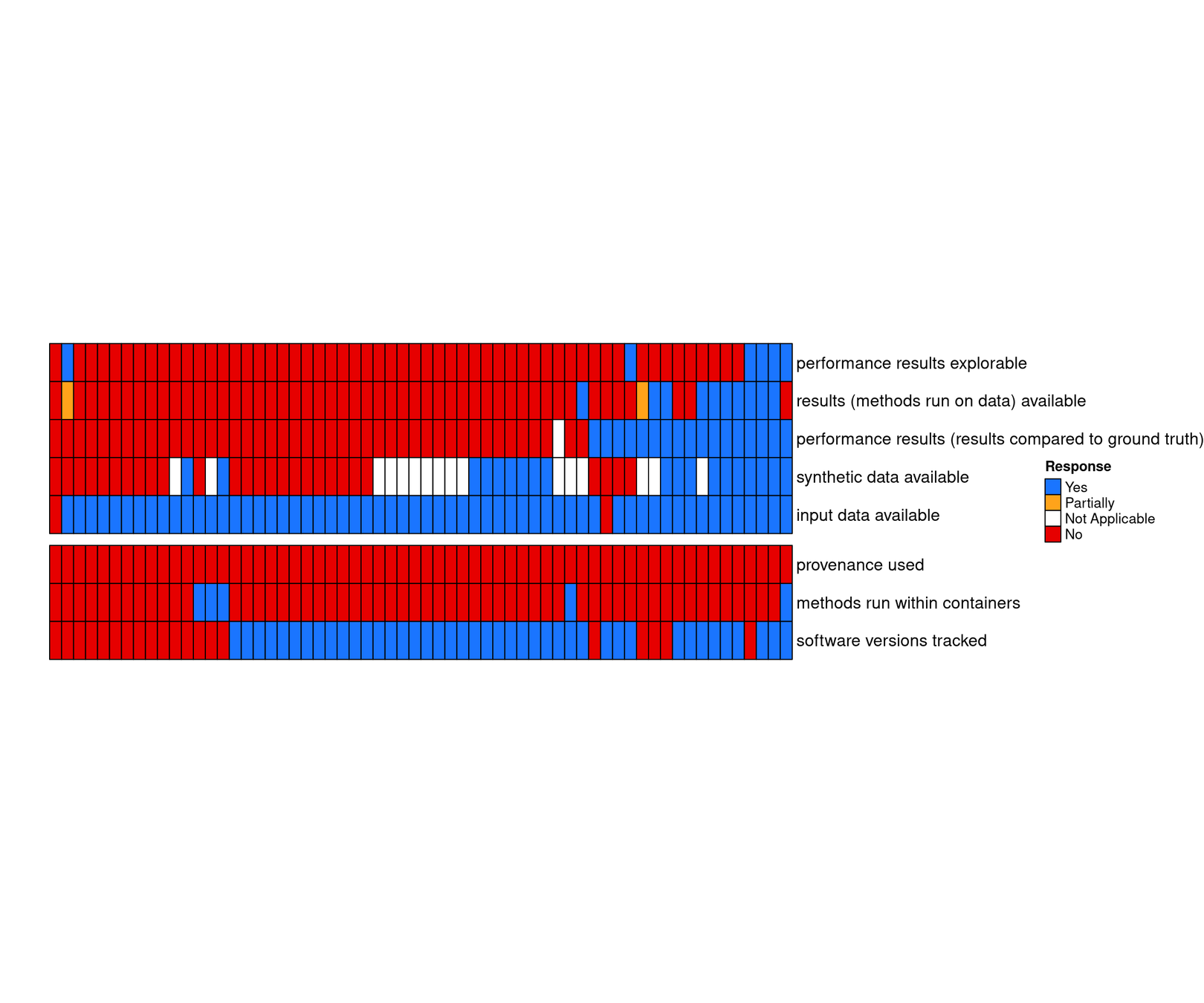
Workflow manager are rarely used
Blocks of open and continuous benchmarking
Code
available
extensible
reusable
conclusion
neutral
community-driven
Reproducibility
code
workflows
enviroments
software versions
time-Scale
static
continuous
Open Data
input data
method results
simulations
performance results
currently part of most benchmarks
not part of current standards
Omnibenchmark:
Open and continuous community benchmarking



Omnibenchmark is a platform for open and continuous community driven benchmarking
Method developer/
Benchmarker
Method user
Methods
Datasets



Metrics
Omnibenchmark
- continuous
- self-contained modules
- all "products" can be accessed
- provenance tracking
- anyone can contribute
Omnibenchmark design
Data
standardized datasets


= 1 "module" (renku project )



Methods
method results

Metrics
metric results

Dashboard
interactive result exploration
Method user
Method developer/
Benchmarker
Omnibenchmark modules are independent and self-contained

GitLab project

Docker container

Workflow

Datasets
Collection of
method* history,
description how to run it, comp. environment, datasets
=
components of omnibenchmark
Omnibenchmark-python

pypy module for workflow and dataset management with renku/KG
orchestrator

CICD Orchestrator to automatically run and update benchmarks
omnibenchmark-graph

Triplet store to perform cross repository queries
Bettr

Shiny app to interactively explore results

module Processes (Benchmark steps) are stored as triplets
Result
Code
Data
generated
used_by
used_by
Data
Code
Result
used_by
generated
Subject
predicate
Object
Triplet
modules are connected via knowledge Base




Module A




Module B
Triplet Store

triplet generation
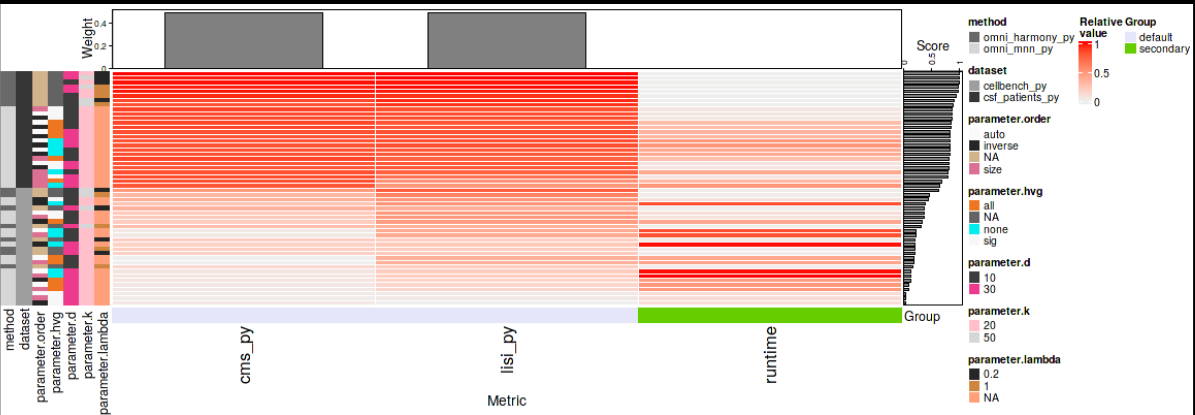
bettr: A better way to explore what is best
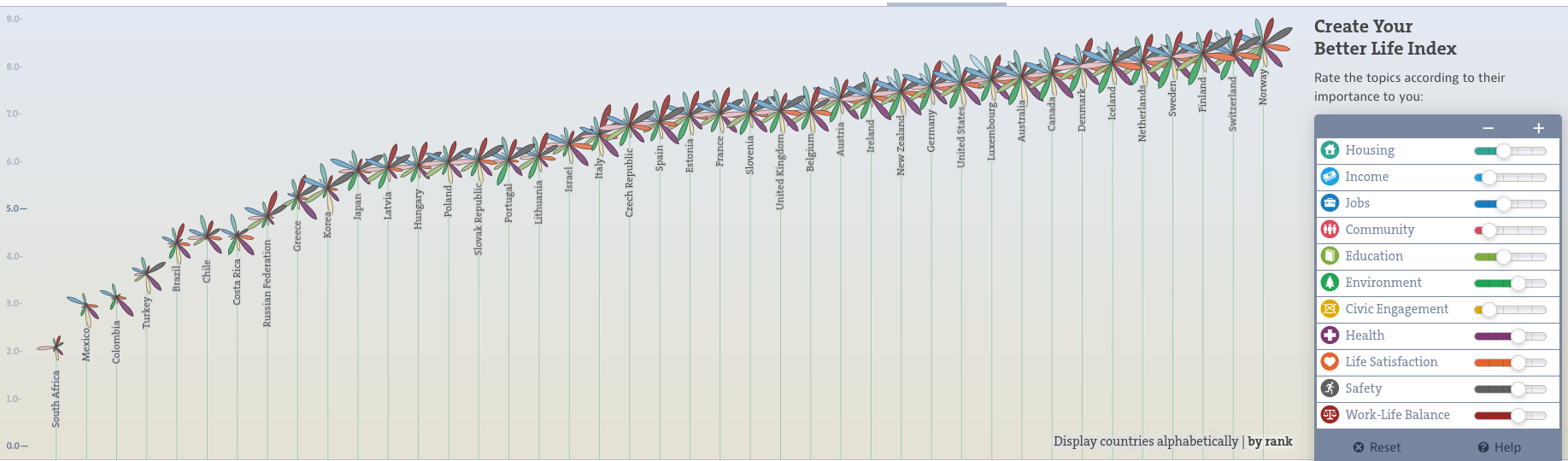
https://www.oecdbetterlifeindex.org
bettr: A better way to explore what is best
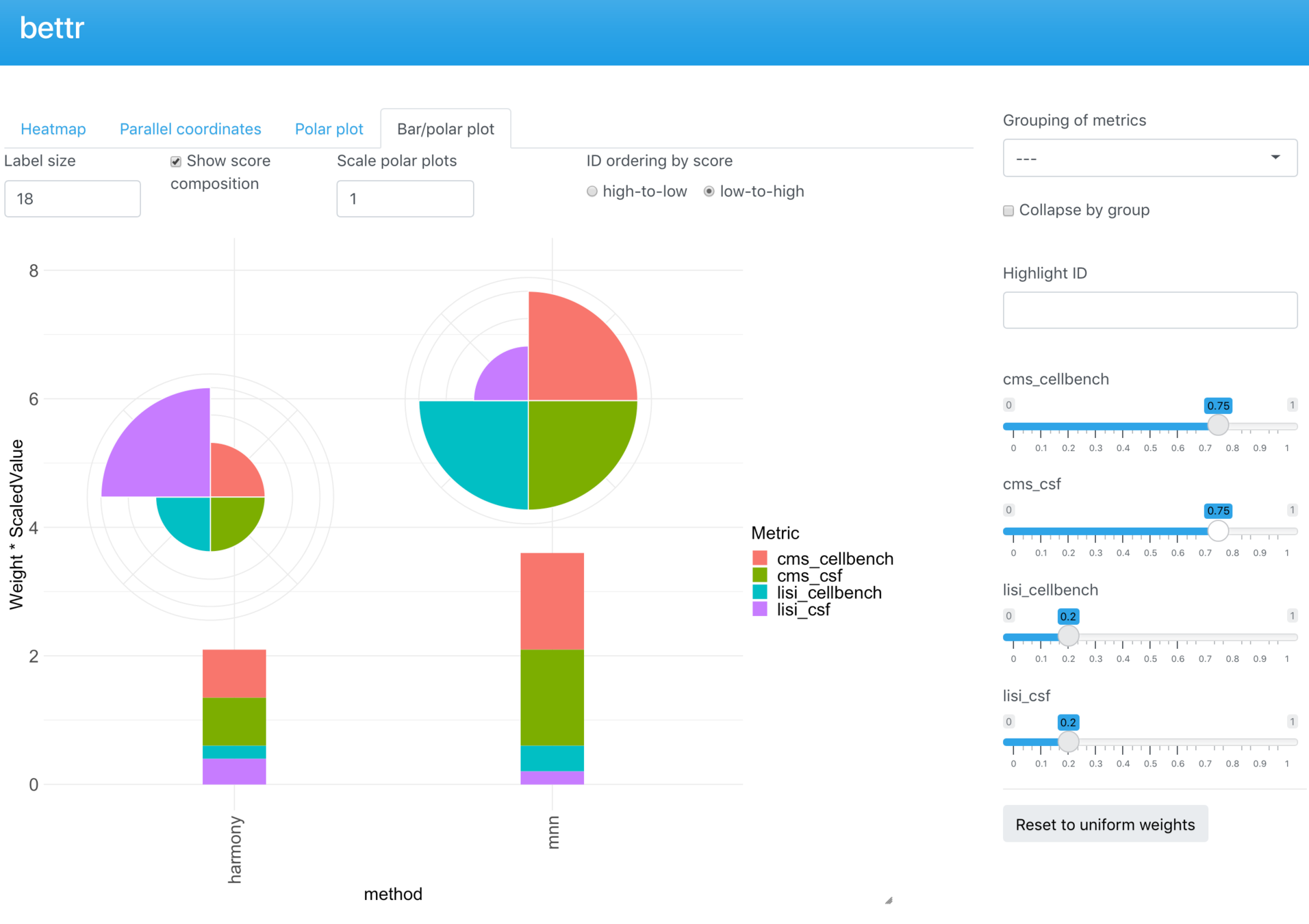
Project status
- 2 continuous prototype benchmarks
- Hackathon
- Dashboard
- Documentation
- Templates
Acknowledgments

Robinson group
Mark Robinson
Anthony Sonrel
Izaskun Mallona
Pierre-Luc Germain

Renku team
Oksana Riba Grognuz
Friedrich Miescher institute
Charlotte Soneson
THANK YOU!

A data analysis platform/system built from a set of microservices
GitLab --> version control/CICD


Apache Jena --> Triple store


Jupyter server --> interactive sessions

Docker/Kubernetes --> software/enviroment management

GitLFS --> File storage
What is renku?

Renku client is a dataset and workflow management system
Renku client
-
Dataset and workflow management system → “renku-python”
-
Knowledge graph tracking → provenance
Renkulab
-
User interface with free interactive sessions
-
GitLab
Renku client is based on a Triplet store (Knowledge graph)

Result
Code
Data
generated
used_by
used_by
Data
Code
Result
used_by
generated
User interaction with renku client
Automatic triplet generation
Triplet store "Knowledge graph"
User interaction with renku client
KG-endpoint queries
orchestrator defines benchmark scopes
Each benchmark has their own orchestrator
Schedules automatic module updates
"Gate-keeping" - controls addition of new modules