Omnibenchmark:
Open and continuous community benchmarking
Almut Lütge & Anthony Sonrel



status quo:
Meta-analysis of 62 method benchmarks in the field of single cell omics

62 single cell omics method benchmarks
2 reviewer per benchmark
Meta-analysis:
-
Title
-
Number of datasets used in evaluations:
-
Number of methods evaluated:
-
Degree to which authors are neutral:
...
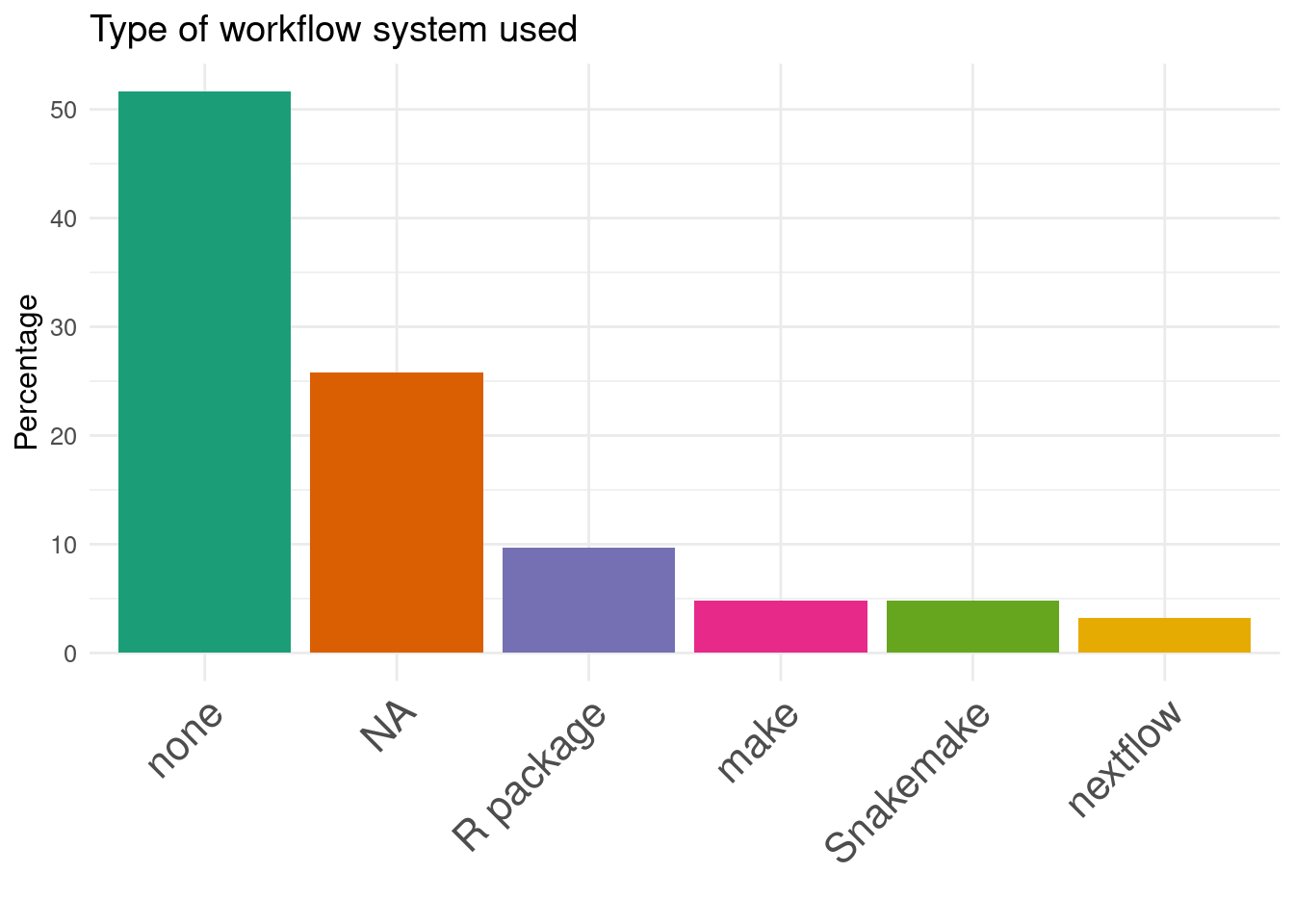
22. Type of workflow system used:
independent harmonization of responses

summaries
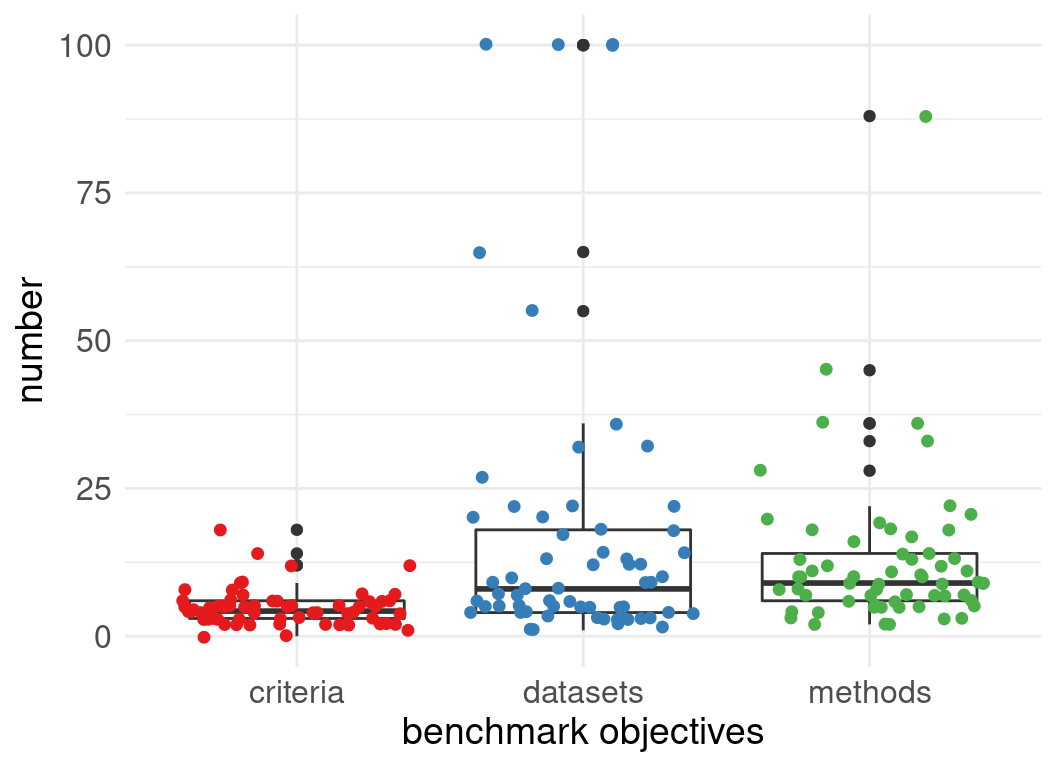
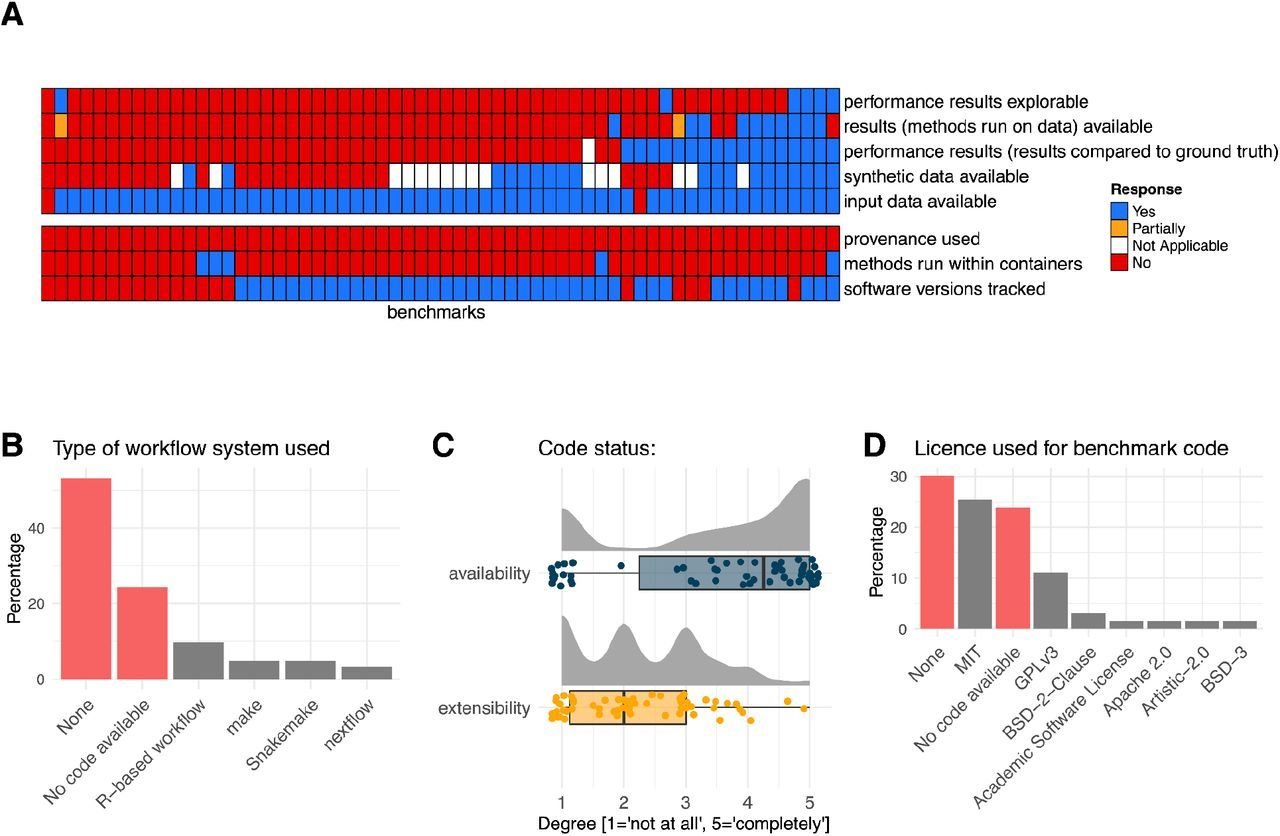
Benchmark designs:
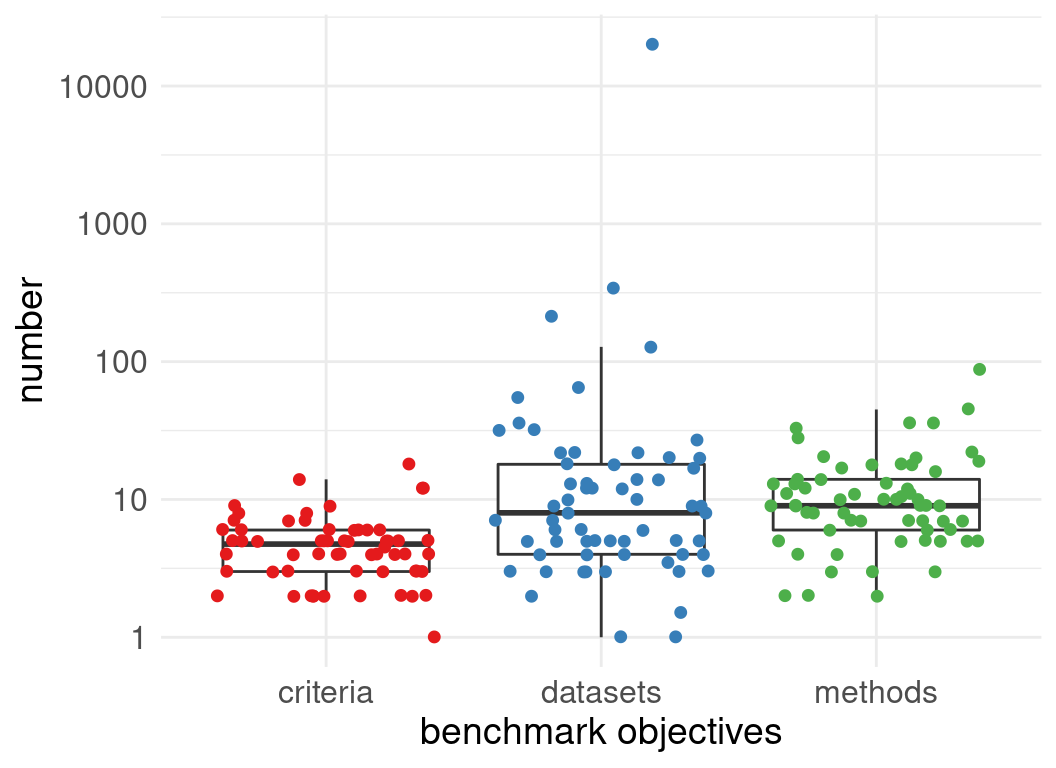
Often Benchmark code is available but not extensible
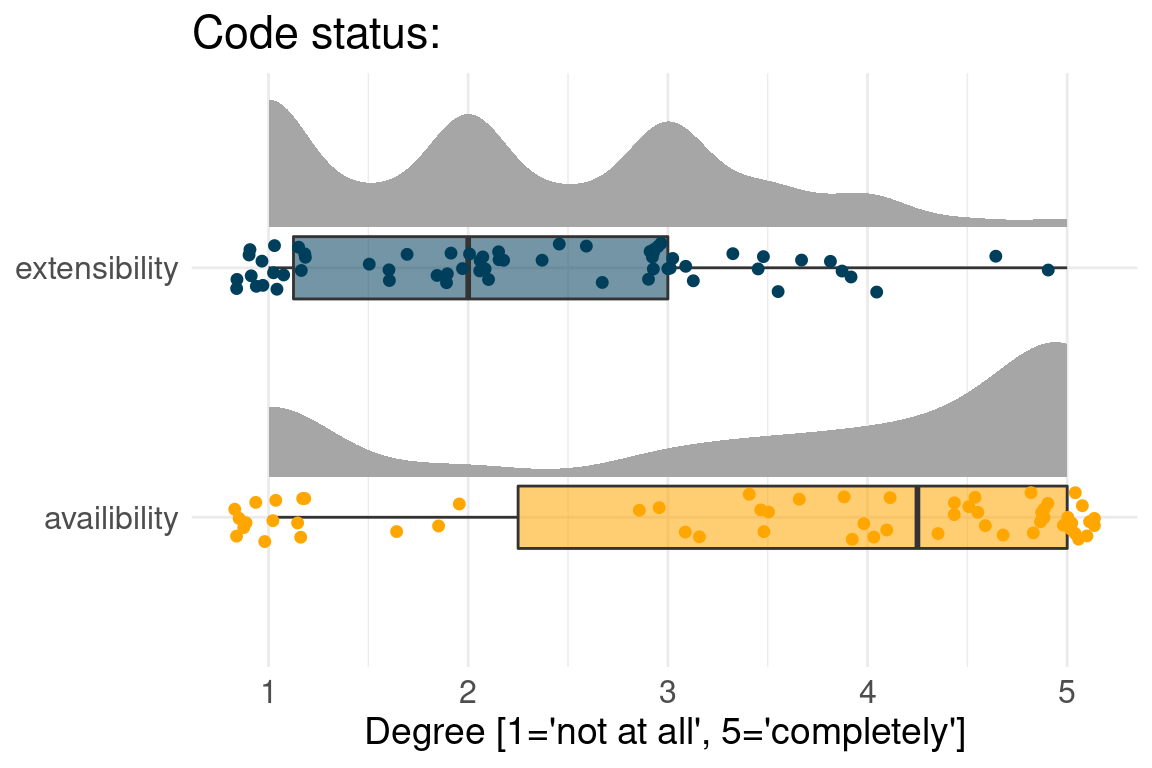
usually input data are available, but not results
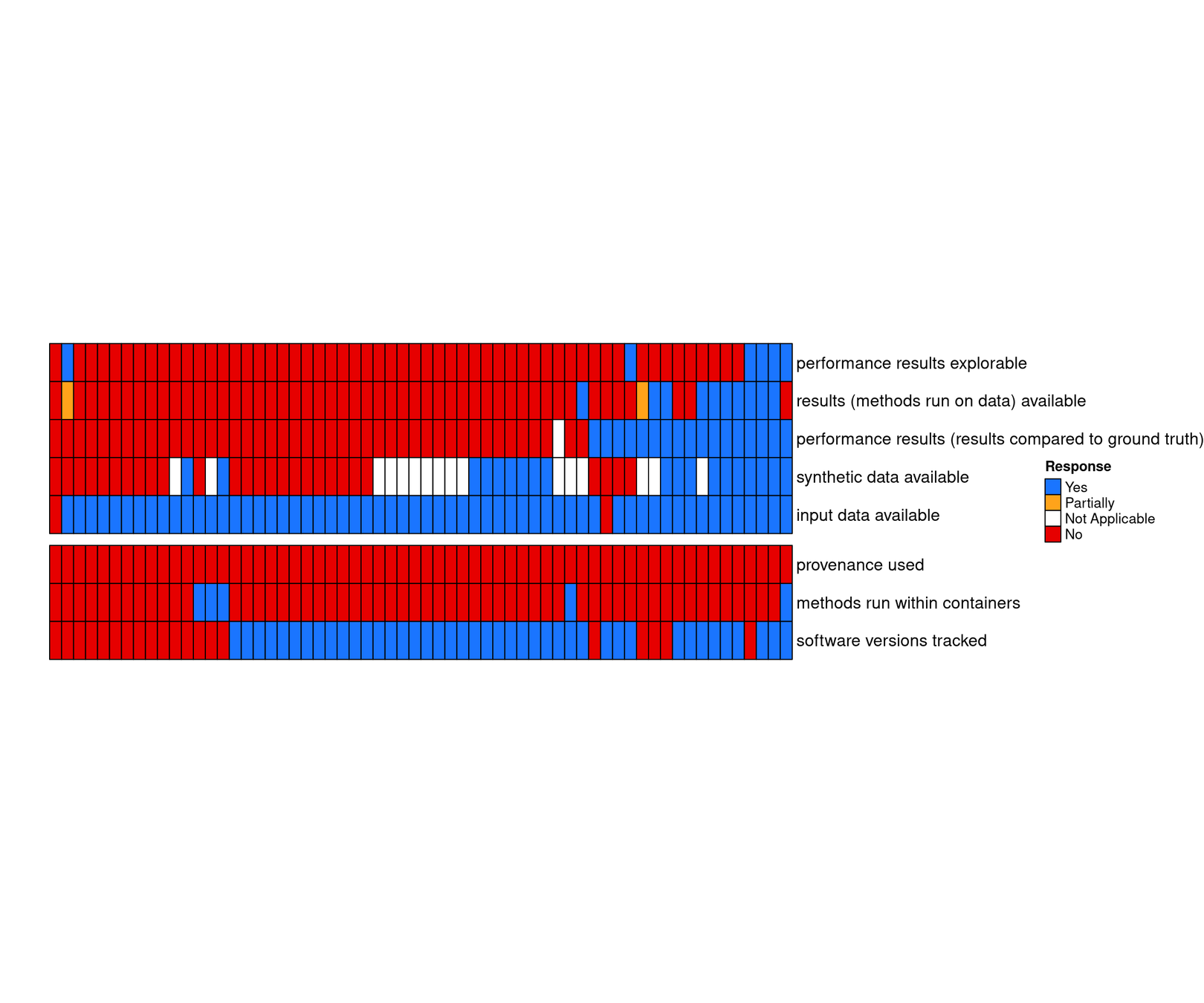
Workflow manager are rarely used
Blocks of open and continuous benchmarking
Code
available
extensible
reusable
conclusion
neutral
community-driven
Reproducibility
code
workflows
enviroments
software versions
time-Scale
static
continuous
Open Data
input data
method results
simulations
performance results
currently part of most benchmarks
not part of current standards
Omnibenchmark is a platform for open and continuous community driven benchmarking
Method developer/
Benchmarker
Method user
Methods
Datasets



Metrics
Omnibenchmark
- continuous
- self-contained modules
- all "products" can be accessed
- provenance tracking
- anyone can contribute

Omnibenchmark design
Data
standardized datasets


= 1 "module" (renku project )



Methods
method results

Metrics
metric results

Dashboard
interactive result exploration
Method user
Method developer/
Benchmarker
Omnibenchmark design
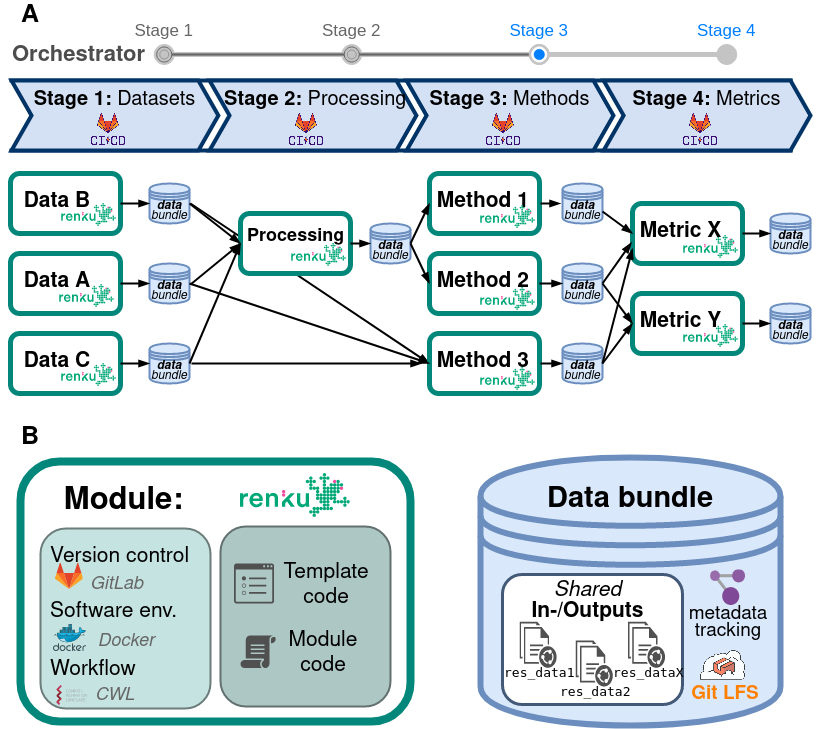

GitLab

Docker

Workflow

Module:
Template code
Module code

Omnibenchmark design


Omnibenchmark components
contributer
user

omnibenchmark-python

omniValidator
benchmarker
projects

templates

omb-site

{
orchestrator

triplestore


omni-sparql
dashboards

Omnibenchmark user I
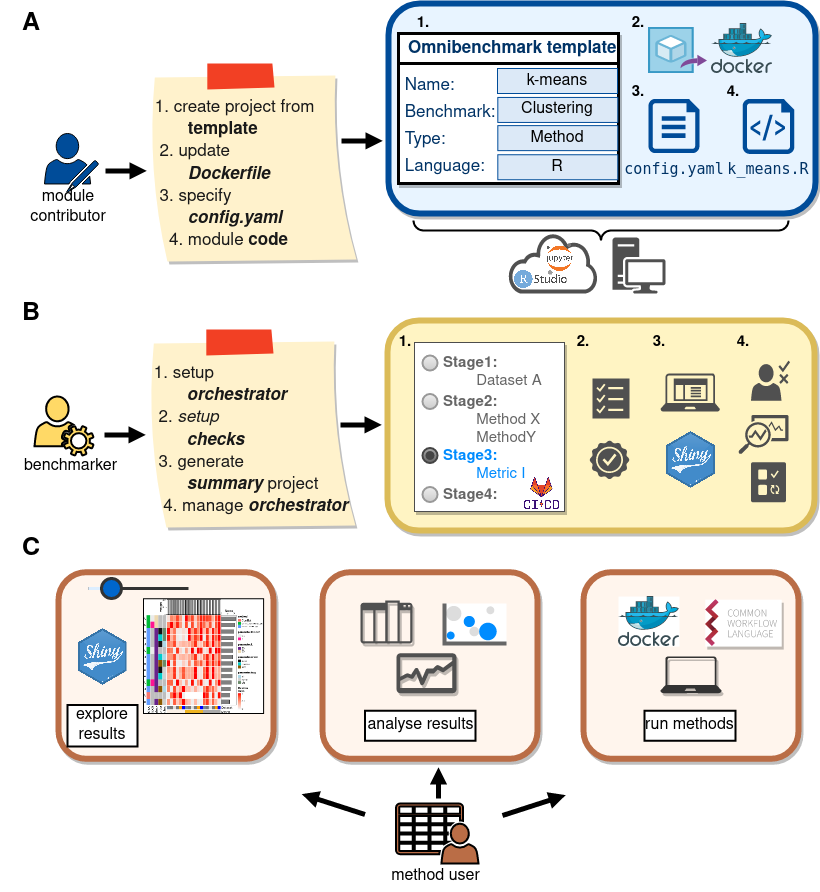
Omnibenchmark user II

Omnibenchmark user III

Data
standardized datasets

new module
Method developer/
Benchmarker
Cumbersome to:
- clone project
- find fields to modify
- connect to other components
Omnibenchmark templates
Data
standardized datasets

templates
Pre-filled projects
Method developer/
Benchmarker
--> Templates allow easy contribution to an Omnibenchmark
Method developer/
Benchmarker
How do templates work ?
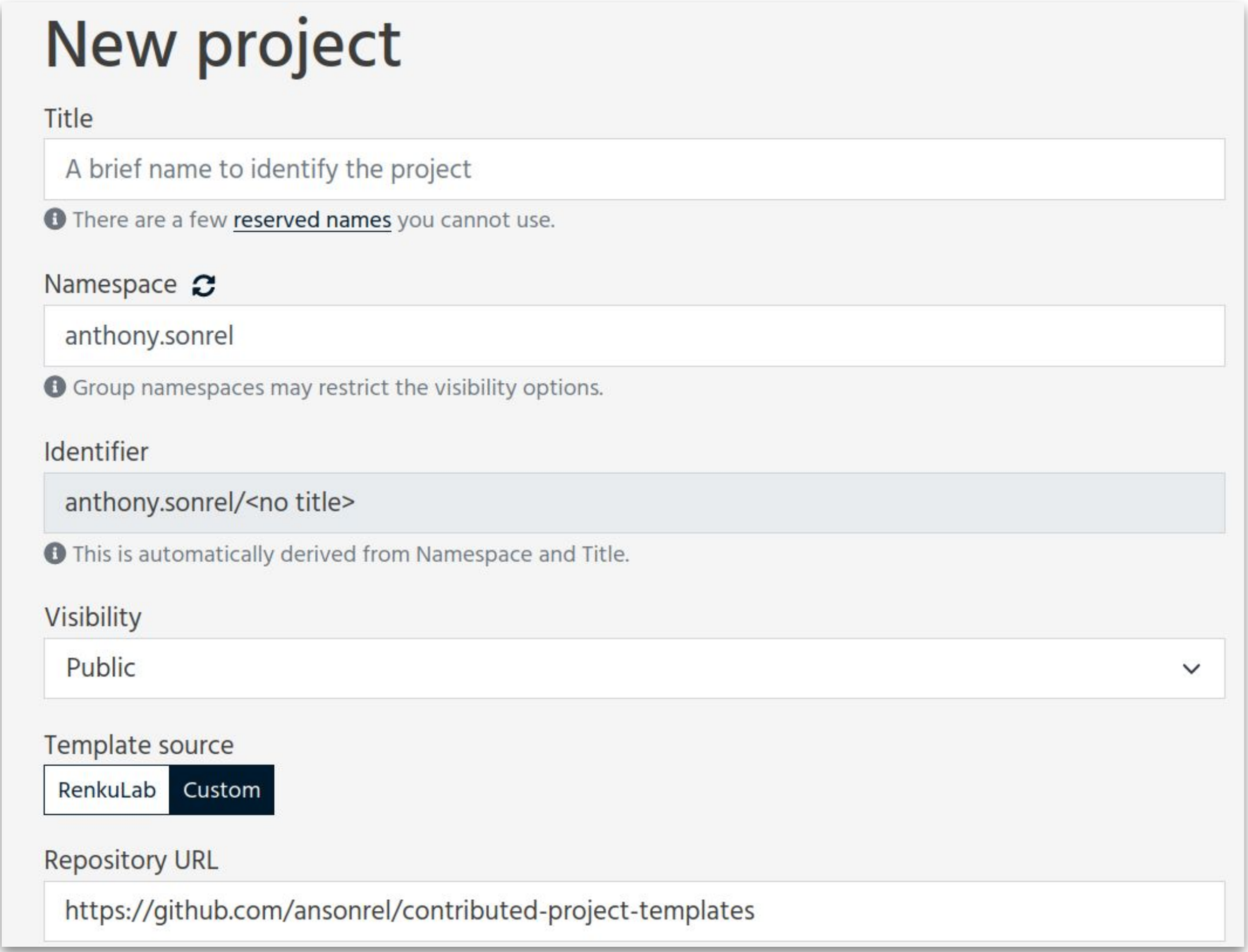
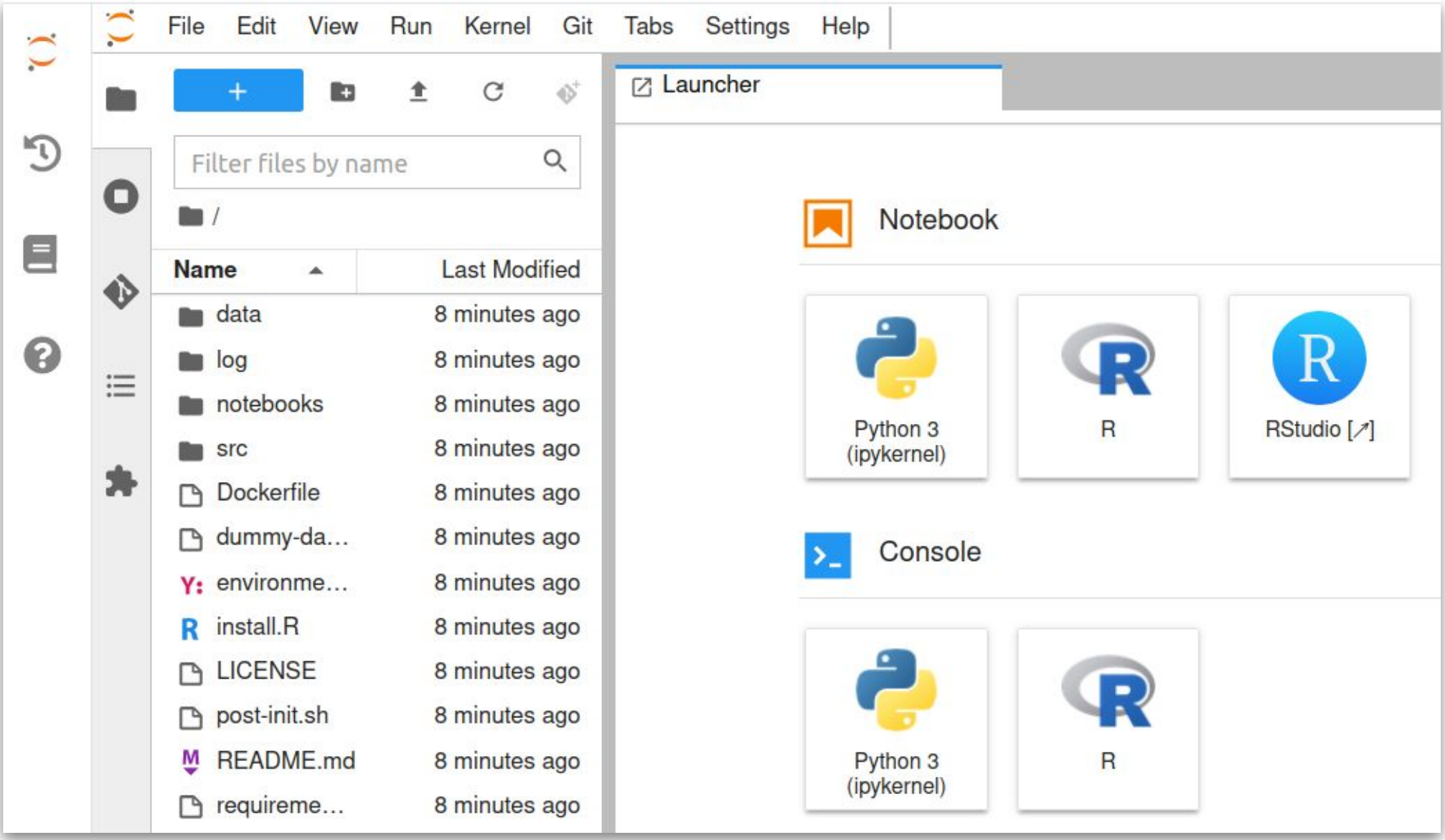
Method developer/
Benchmarker


bettr: A better way to explore what is best
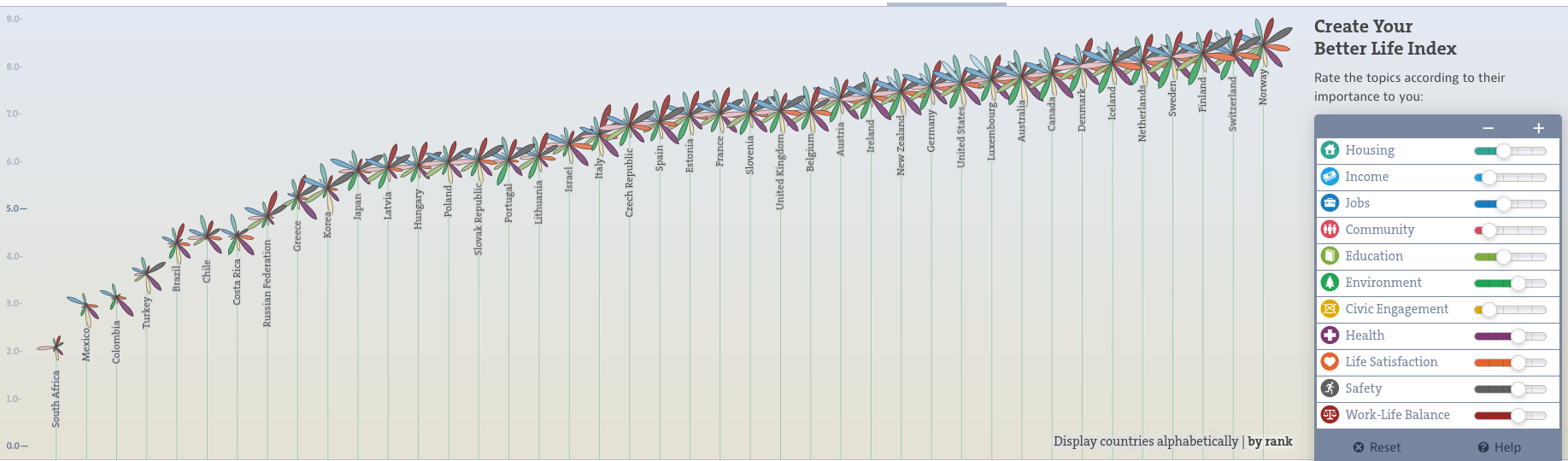
https://www.oecdbetterlifeindex.org
bettr: A better way to explore what is best
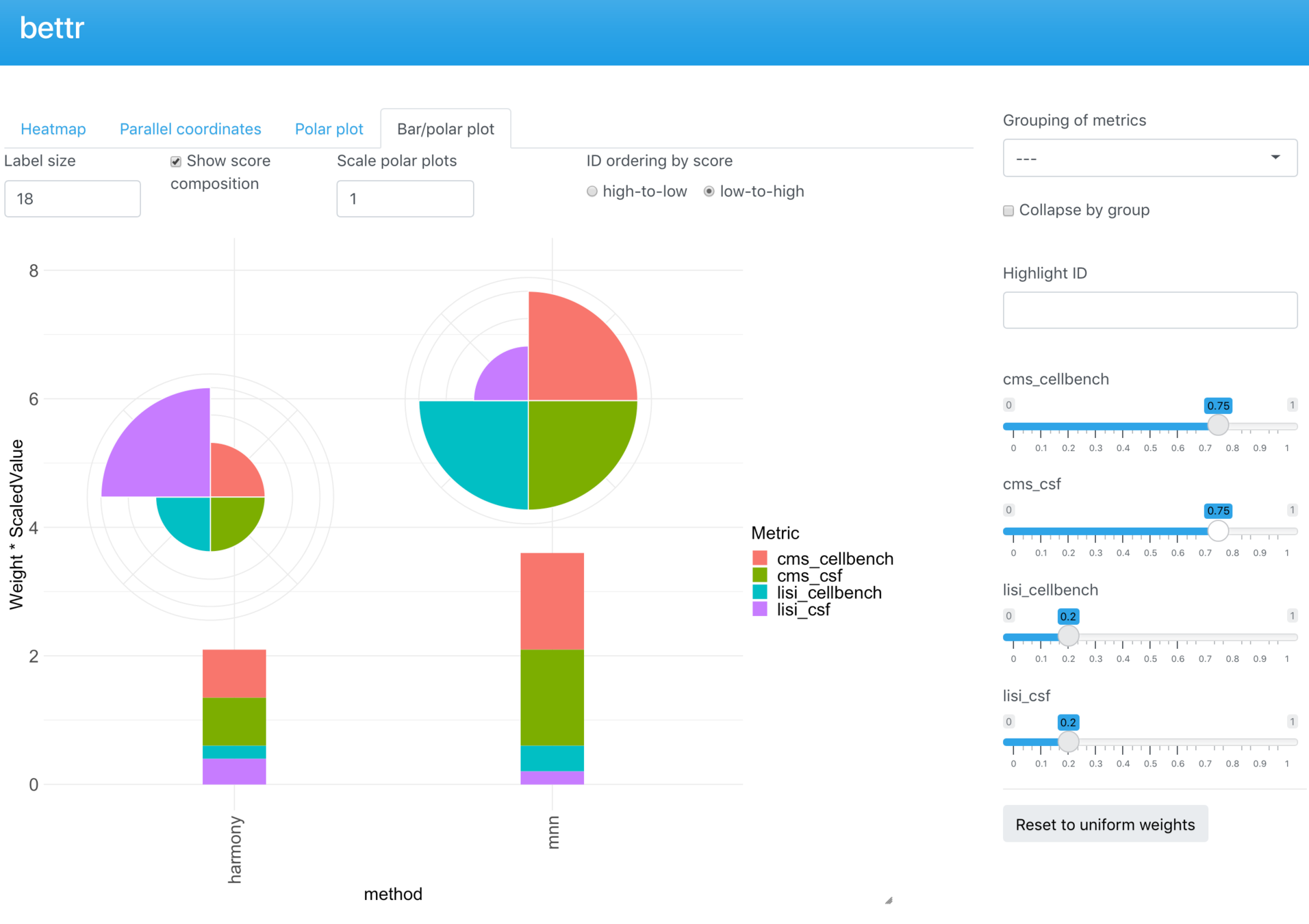
bettr exploration of prototype
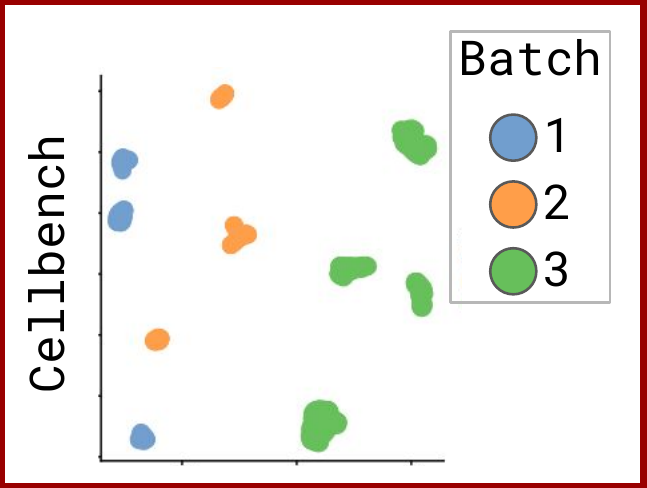
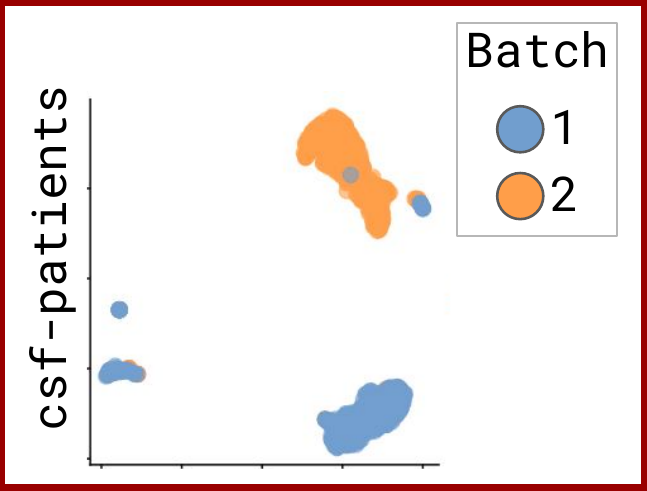
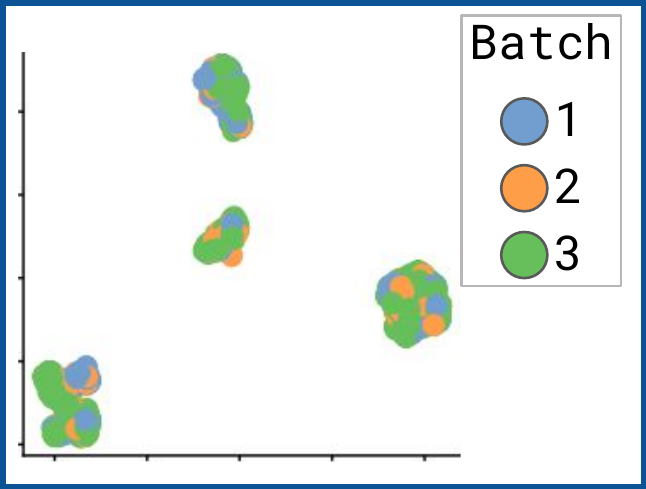
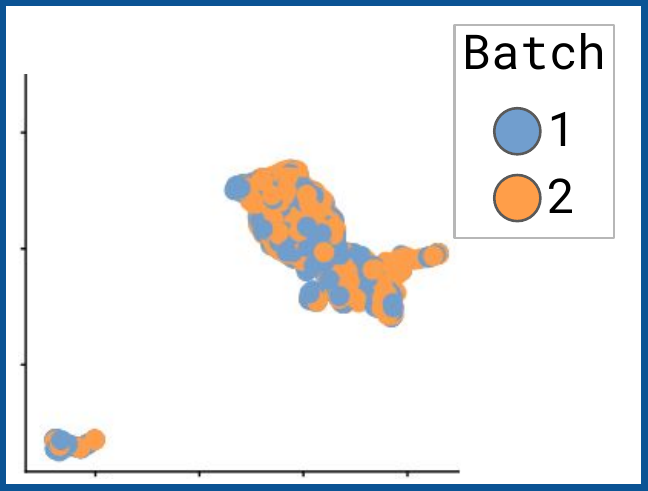
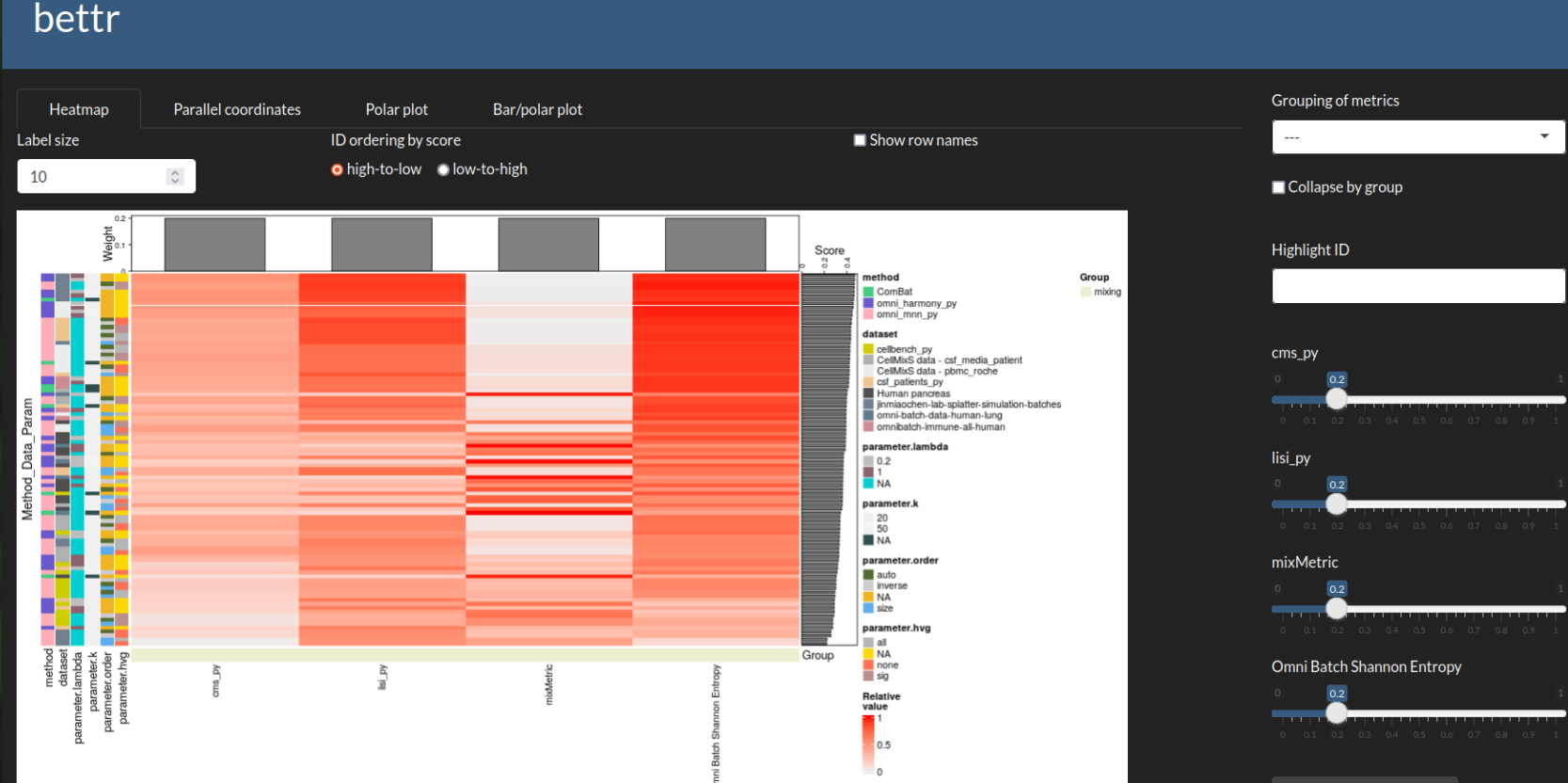
Omnibenchmark prototype
Thank you!

A data analysis platform/system built from a set of microservices
GitLab --> version control/CICD


Apache Jena --> Triple store


Jupyter server --> interactive sessions

Docker/Kubernetes --> software/enviroment management

GitLFS --> File storage
What is renku?

Renku client is a dataset and workflow management system
Renku client
-
Dataset and workflow management system → “renku-python”
-
Knowledge graph tracking → provenance
Renkulab
-
User interface with free interactive sessions
-
GitLab
Renku client is based on a Triplet store (Knowledge graph)

Result
Code
Data
generated
used_by
used_by
Data
Code
Result
used_by
generated
User interaction with renku client
Automatic triplet generation
Triplet store "Knowledge graph"
User interaction with renku client
KG-endpoint queries
module code
- Flexible language, code, etc.
- Minimal inputs and outputs are predefined depending on benchmark and module type! Check here!
- Automatic workflow generation, multiple parameter runs , updates etc.



Input types
output types
Omnibenchmark python module
from omnibenchmark import get_omni_object_from_yaml, renku save
## Load config
omni_obj = get_omni_object_from_yaml('src/config.yaml')
## Check for new/updates of input datasets
omni_obj.update_object()
renku_save()
## Create output dataset
omni_obj.create_dataset()
## Generate and run workflow
omni_obj.run_renku()
renku_save()
## Store results in output dataset
omni_obj.update_result_dataset()
renku_save()Methods
Metrics
Omnibenchmark design
Data
standardized datasets


= 1 "module" (renku project )



Methods
method results

Metrics
metric results

Dashboard
interactive result exploration
Method user
Method developer/
Benchmarker