Omnibenchmark:
(Re-)design and future plans
Science talk
Zürich, 04.10.23




Omnibenchmark:
General design and concepts

GitLab

Docker

Workflow

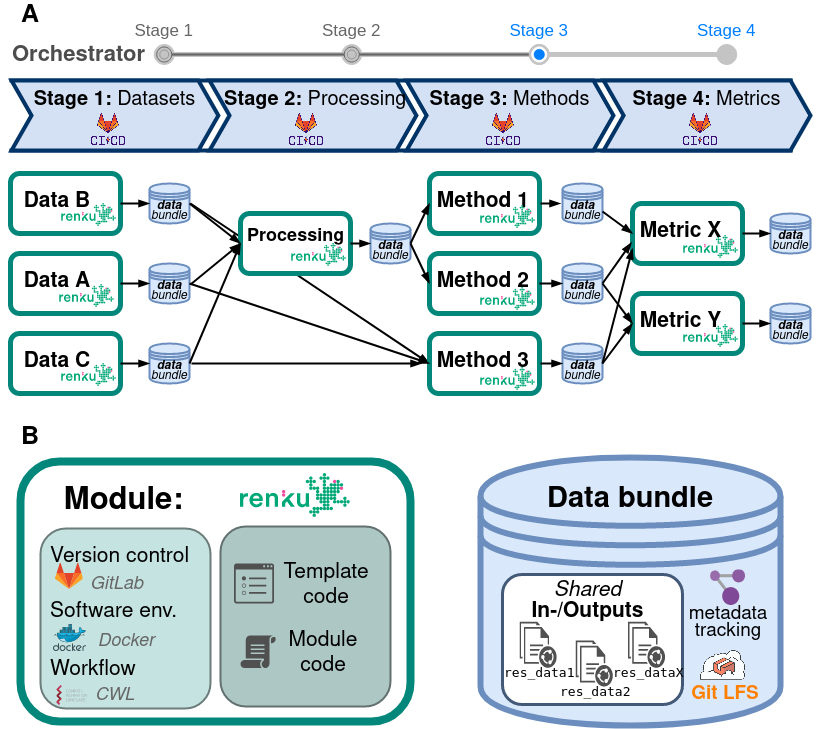
Module:
Template code
Module code

Omnibenchmark components
Omnibenchmark design



Omnibenchmark:
Current limitations
Design Limitations
1. Limited project "cross-talk"

Design Limitations
1. Limited project "cross-talk"

Renku rest api
-
GET /knowledge-graph/entities
Allows finding projects, datasets, workflows, and persons
--> omb is "file-centric" (activity-centric)
-
GET /knowledge-graph/projects/:namespace/:name
Finds details of the project with the given namespace/name
Renku rest api
-
version
-
scope and speed of queries
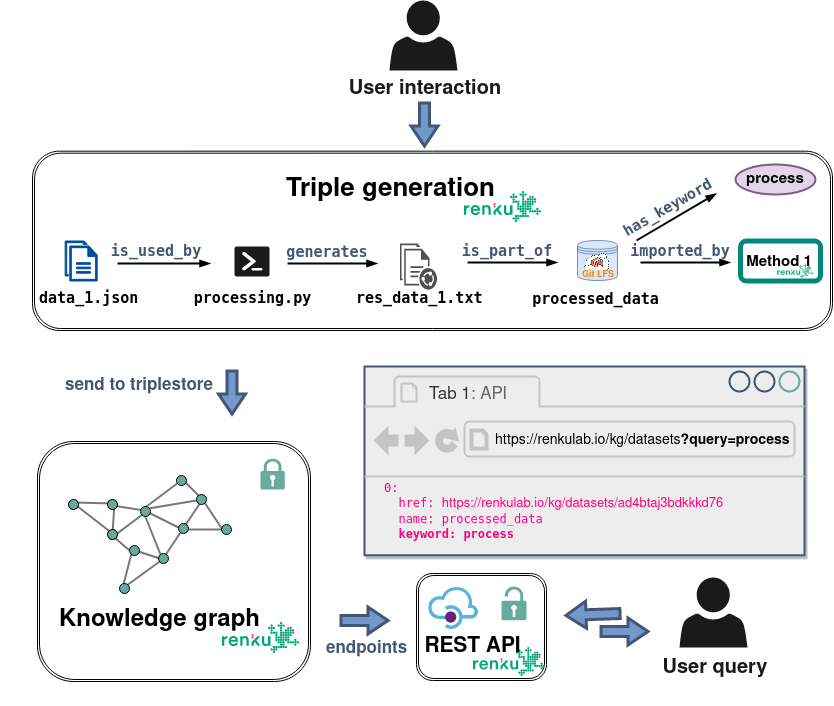
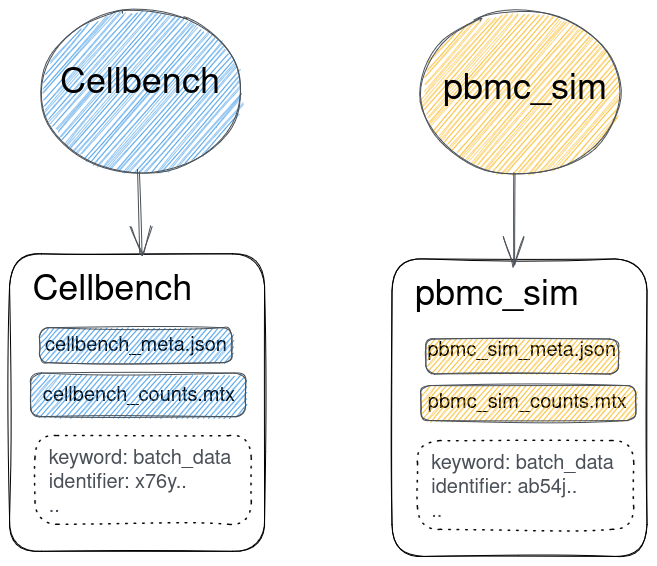
Why project "cross-talk"?
Why project "cross-talk"?


Why project "cross-talk"?


Why project "cross-talk"?


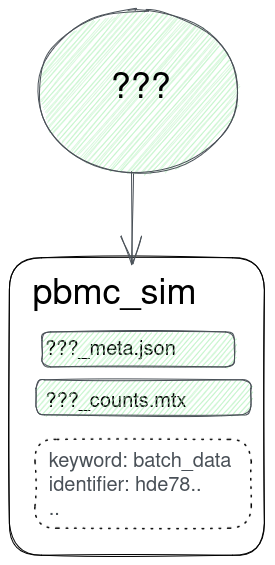
Why project "cross-talk"?
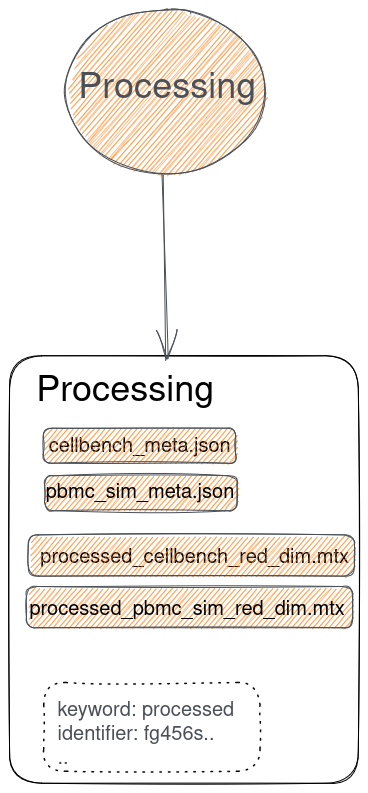

Why project "cross-talk"?


Why project "cross-talk"?


How to implicitly match files from the same origin dataset?
Why project "cross-talk"?




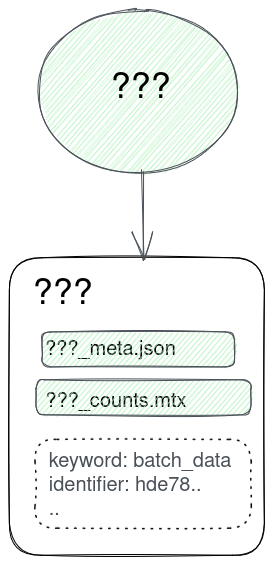


Lineages!
Lineages
-
fast and performant
--> renku ontology is not file-centric

Lineages
-
fast and performant
--> renku ontology is not file-centric

Lineages
-
ontology (Izaskun)
-
generate + send triples during benchmark execution (plugin, omni-cli, Ben)
-
queries (Izaskun)
-
adapt omnibenchmark-python (Almut)
-
integrate into orchestrator runs
Design Limitations
1. Limited project "cross-talk"
2. Missing concept of versions








Where do versions help?
- Define scope (storage policies etc.)
- Reduce search space in queries
- Reproducibility/ Releases
- Prevent race conditions

- (Local benchmarks)
Epochs to version benchmarks
epoch == 1 (full?) orchestrator run
--> benchmarks are defined by the orchestrator (scope + order + epoch)
--> epoch as part of omni-ontology (cross-project communication)
--> orchestrator as .yaml file (local graph/epochs)
Design Limitations
1. Limited project "cross-talk"
2. Missing concept of versions
3. Renku dependency

Omnibenchmark-Renku Relationship
Renku is a set of microservices:

| RENKU SERVICE | RELATIONSHIP STATUS |
|---|---|
| renku-python | complicated |
| renku graph | in separation |
| renku GUI | open relationship |
| renkuLab | long distance |
Omnibenchmark-Renku strategy
- reduce dependencies

- increase interaction
Omnibenchmark-Renku strategy
- reduce dependencies

- increase interaction/communication
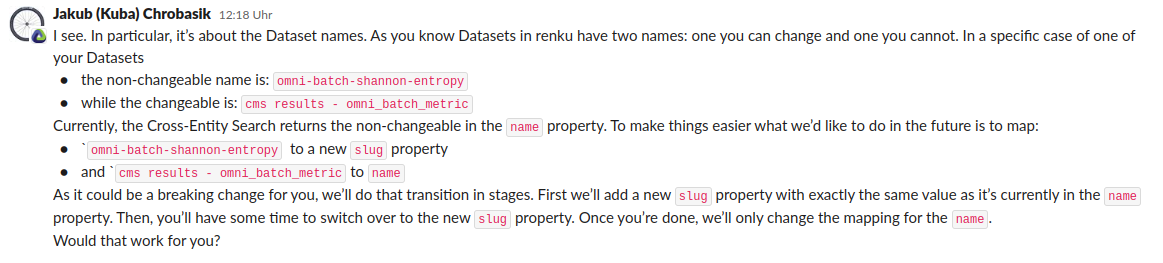
Omnibenchmark-Renku strategy

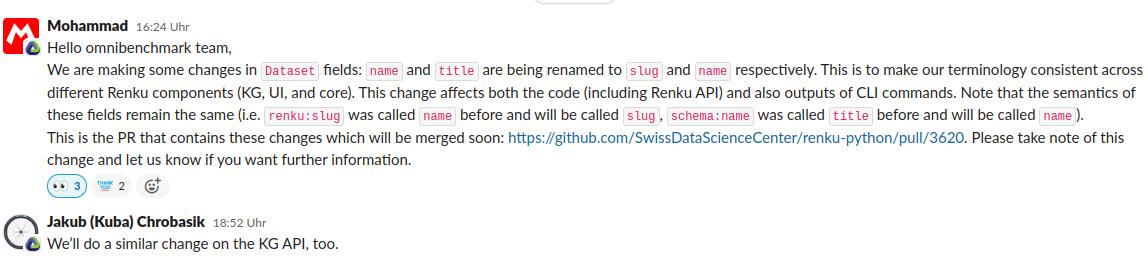
Summary

- some OMB base changes in progress
- OMB ontology, epoch concept
- Reduce some renku dependency
- user facing changes?
Design: Benchmark modules
Data
standardized datasets


= 1 "module" (renku project )



Methods
method results

Metrics
metric results

Dashboard
interactive result exploration
Method user
Method developer/
Benchmarker
The omnibenchmark ecosystem
Omnibenchmark Python
1. User module specification

# src/config.yaml
---
data:
name: "out_dataset"
title: "Output of an example OmniObject"
description: "describe module here"
keywords: ["example_dataset"]
script: "path/to/method/dataset/metric/script.py"
benchmark_name: "omni_celltype"
inputs:
keywords: ["import_this", "import_that"]
files: ["count_file", "dim_red_file"]
prefix:
count_file: "counts"
dim_red_file: ["features", "genes"]
outputs:
files:
corrected_counts:
end: ".mtx.gz"
meta:
end: ".json"
parameter:
names: ["param1", "param2"]
keywords: ["param_dataset"]
Omnibenchmark Python
1. User module specification

# src/run_workflow.py
from omnibenchmark import get_omni_object_from_yaml, renku save
## Load config
omni_obj = get_omni_object_from_yaml('src/config.yaml')
## Check for new/updates of input datasets
omni_obj.update_object()
renku_save()
## Create output dataset
omni_obj.create_dataset()
## Generate and run workflow
omni_obj.run_renku()
renku_save()
## Store results in output dataset
omni_obj.update_result_dataset()
renku_save()
Omnibenchmark Python
1. User module specification

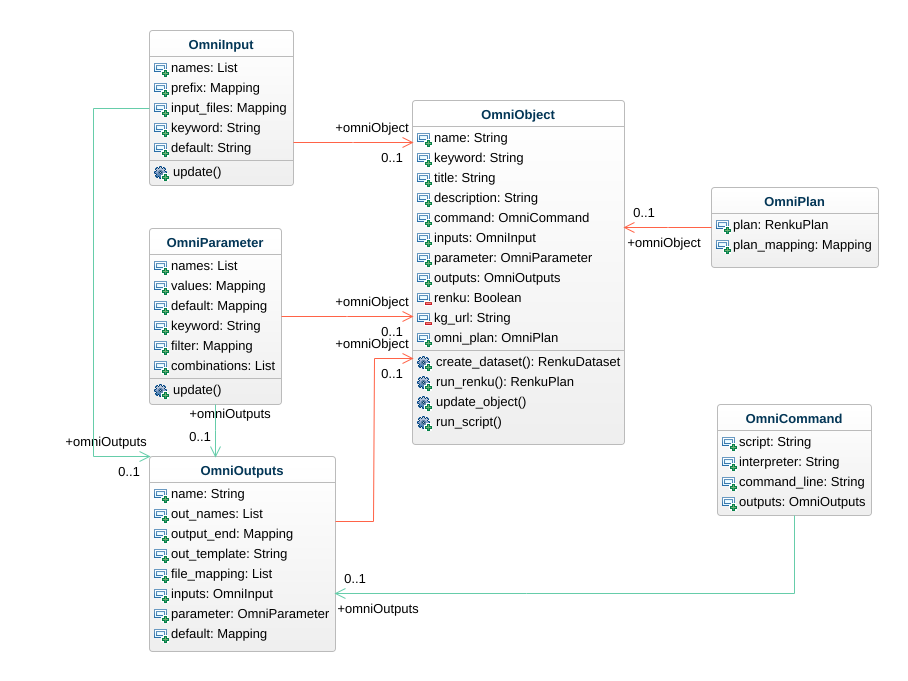
Omnibenchmark Python
2. Manage input import/updates

data:
name: "iris_random_forest"
title: "Random Forest"
description: "Random forest applied on the iris dataset"
keywords: ["iris_method"]
script: "src/iris-random-forest.R"
benchmark_name: "iris_example"
inputs:
keywords: ["iris_filtered"]
files: ["iris_input"]
prefix:
iris_input: "_filt_dataset"
outputs:
template: "data/${name}/${name}_${unique_values}_${out_name}.${out_end}"
files:
rf_model:
end: ".rds"
parameter:
keywords: ["iris_parameters"]
names: ["train_split", "rseed"]
Example: Method module
Omnibenchmark Python
2. Manage input import/updates

Example: Method module
-
benchmark: iris_example
-
keyword: iris_filtered


"endpoints"
1.
Omnibenchmark Python
2. Manage input import/updates

Example: Method module
-
benchmark: iris_example
-
keyword: iris_filtered
Omnibenchmark Python
2. Manage input import/updates

Example: Method module
-
benchmark: iris_example
-
keyword: iris_filtered


"endpoints"
1.
2.

Orchestrator



Omnibenchmark Python
2. Manage input import/updates

data:
name: "iris_random_forest"
title: "Random Forest"
description: "Random forest applied on the iris dataset"
keywords: ["iris_method"]
script: "src/iris-random-forest.R"
benchmark_name: "iris_example"
inputs:
keywords: ["iris_filtered"]
files: ["iris_input"]
prefix:
iris_input: "_filt_dataset"
outputs:
template: "data/${name}/${name}_${unique_values}_${out_name}.${out_end}"
files:
rf_model:
end: ".rds"
parameter:
keywords: ["iris_parameters"]
names: ["train_split", "rseed"]
Example: Method module
Omnibenchmark Python
2. Generalize user specifications

data:
name: "iris_random_forest"
title: "Random Forest"
description: "Random forest applied on the iris dataset"
keywords: ["iris_method"]
script: "src/iris-random-forest.R"
benchmark_name: "iris_example"
inputs:
keywords: ["iris_filtered"]
files: ["iris_input"]
prefix:
iris_input: "_filt_dataset"
outputs:
template: "data/${name}/${name}_${unique_values}_${out_name}.${out_end}"
files:
rf_model:
end: ".rds"
parameter:
keywords: ["iris_parameters"]
names: ["train_split", "rseed"]
Example: Method module
Omnibenchmark Python
3. Generalize user specifications

Example: Method module

iris-random-forest.R
*._filt_dataset*.

-
benchmark: iris_example
-
keyword: iris_filtered
Omnibenchmark Python
2. Generalize user specifications

The omnibenchmark ecosystem

omni-validator


omni-sparql

triple store

omni-cli
omni-utils

The omnibenchmark Infrastructure



- omni ontology triples
- project-specific renku ontology triples

GitLFS

File storage
?
imlsomnibenchmark
- Version control (git)
- CICD
- UI
- API

renkuLab

renku GUI
- Interactive sessions
- Authentication (?)
- Queries
- Boards(?)