Soss: Lightweight Probabilistic Programming in Julia

Chad Scherrer
Senior Data Scientist, Metis


About Me
Served as Technical Lead for language evaluation for the DARPA program
Probabilistic Programming for Advancing Machine Learning (PPAML)
PPL Publications
-
Scherrer, Diatchki, Erkök, & Sottile, Passage : A Parallel Sampler Generator for Hierarchical Bayesian Modeling, NIPS 2012 Workshop on Probabilistic Programming
-
Scherrer, An Exponential Family Basis for Probabilistic Programming, POPL 2017 Workshop on Probabilistic Programming Semantics
- Westbrook, Scherrer, Collins, and Mertens, GraPPa: Spanning the Expressivity vs. Efficiency Continuum, POPL 2017 Workshop on Probabilistic Programming Semantics
Making Rational Decisions
Managed Uncertainty
Rational Decisions
Bayesian Analysis
Probabilistic Programming
- Physical systems
- Hypothesis testing
- Modeling as simulation
- Medicine
- Finance
- Insurance
Risk
Custom models
Business Applications
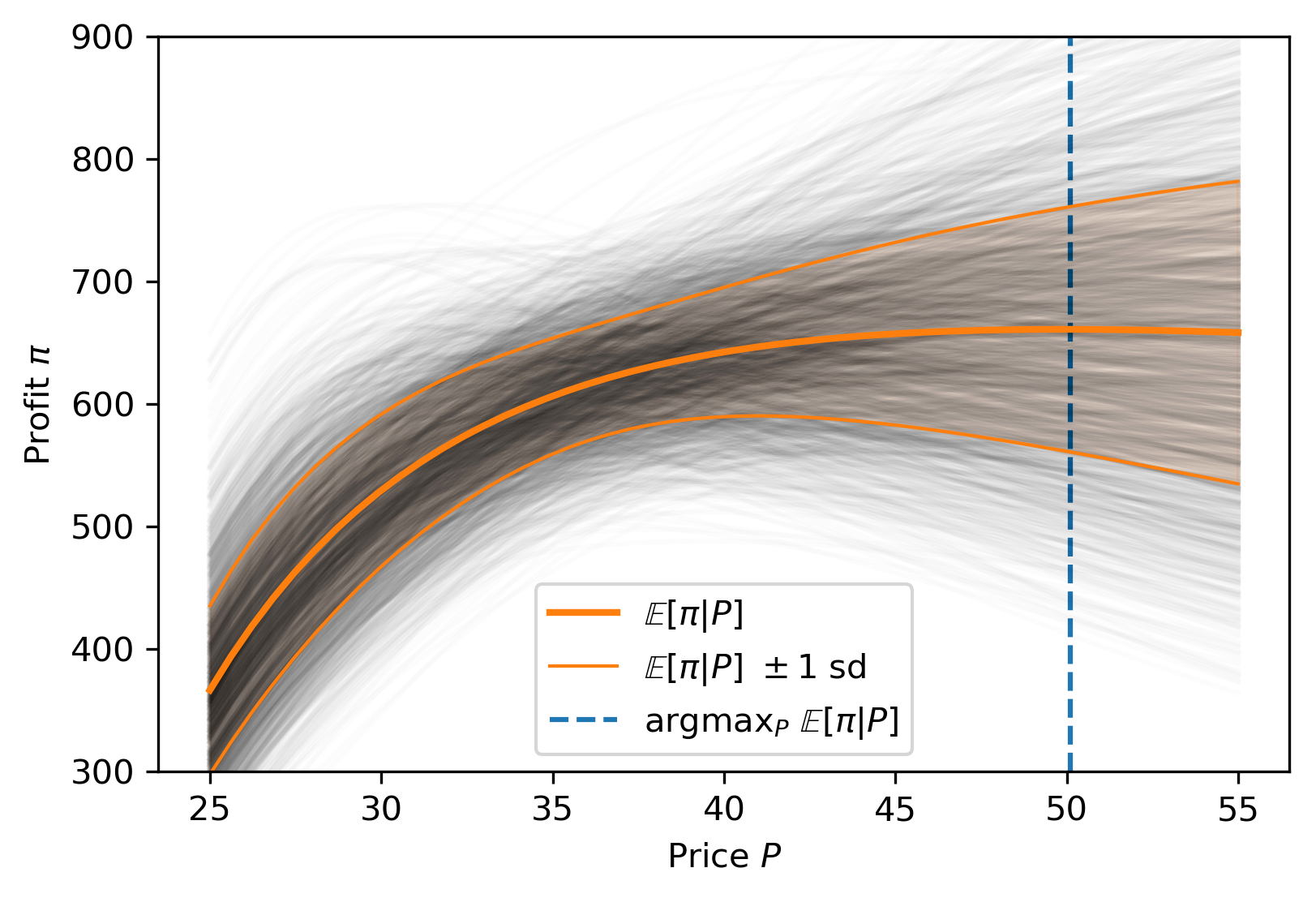
Missing Data






The Two-Language Problem
A disconnect between the "user language" and "developer language"
X
3
Python
C
Deep Learning Framework
- Harder for beginner users
- Barrier to entry for developers
- Limits extensibility
?
A New Approach in Julia
- Give an easy way to specify models
- Code generation for each (model, type of data, inference primitive)
- Composability
A (Very) Simple Example
P(\mu,\sigma|x)\propto P(\mu,\sigma)P(x|\mu,\sigma)
\begin{aligned}
\mu &\sim \text{Normal}(0,5)\\
\sigma &\sim \text{Cauchy}_+(0,3) \\
x_j &\sim \text{Normal}(\mu,\sigma)
\end{aligned}
Theory
Soss
julia> Soss.sourceLogpdf()(m)
quote
_ℓ = 0.0
_ℓ += logpdf(Normal(0, 5), μ)
_ℓ += logpdf(Normal(μ, σ), σ)
_ℓ += logpdf(Normal(μ, σ) |> iid(N), x)
return _ℓ
endm = @model N begin
μ ~ Normal(0,5)
σ ~ Normal(μ,σ)
x ~ Normal(μ,σ) |> iid(N)
end;A Simple Linear Model
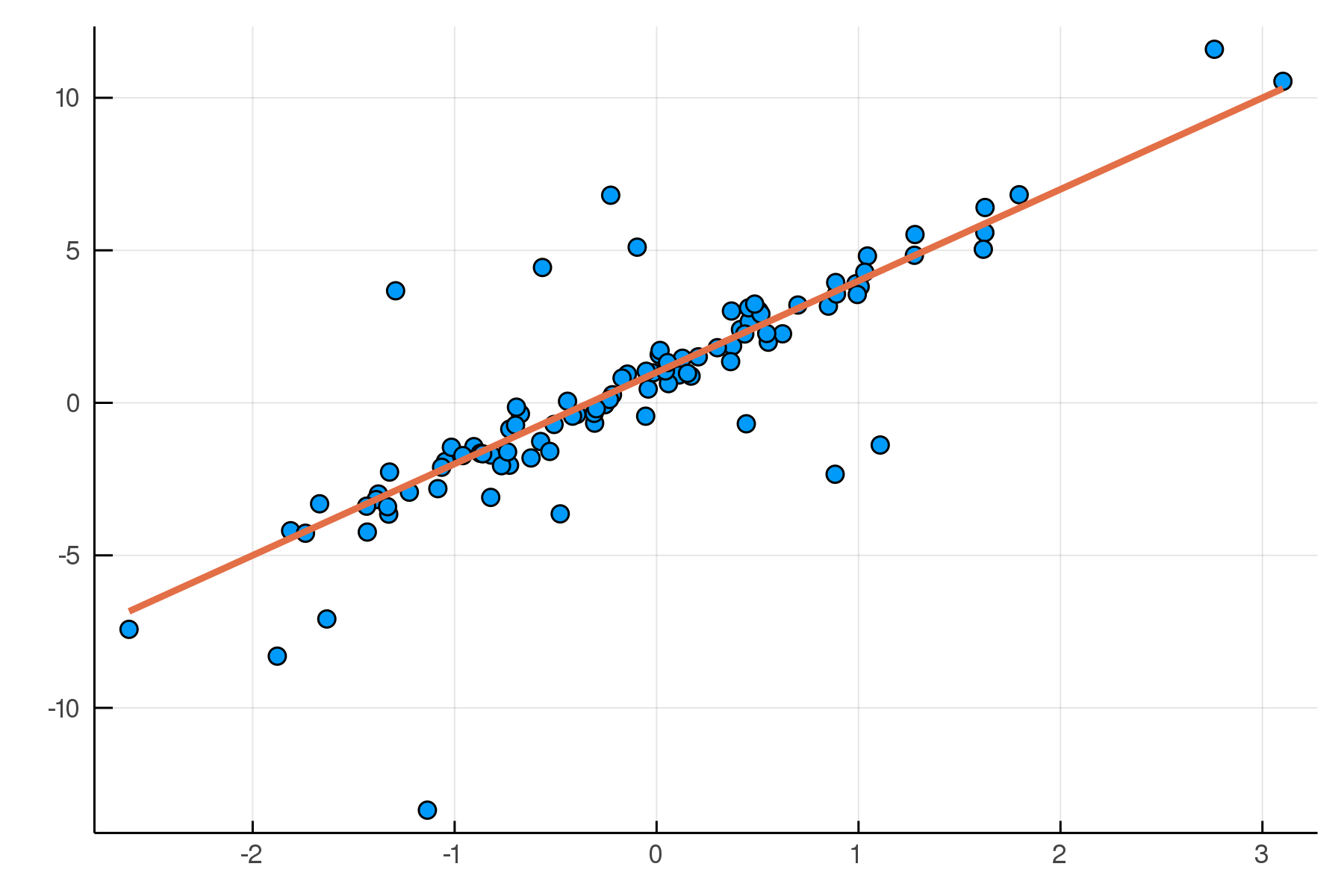
A Simple Linear Model
m = @model x begin
α ~ Cauchy()
β ~ Normal()
σ ~ HalfNormal()
yhat = α .+ β .* x
y ~ For(eachindex(x)) do j
Normal(yhat[j], σ)
end
endjulia> m(x=truth.x)
Joint Distribution
Bound arguments: [x]
Variables: [σ, β, α, yhat, y]
@model x begin
σ ~ HalfNormal()
β ~ Normal()
α ~ Cauchy()
yhat = α .+ β .* x
y ~ For(eachindex(x)) do j
Normal(yhat[j], σ)
end
endObserved data is not specified yet!
Sampling from the Posterior Distribution
julia> post = dynamicHMC(m(x=truth.x), (y=truth.y,)) |> particles
(σ = 2.02 ± 0.15, β = 2.99 ± 0.19, α = 0.788 ± 0.2)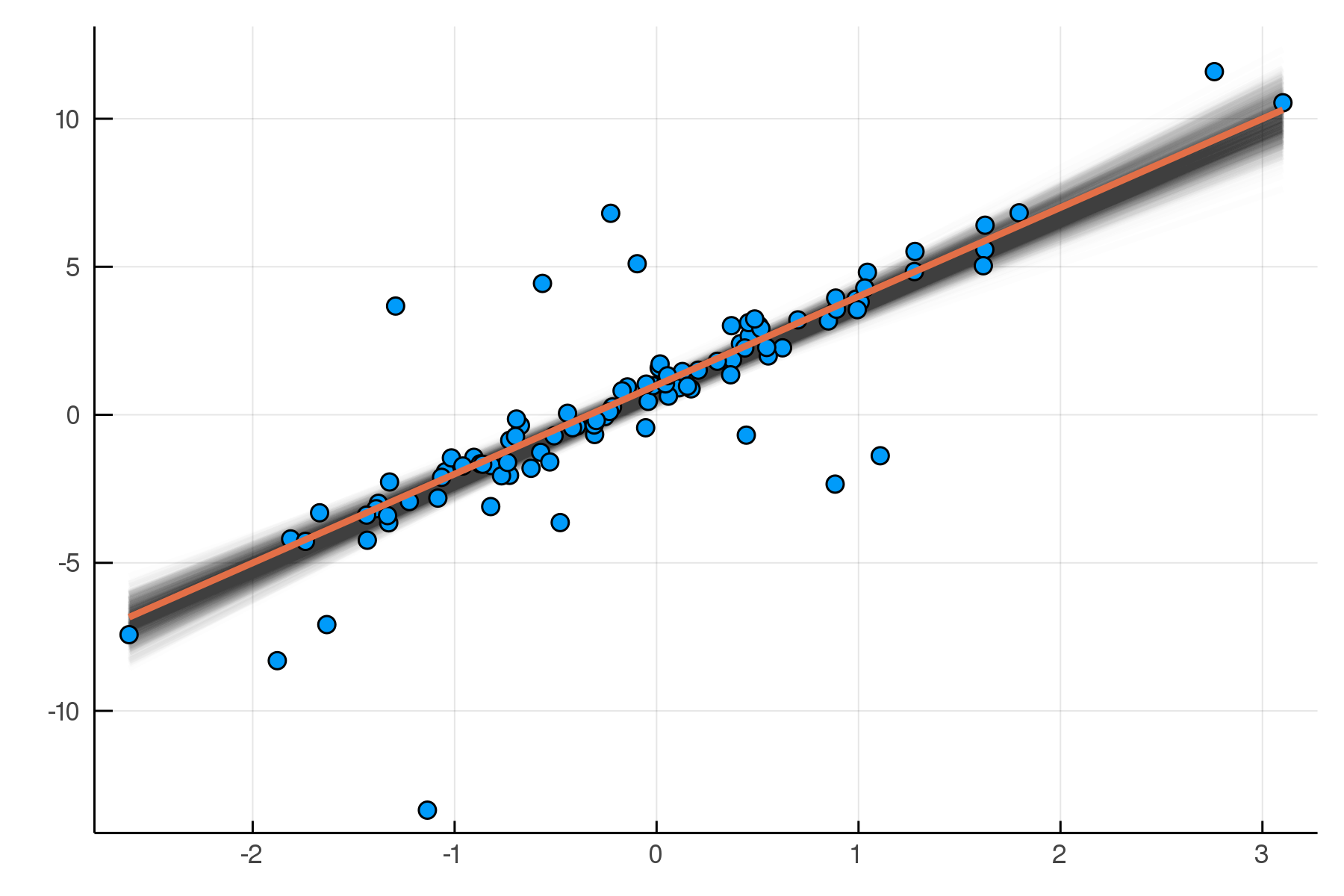
Posterior distribution
Possible best-fit lines
The Posterior Predictive Distribution
Start with Data
Sample Parameters|Data
Sample Data|Parameters
Real Data
Replicated Fake Data
Compare
Posterior
Distribution
Predictive
Distribution
The Posterior Predictive Distribution
julia> pred = predictive(m, :α, :β, :σ)
@model (x, α, β, σ) begin
yhat = α .+ β .* x
y ~ For(eachindex(x)) do j
Normal(yhat[j], σ)
end
endm = @model x begin
α ~ Cauchy()
β ~ Normal()
σ ~ HalfNormal()
yhat = α .+ β .* x
y ~ For(eachindex(x)) do j
Normal(yhat[j], σ)
end
endpostpred = [pred(θ)((x=x,)) for θ ∈ post] .|> rand |> particlespredictive makes a new model!
posterior predictive distributions
draw samples
convert to particles
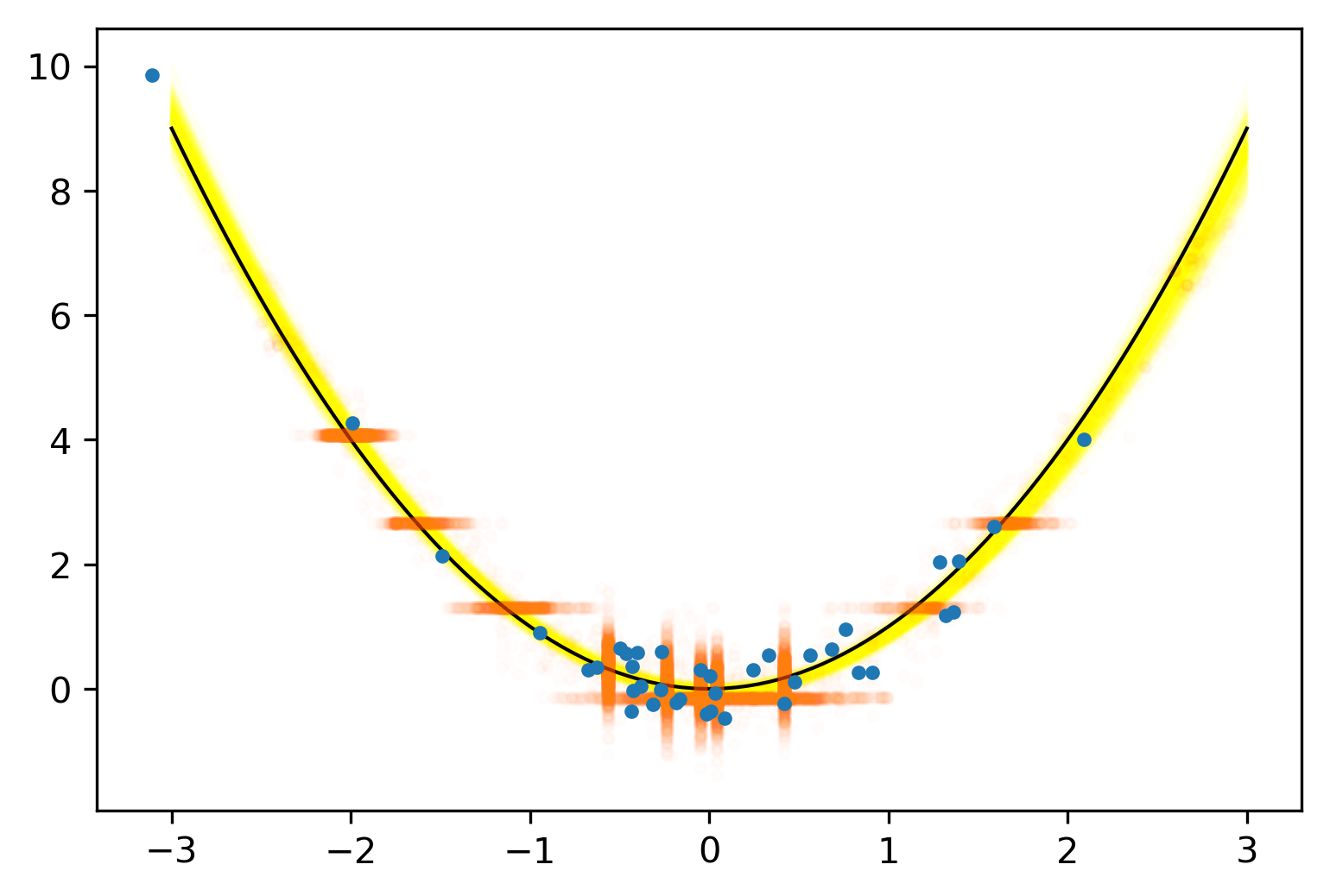
Posterior Predictive Checks
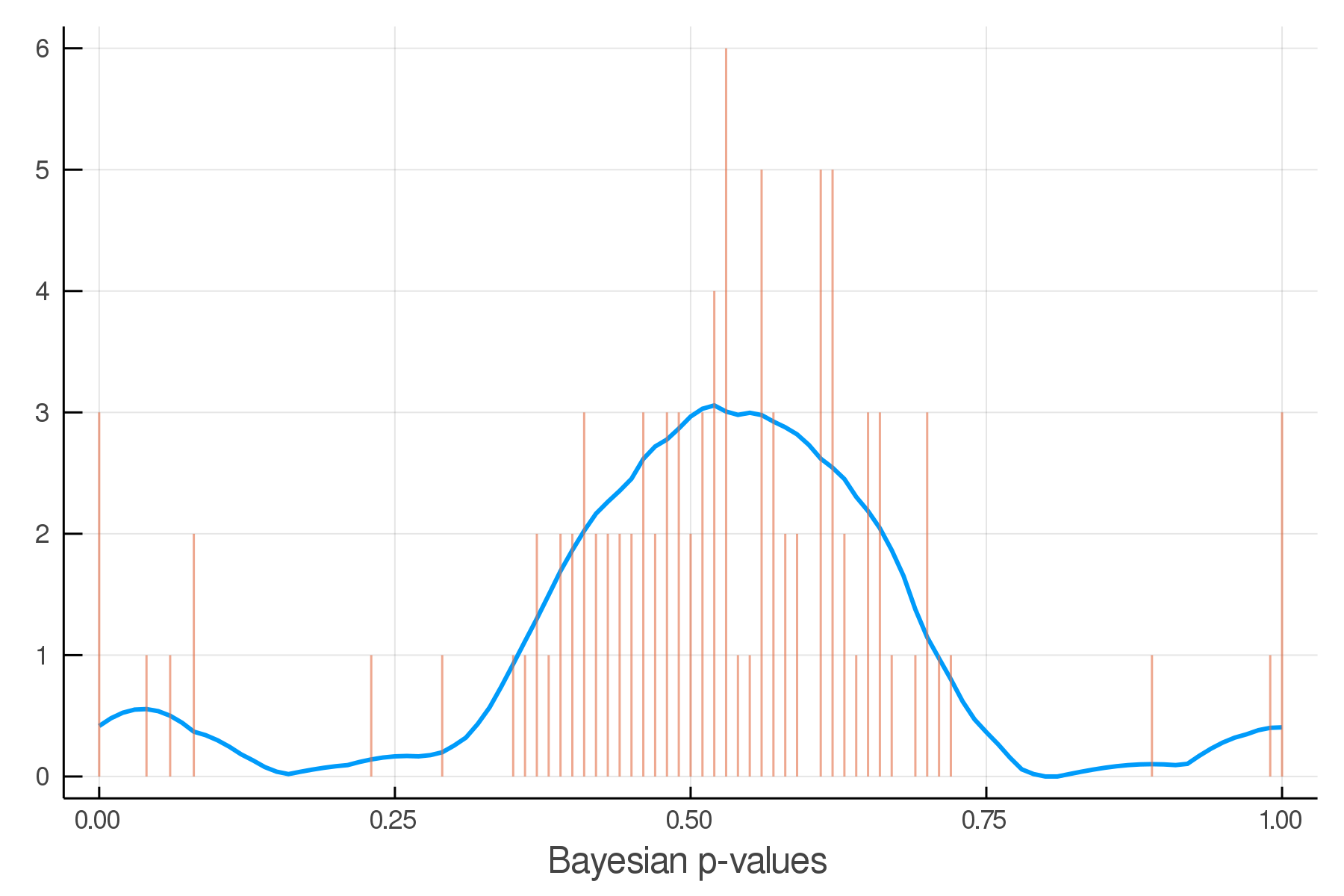
pvals = mean.(truth.y .> postpred.y)Where we expect the data
Where we see the data
Making it Robust
m2 = @model x begin
α ~ Cauchy()
β ~ Normal()
σ ~ HalfNormal(10)
νinv ~ HalfNormal()
yhat = α .+ β .* x
y ~ For(eachindex(x)) do j
StudentT(1/νinv,yhat[j],σ)
end
end;julia> post2 = dynamicHMC(m2(x=truth.x), (y=truth.y,)) |> particles
(σ = 0.444 ± 0.065, νinv = 0.807 ± 0.15
, β = 3.07 ± 0.064, α = 0.937 ± 0.062)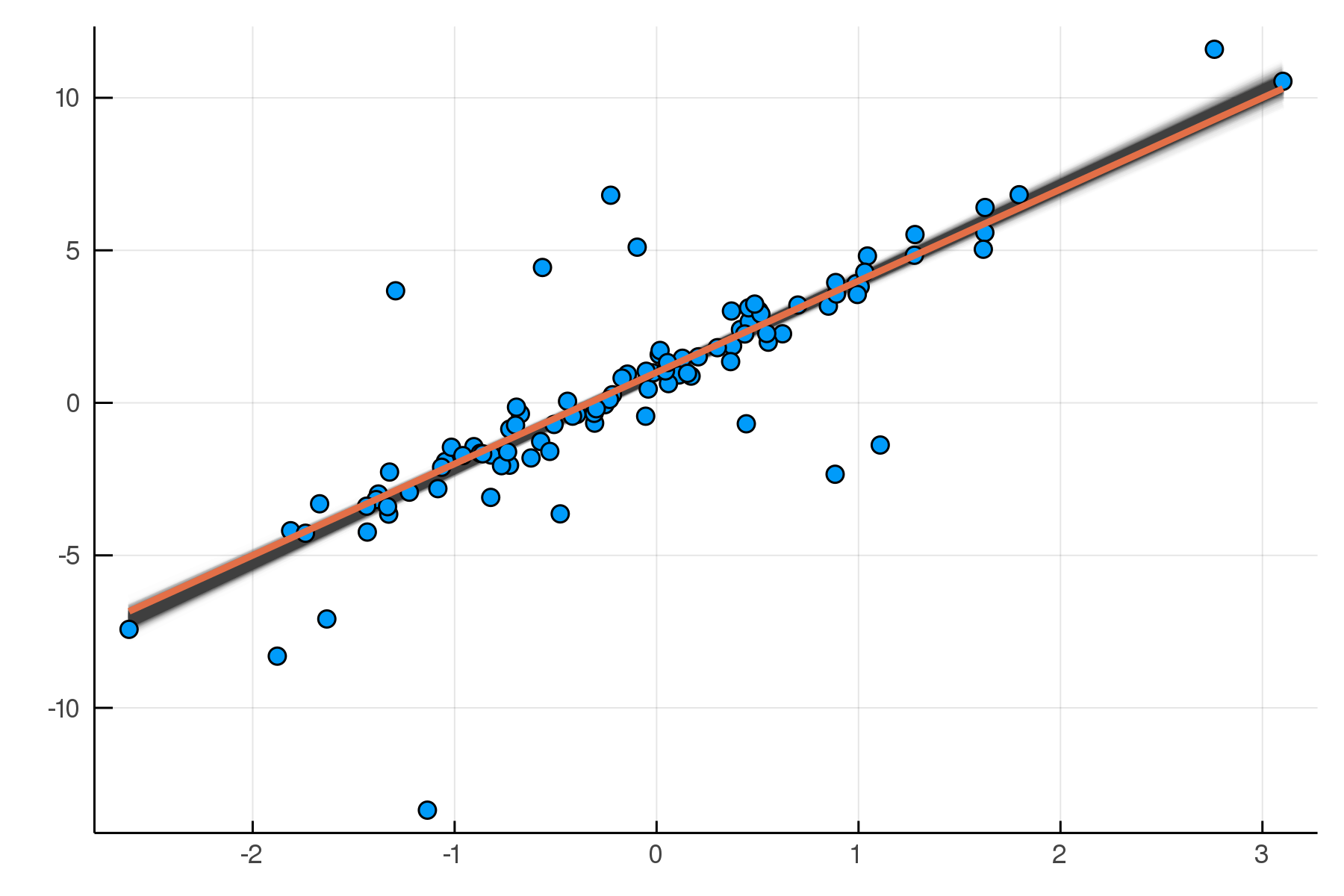
Updated Posterior Predictive Checks
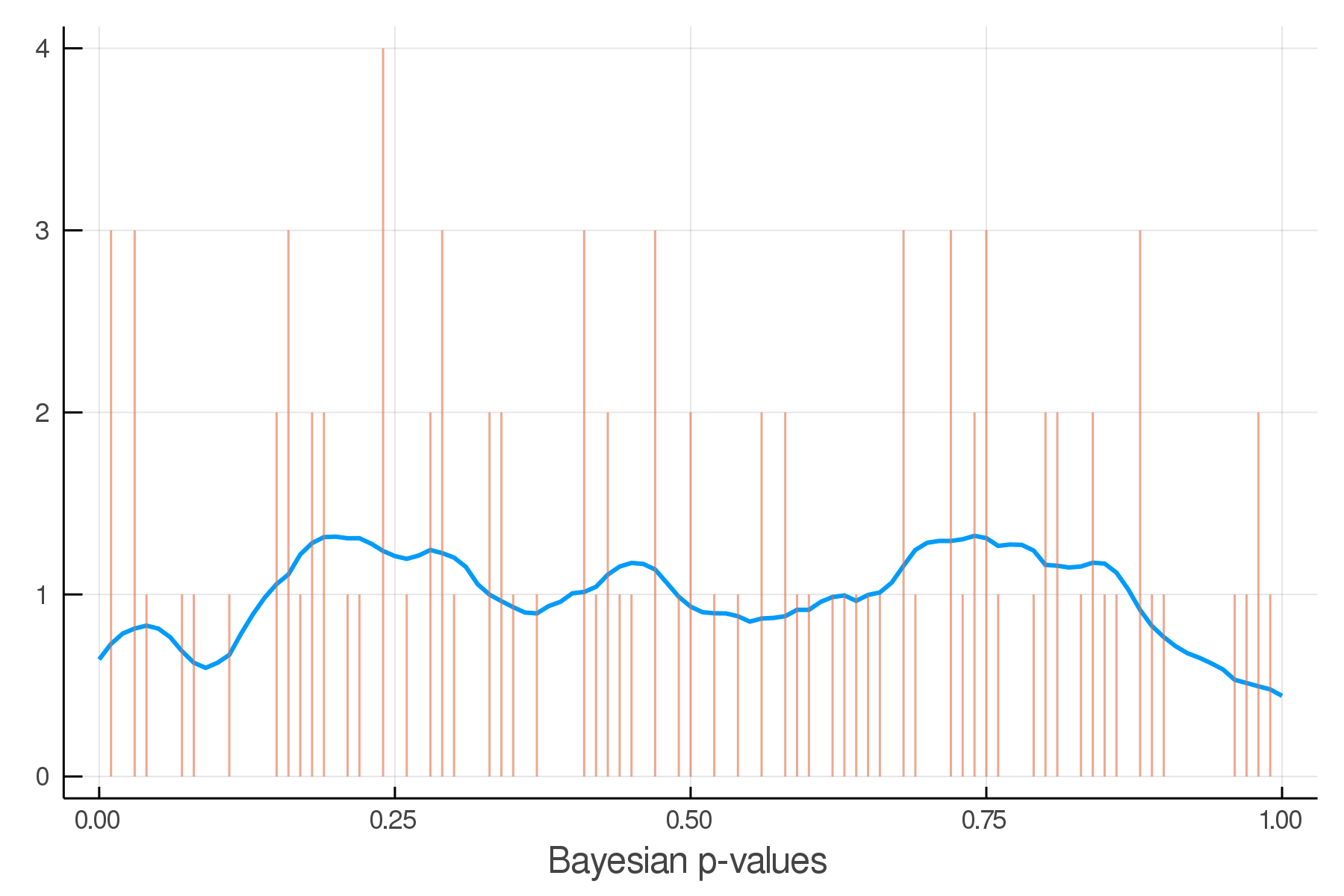

Architecture Overview
Inference Primitives
- xform, rand, particles, logpdf, weightedSample
Inference Algorithms
- stream(sampler, myModel(args), data)
Stream Combinators
- Rejection sampling
- Approximate Bayes (ABC)
- Expectation
Inference Primitive: rand
julia> Soss.sourceRand()(m)
quote
σ = rand(HalfNormal())
β = rand(Normal())
α = rand(Cauchy())
yhat = α .+ β .* x
y = rand(For(((j,)->begin
Normal(yhat[j], σ)
end), eachindex(x)))
(x = x, yhat = yhat, α = α
, β = β, σ = σ, y = y)
endm = @model x begin
α ~ Cauchy()
β ~ Normal()
σ ~ HalfNormal()
yhat = α .+ β .* x
y ~ For(eachindex(x)) do j
Normal(yhat[j], σ)
end
endInference Primitive: logpdf
julia> Soss.sourceLogpdf()(m)
quote
_ℓ = 0.0
_ℓ += logpdf(HalfNormal(), σ)
_ℓ += logpdf(Normal(), β)
_ℓ += logpdf(Cauchy(), α)
yhat = α .+ β .* x
_ℓ += logpdf(For(eachindex(x)) do j
Normal(yhat[j], σ)
end, y)
return _ℓ
endm = @model x begin
α ~ Cauchy()
β ~ Normal()
σ ~ HalfNormal()
yhat = α .+ β .* x
y ~ For(eachindex(x)) do j
Normal(yhat[j], σ)
end
endFirst-Class Models
julia> m = @model begin
a ~ @model begin
x ~ Normal()
end
end;
julia> rand(m())
(a = (x = -0.20051706307697828,),)julia> m2 = @model anotherModel begin
y ~ anotherModel
z ~ anotherModel
w ~ Normal(y.a.x / z.a.x, 1)
end;
julia> rand(m2(anotherModel=m)).w
-1.822683102320004Coming Soon
\sum_j\left(a + b x_j\right) \rightarrow Na + b \sum_j x_j
- Stream combinators via Transducers.jl and OnlineStats.jl
- Connection to other PPLs: Turing.jl, Gen.jl
- Normalizing flows with Bijectors.jl
- Deep learning with Flux.jl
- Gaussian processes
- Symbolic simplification
Thank You!

Special Thanks for