Towards an open source "Google Maps"-like interface for neuroanatomical data
Daniel Fürth
Meletis Lab
Lee lab meeting
22th July 2016
daniel.furth@ki.se

Towards an open source "Google Maps"-like interface for neuroanatomical data
Daniel Fürth
Meletis Lab
Zador lab meeting
11th April 2016
daniel.furth@ki.se
Content:
- Neuroanatomical Information System (NIS)
- Input to serotonergic raphe nuclei
- Cortico-striatal networks
- Scalable monosynaptic architecture
- Spatial mapping of single-cell RNAseq data to tissue of origin
- Multiplexed padlock probes
Neuroanatomical Information System (NIS)

Geographical Information System (GIS)
Web-interface



similar to...
works with...
Surface primitives
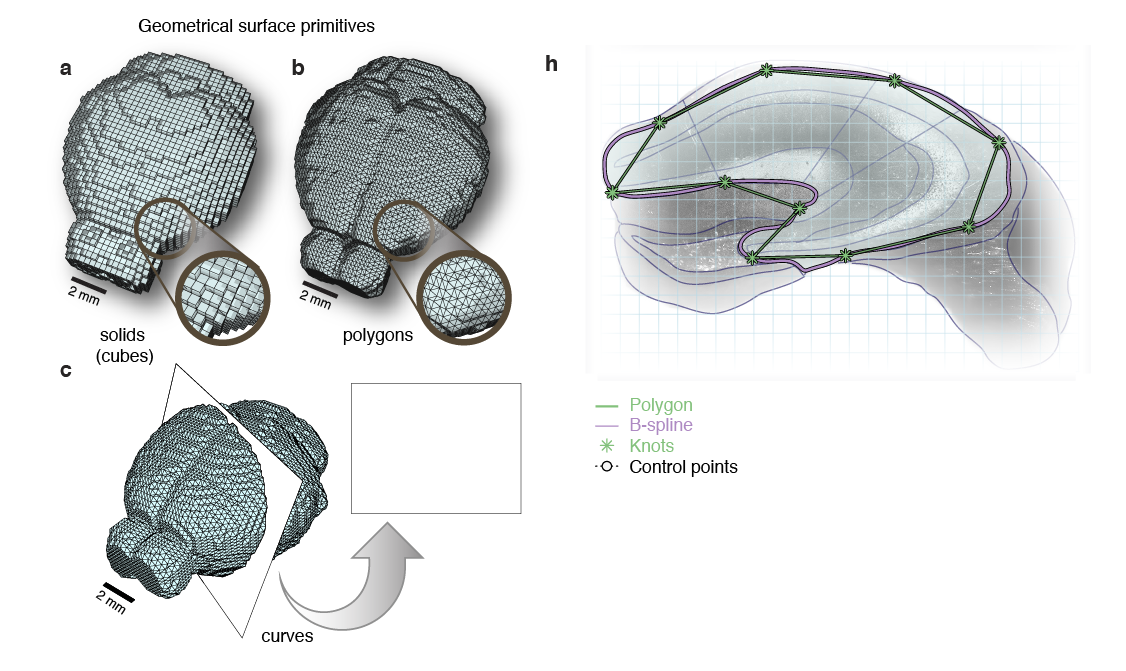
Non-Uniform Rational B-Splines (NURBS)
Surface primitives
Surface primitives

www.wholebrainsoftware.org
Tracing the network

Reconstructing brain from sectioned tissue
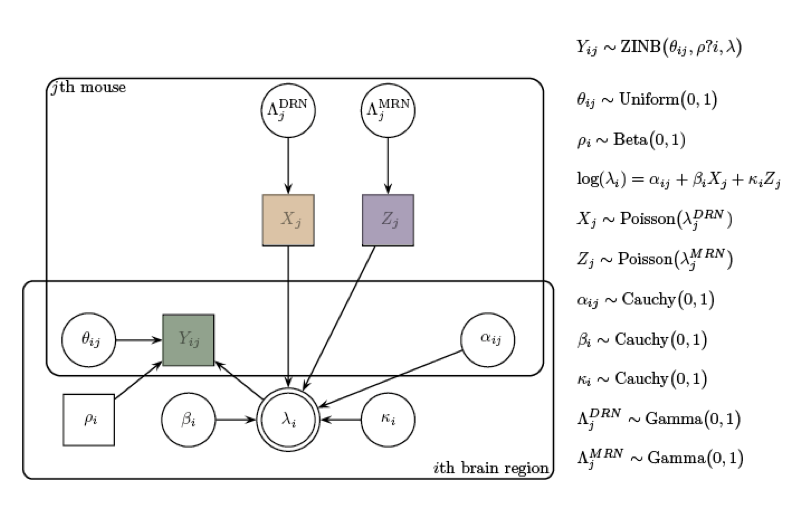
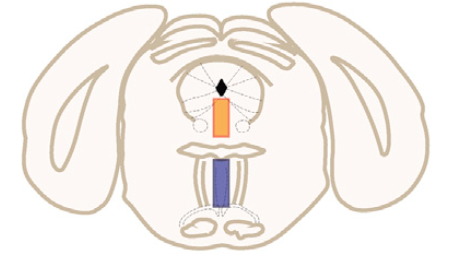
Pollak et al. 2014
www.mcstan.org

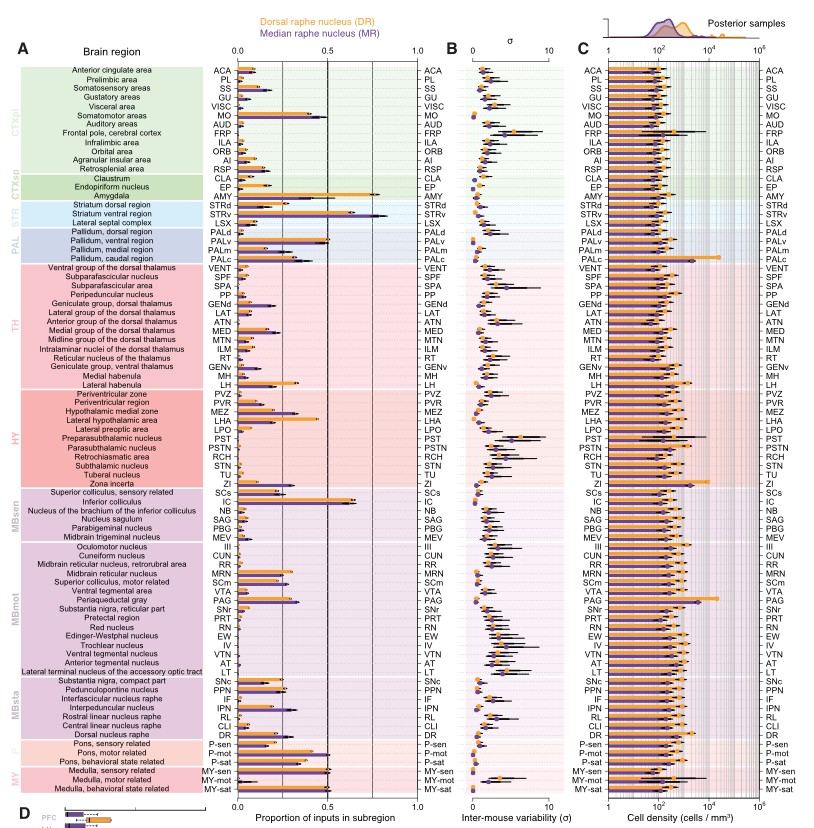
Whole-Brain Reconstruction
Pollak Dorocic et al. 2014


understanding behavior
striatal models
Graybiel, A. et al. (1990) PNAS
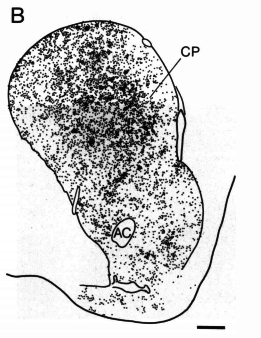

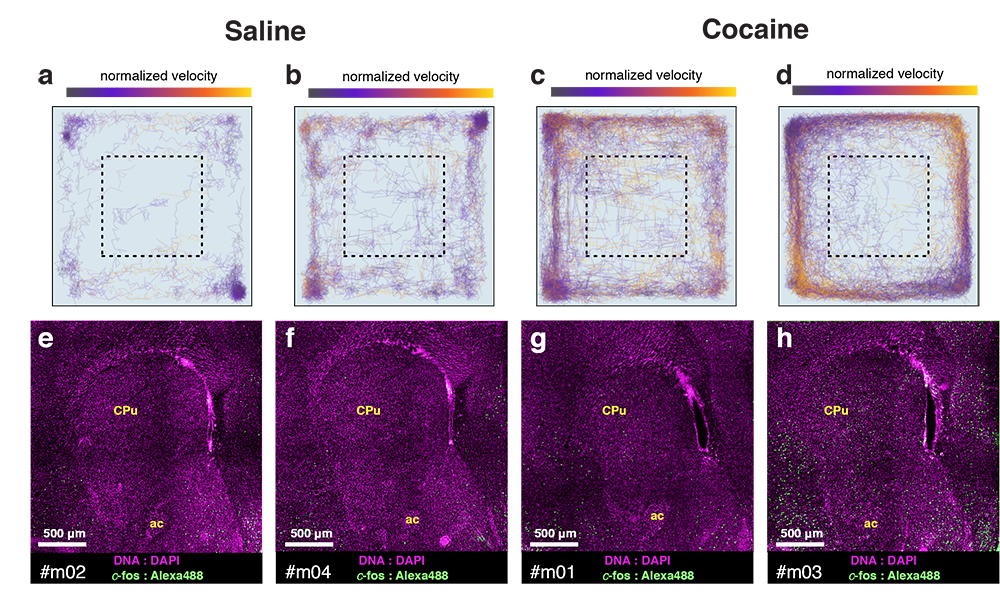
cfos
cocaine
saline
striatal models
Albin, Young & Penney, 1989
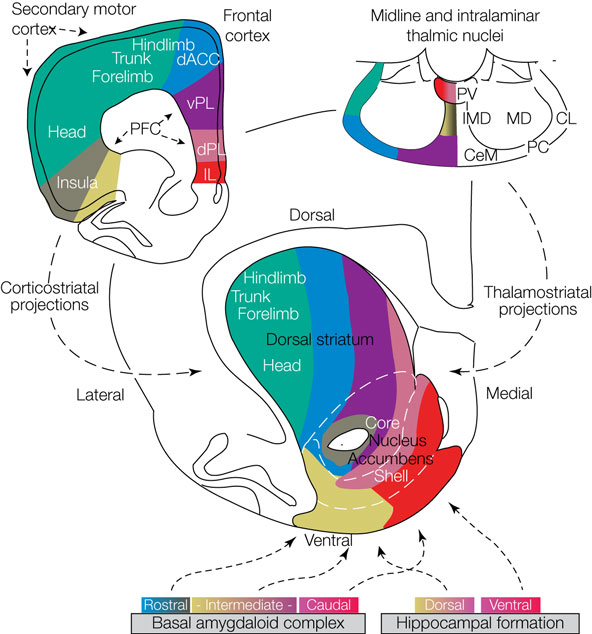

Alexander, DeLong & Stric 1986
striatal models
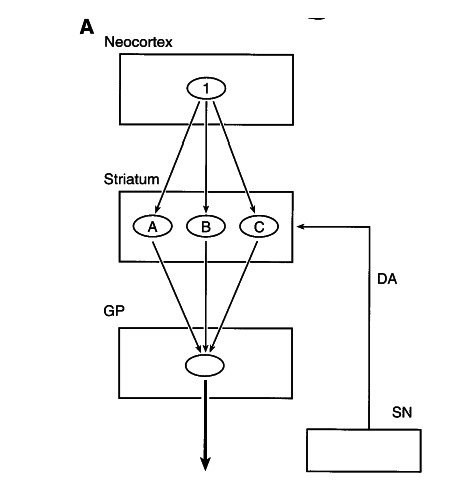
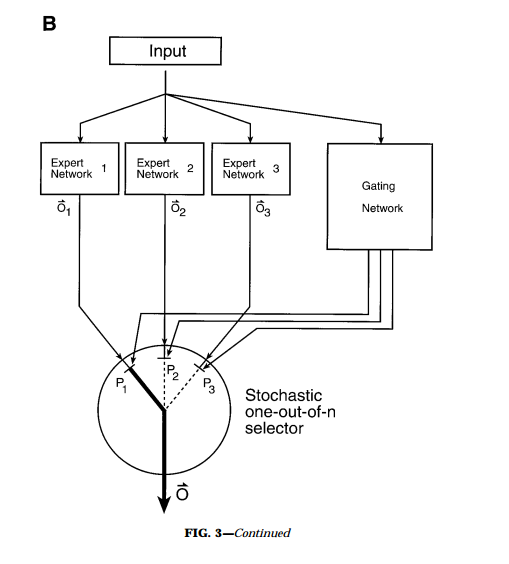
Graybiel, A. et al. (1990)
Jordan, M. I. et. al. (1991). Adaptive mixtures of local experts.
Tracing the network
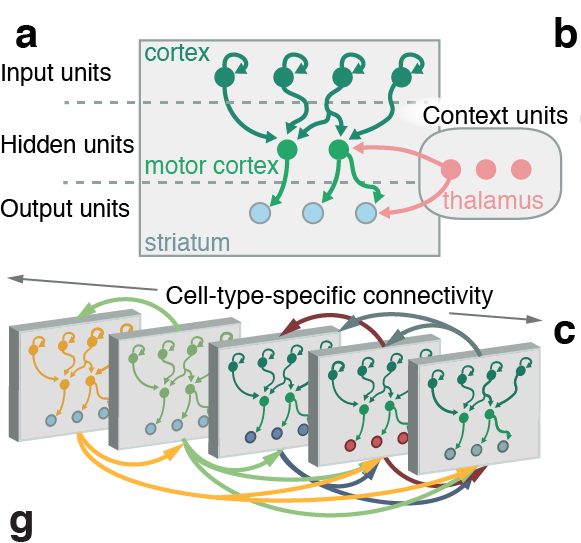
Tracing the network
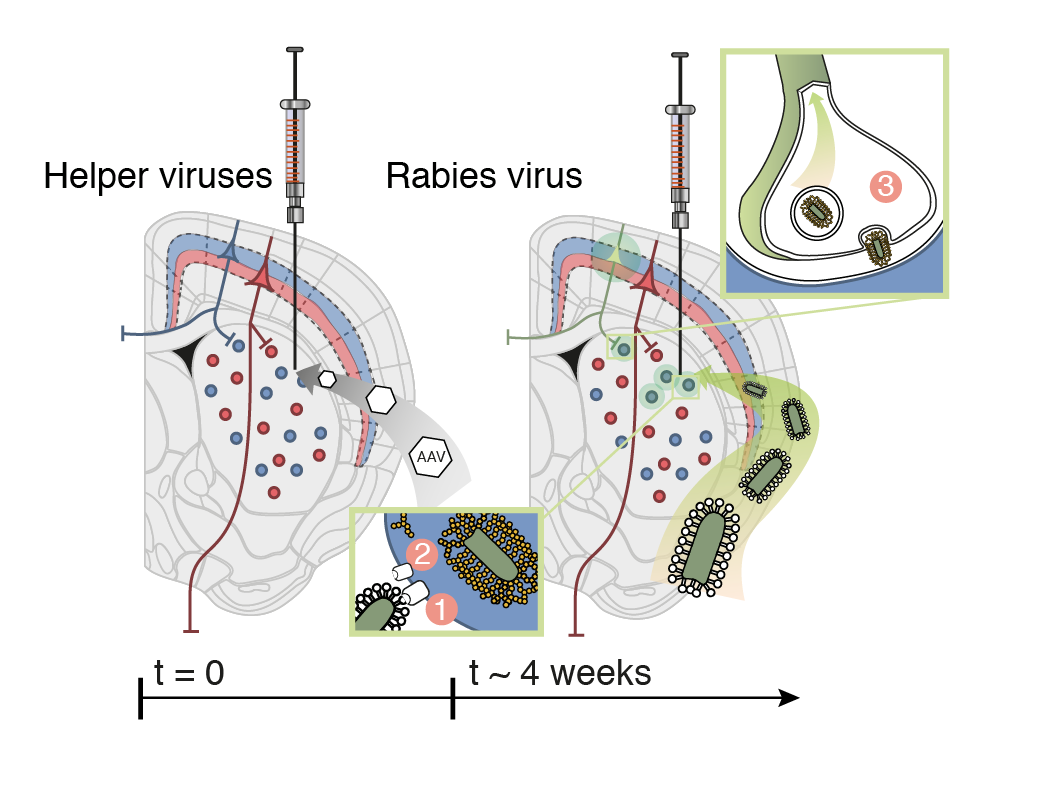
DRD2 film
56,710 neurons
Tracing the network

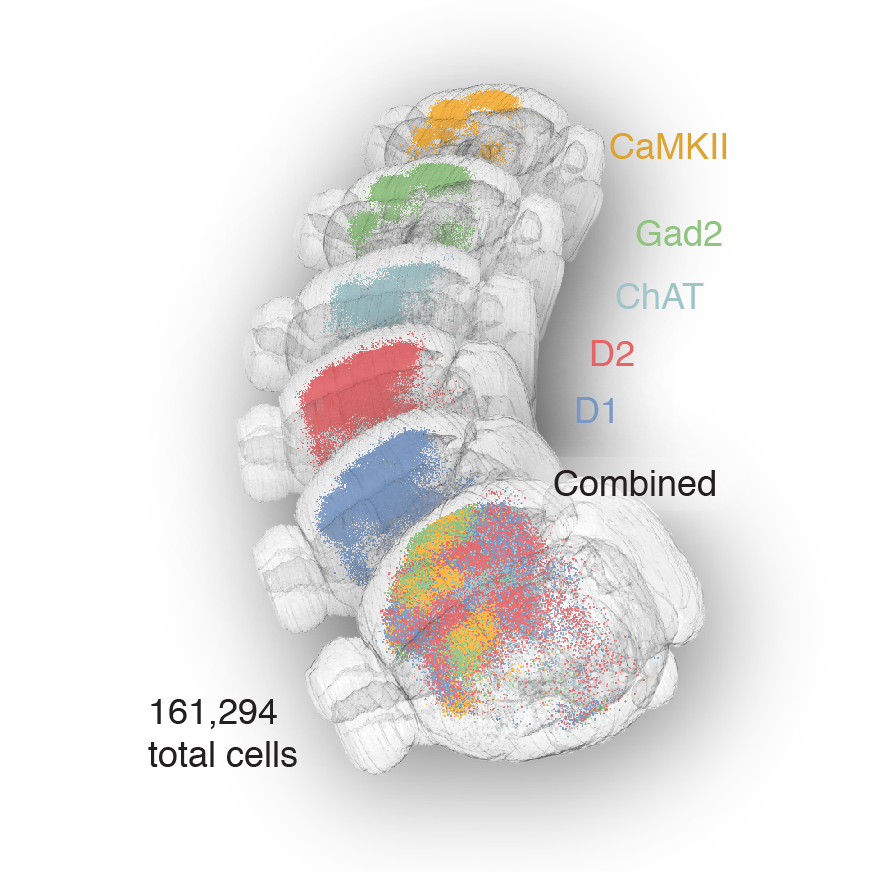
161,294 pooled neurons
Tracing the network
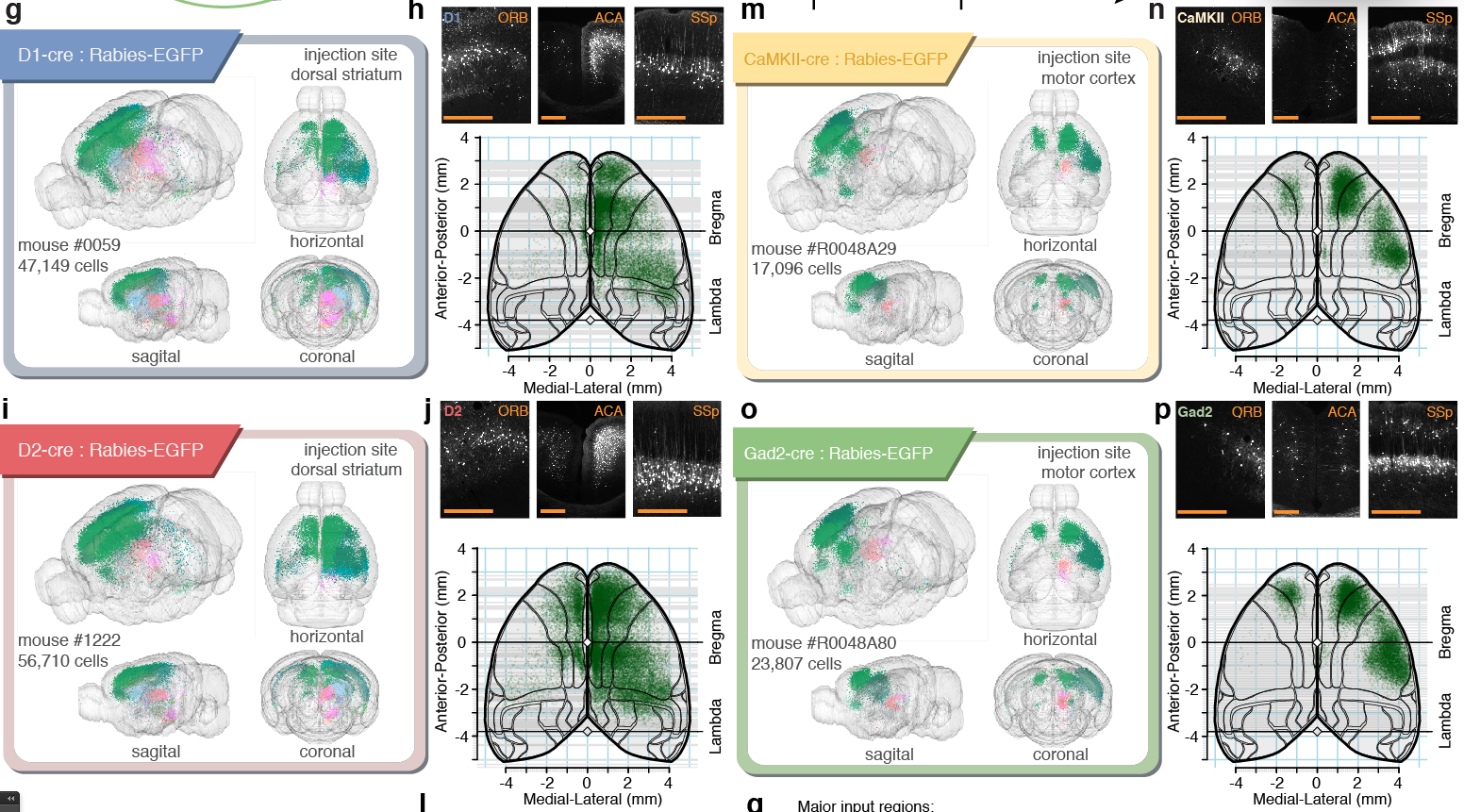
Tracing the network
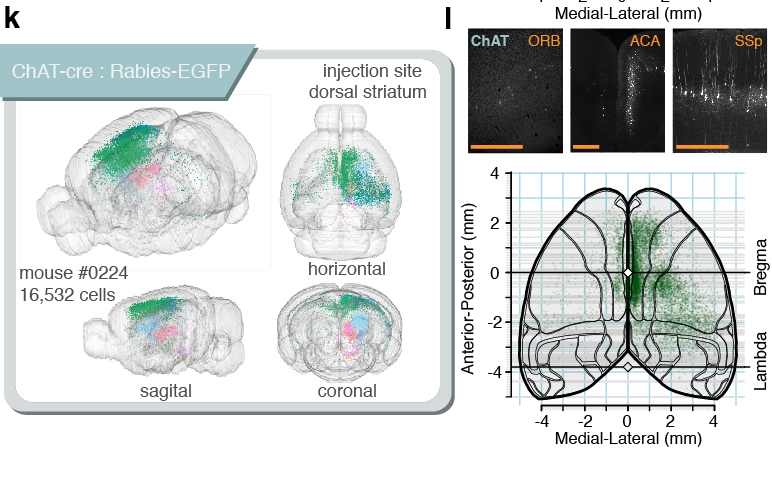
Tracing the network
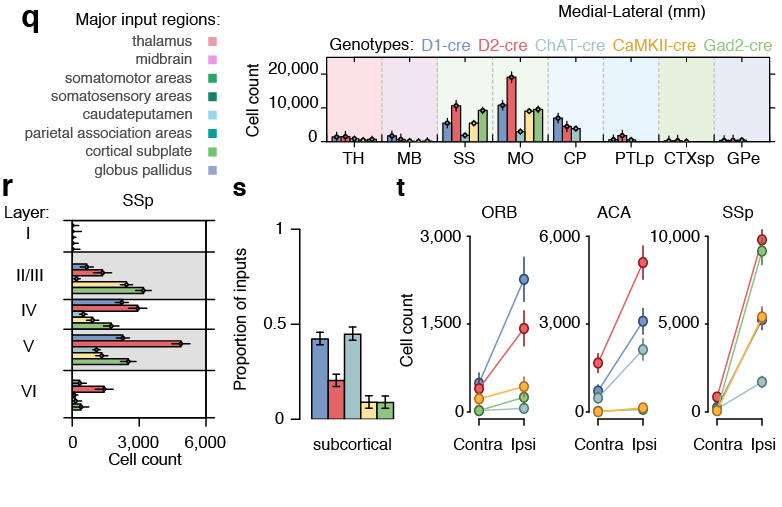
Cocaine induced locomotoric activity
Multiresolution decomposition
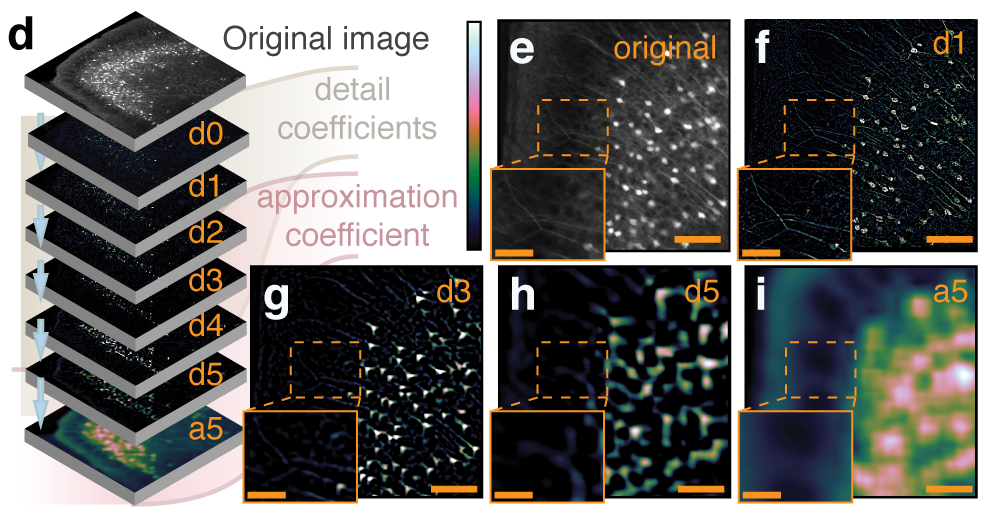
Multiresolution decomposition
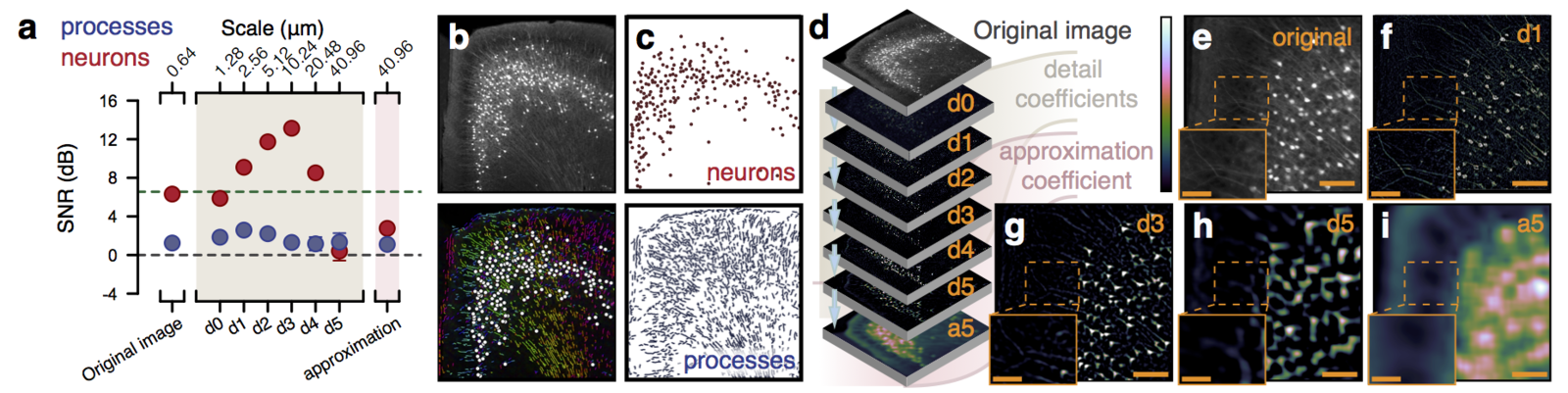
Multiresolution decomposition
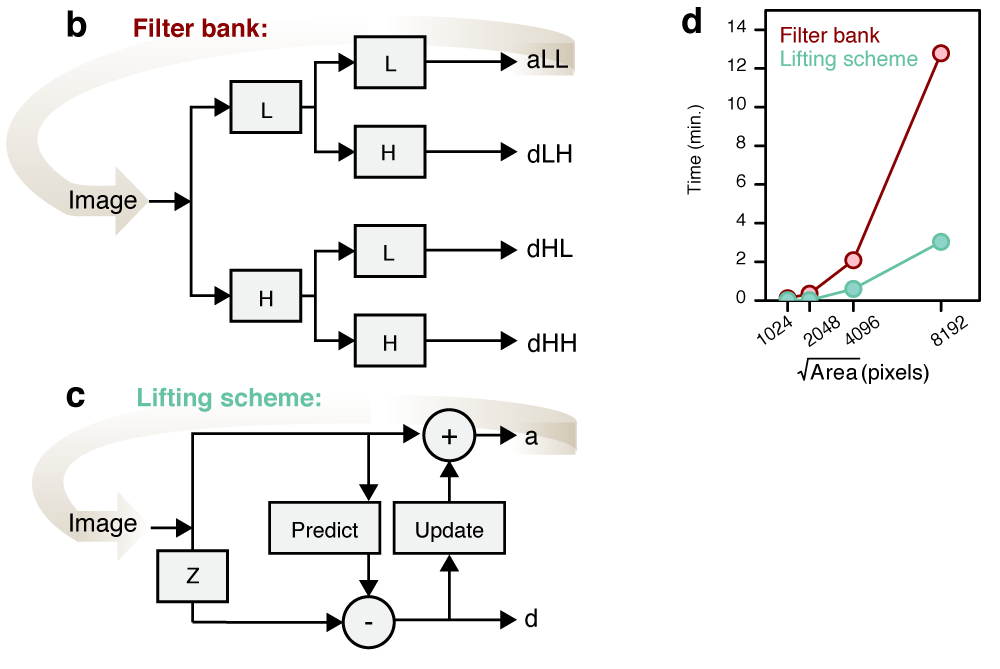
Performance
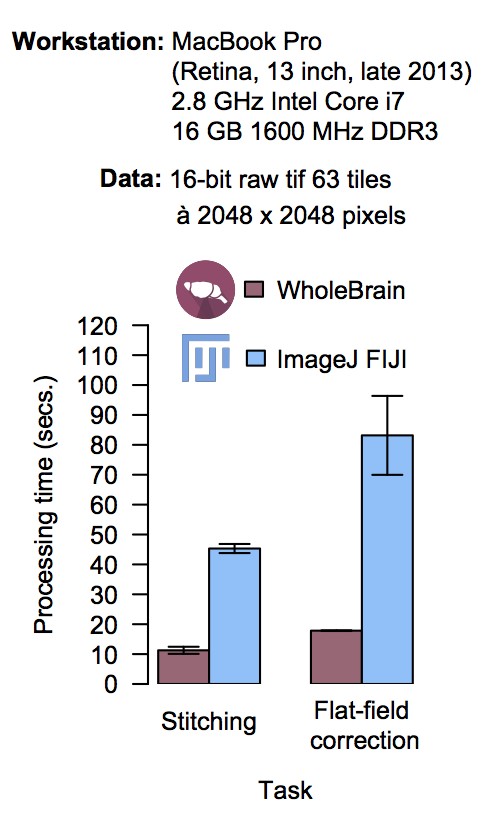
Multiresolution decomposition
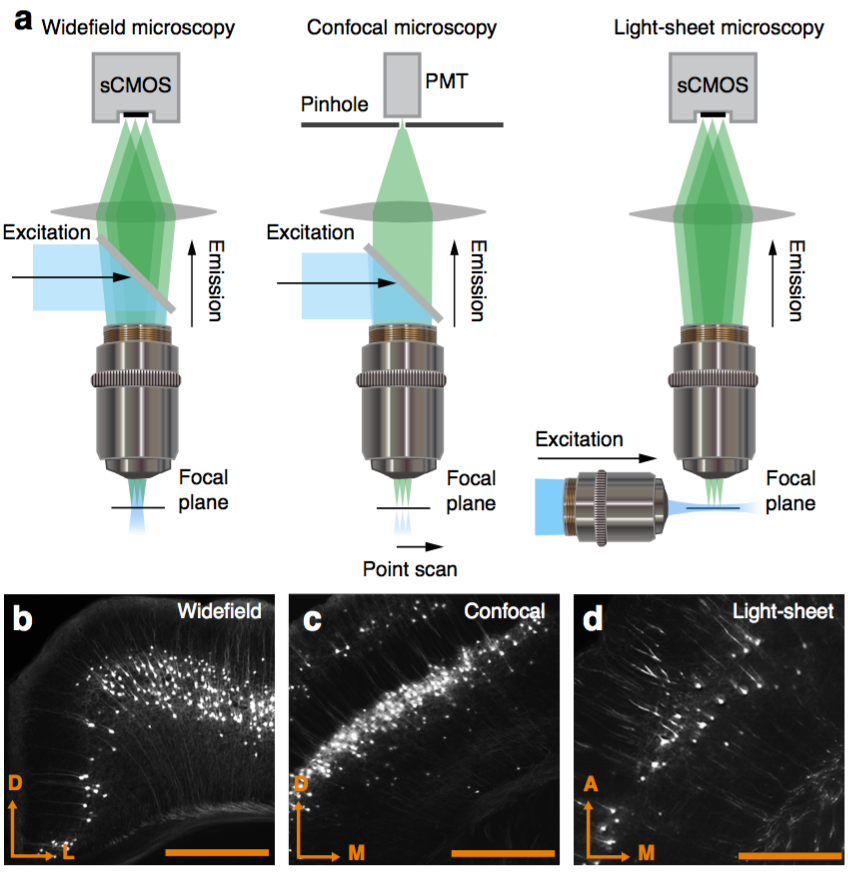
Multiresolution decomposition
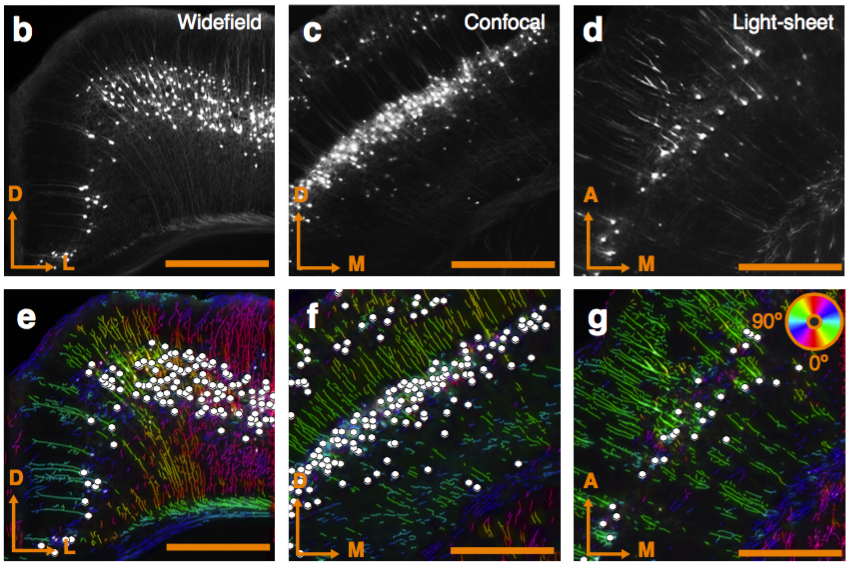
Multiresolution decomposition
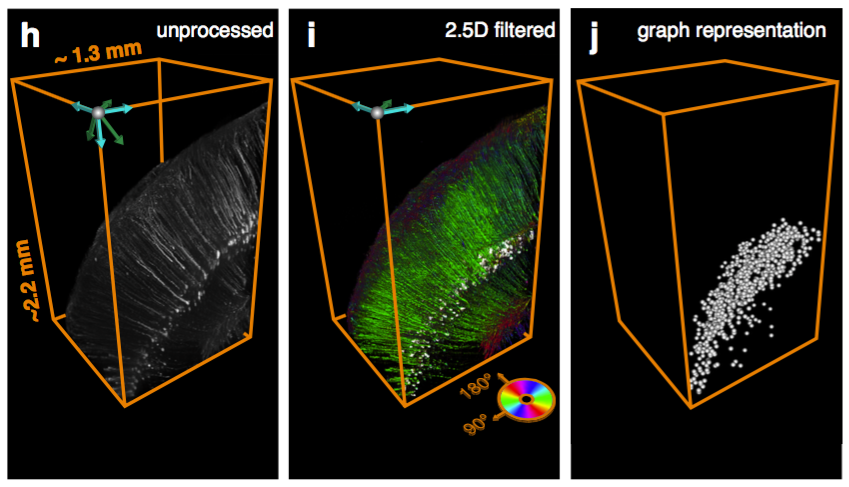
Segmentation of nuclei
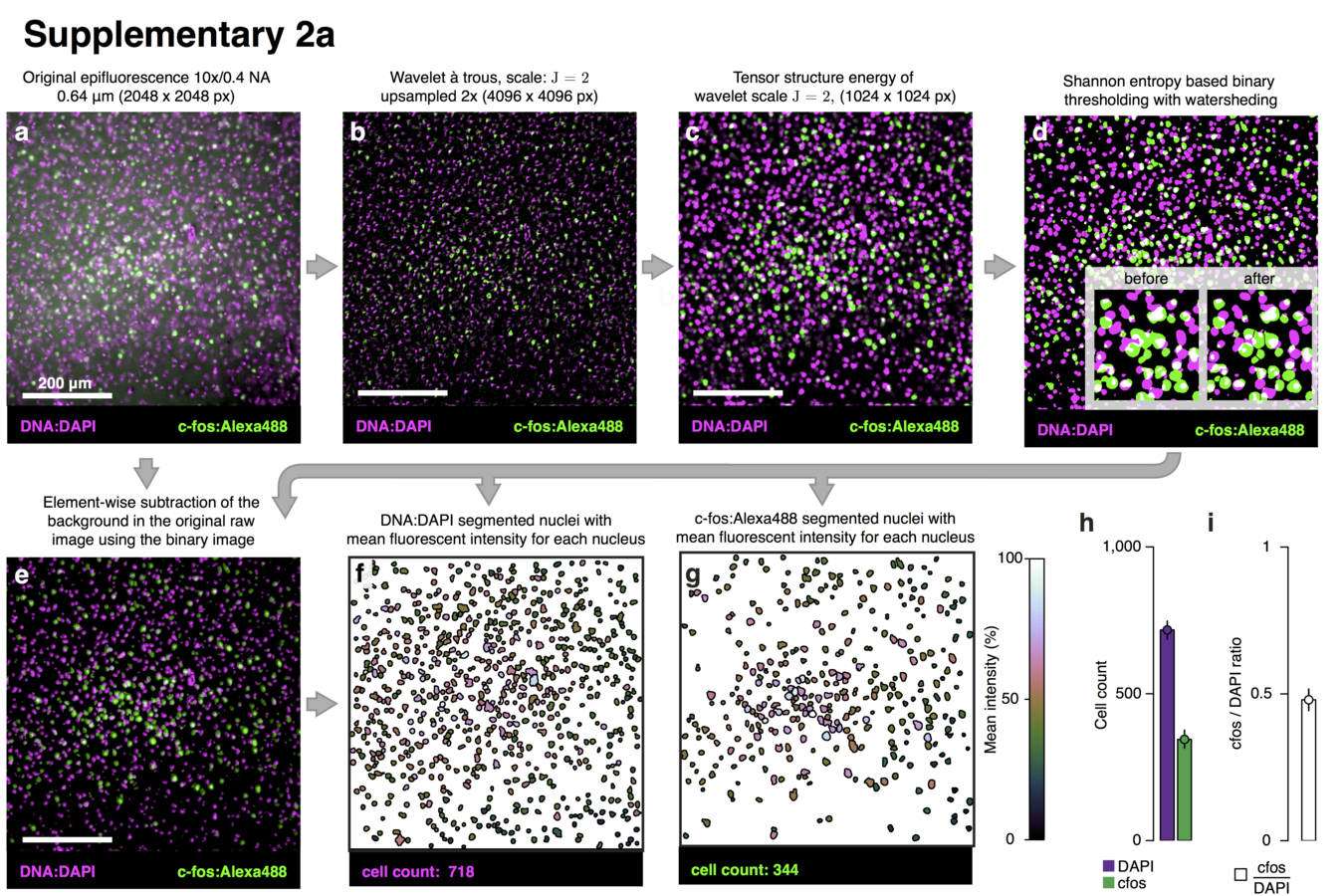
Segmentation of nuclei
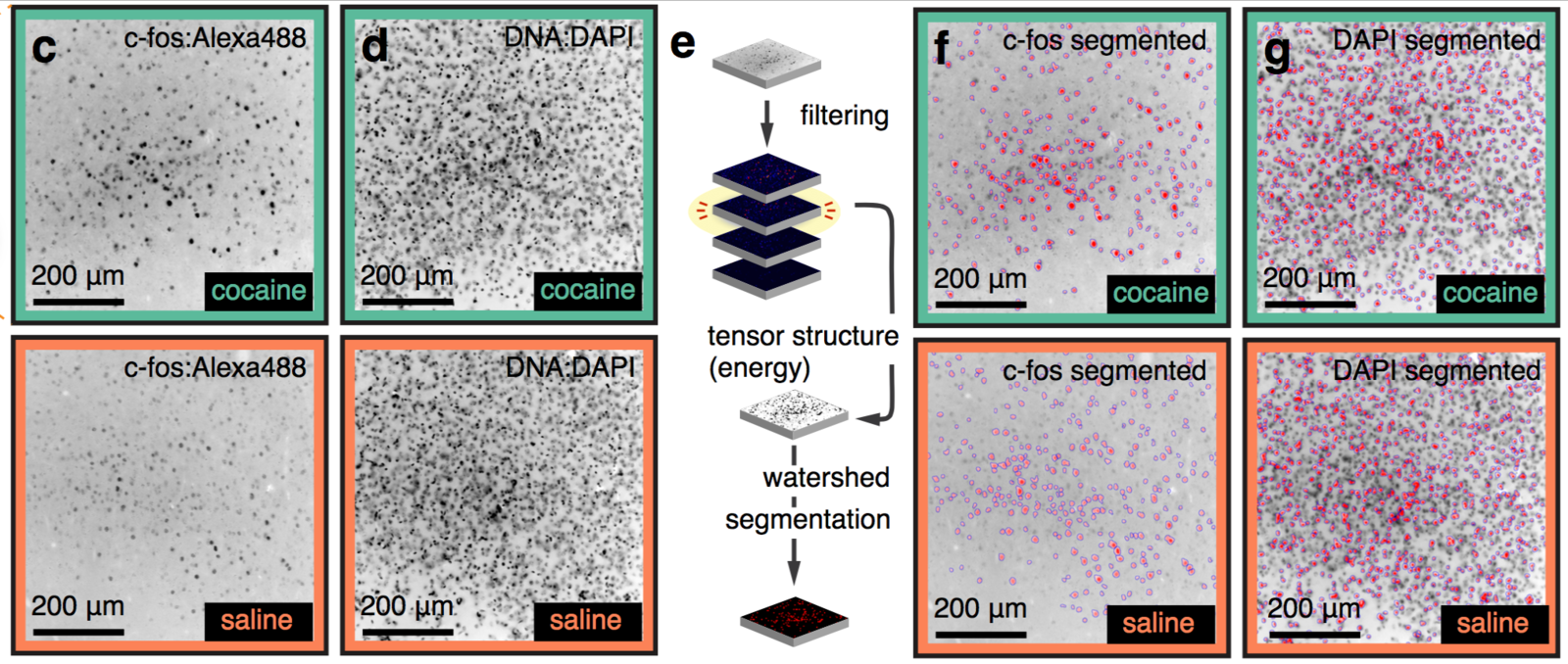
Best measurement of c-fos activity?
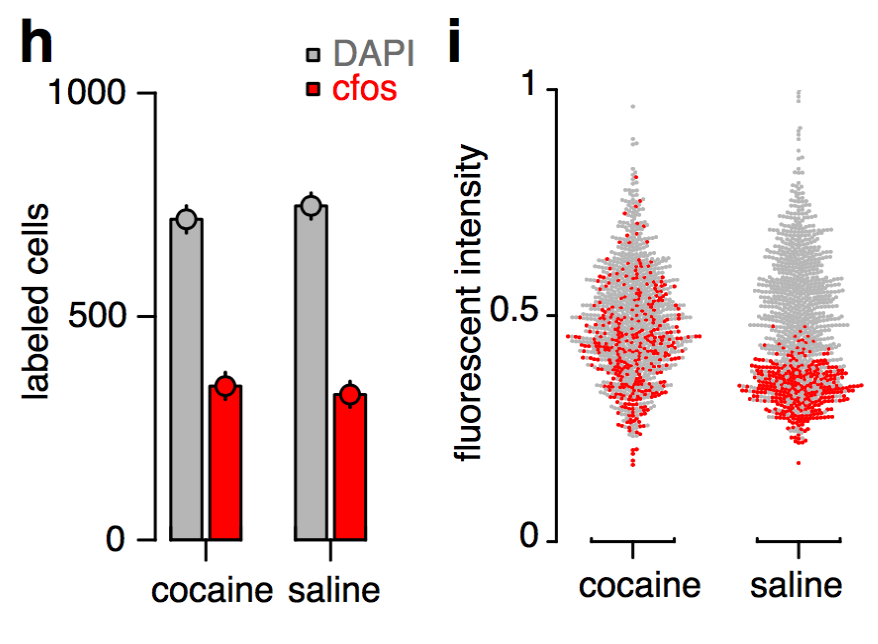
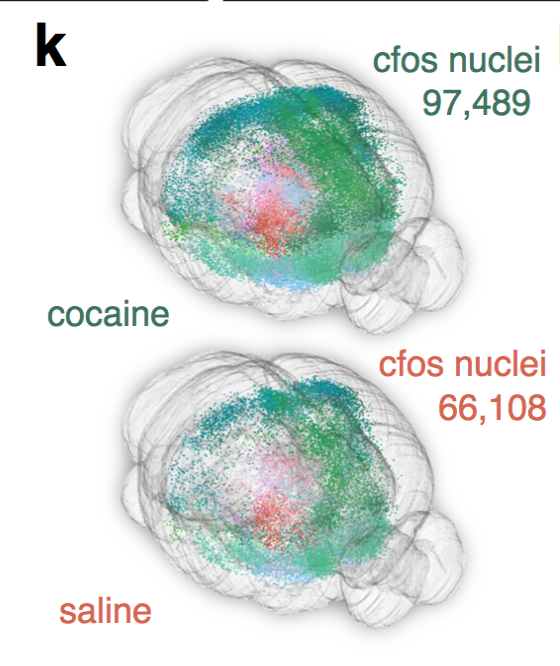
Monotonic function of IEG activity?
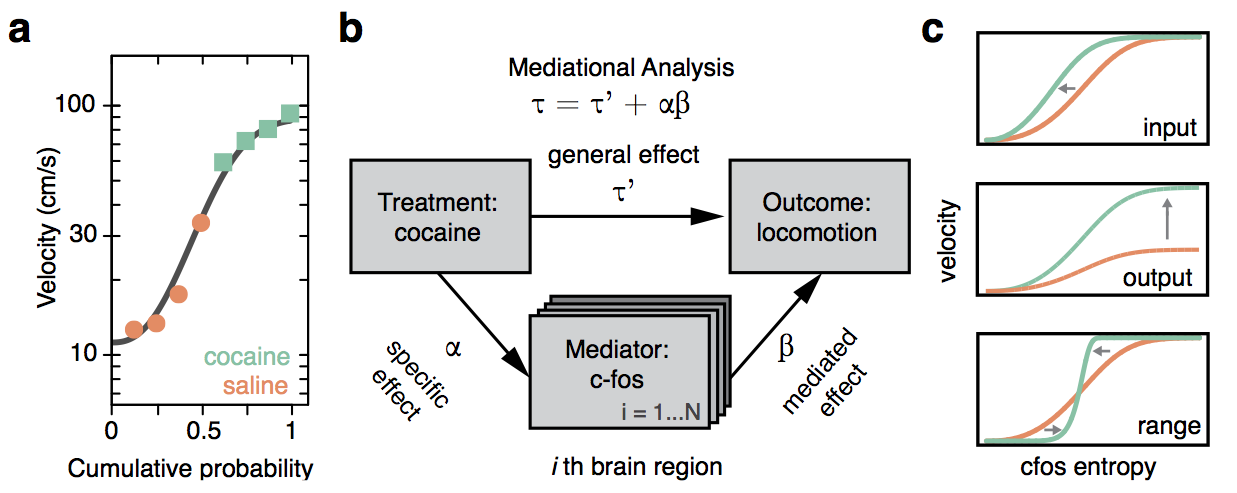
Functional validation
Can be used to segment processes and their direction.
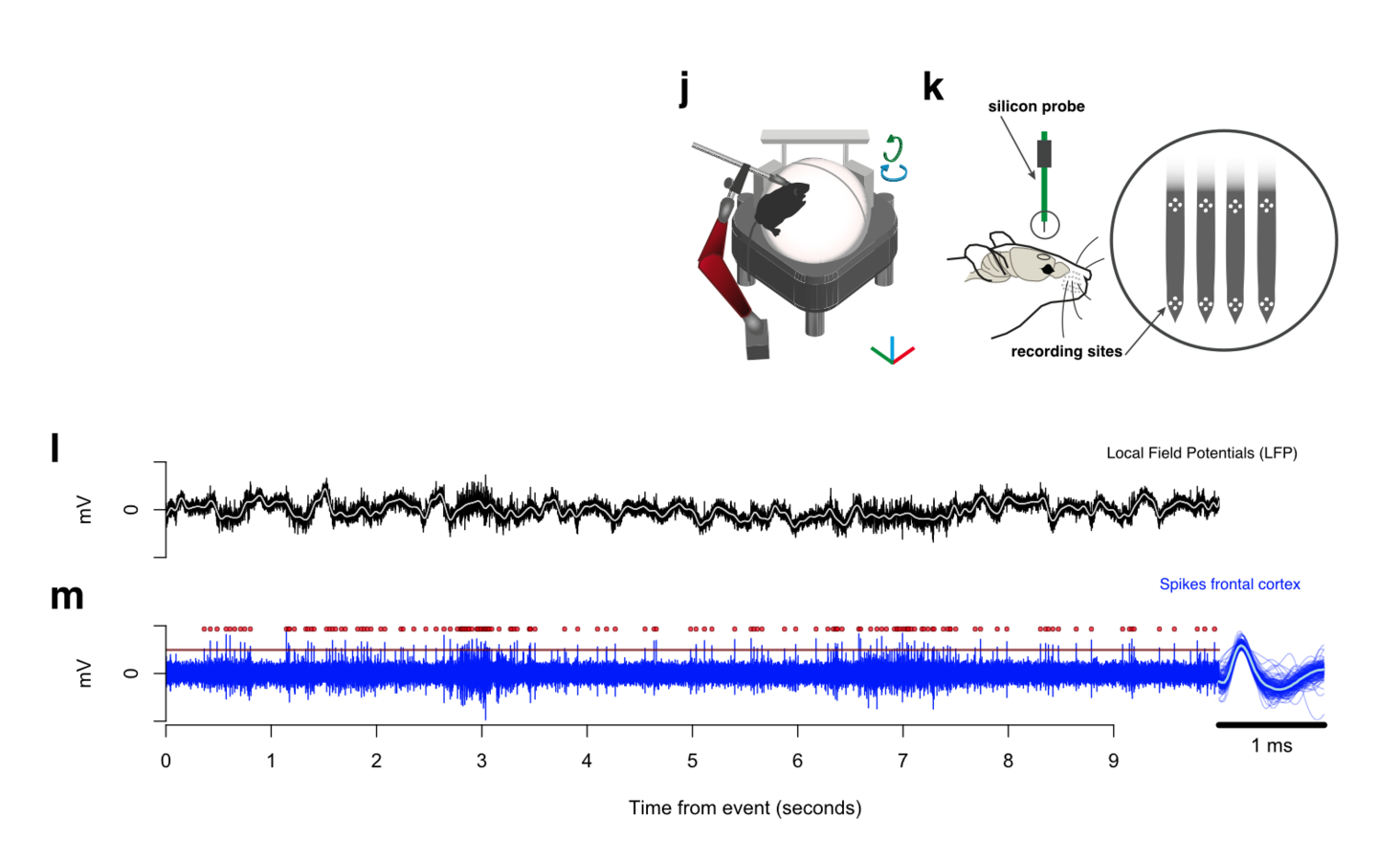
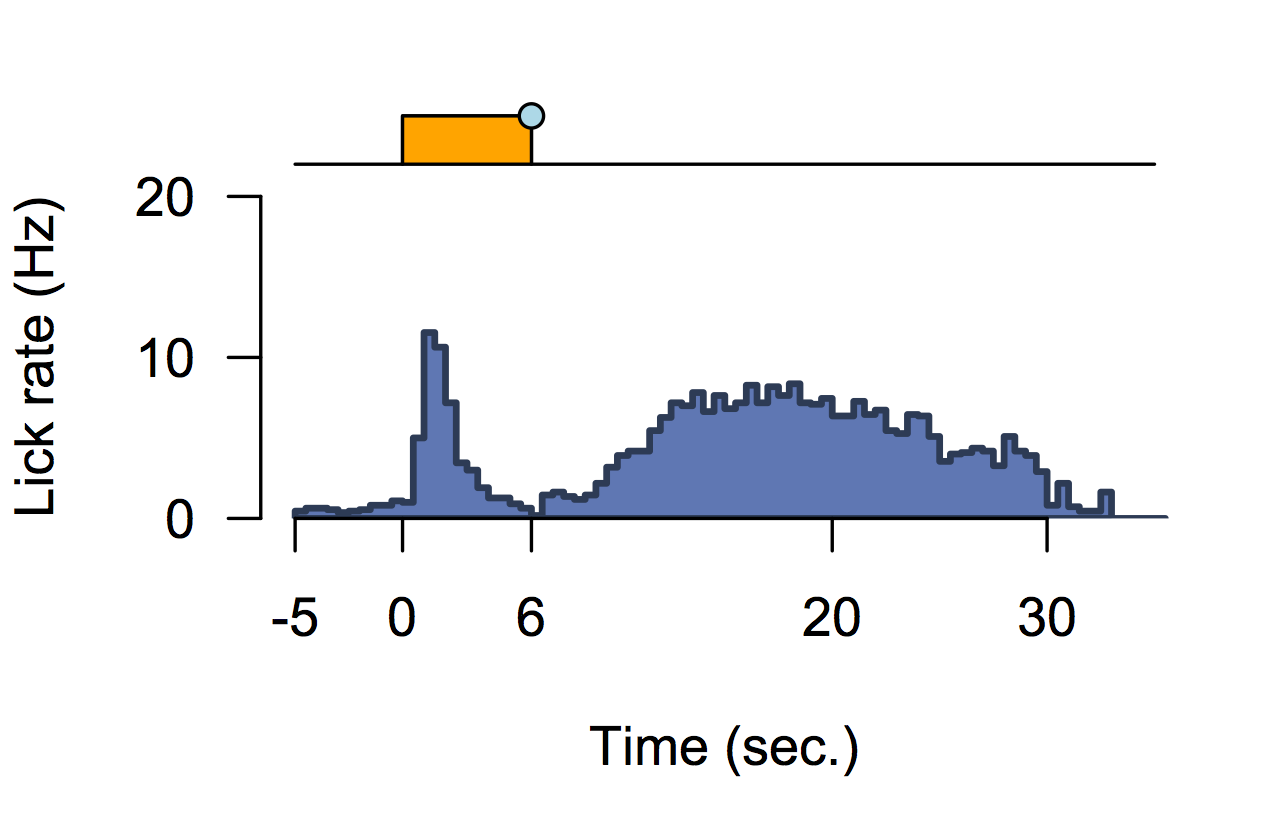
Functional validation
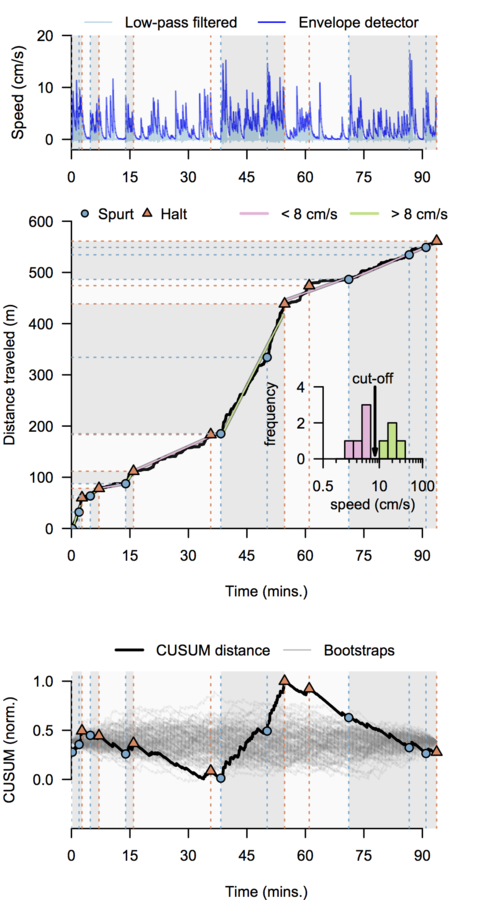
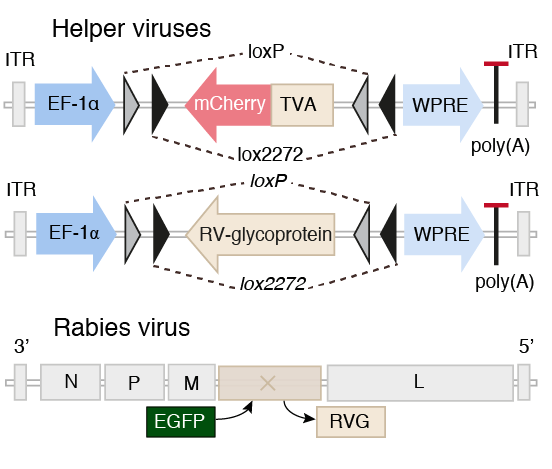
Scalable monosynaptic architecture
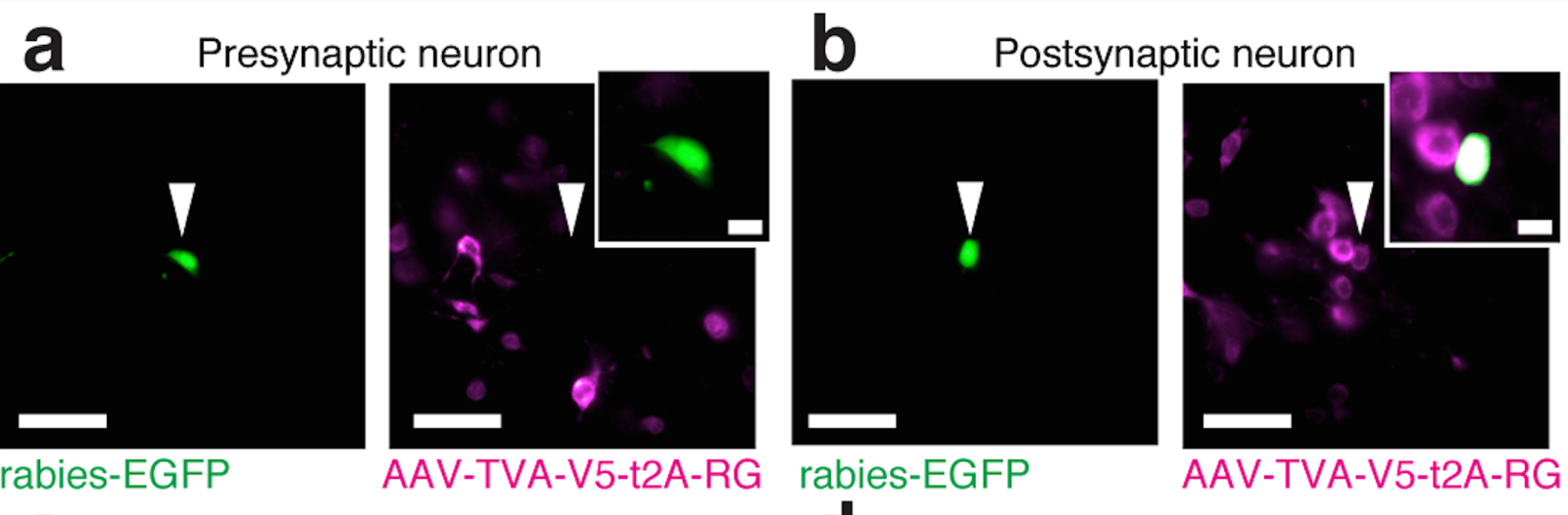
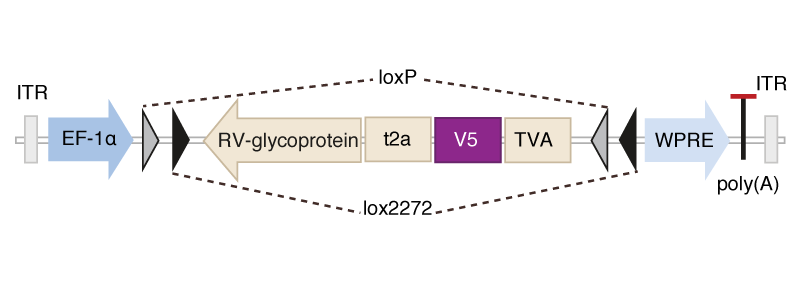
Scalable monosynaptic architecture
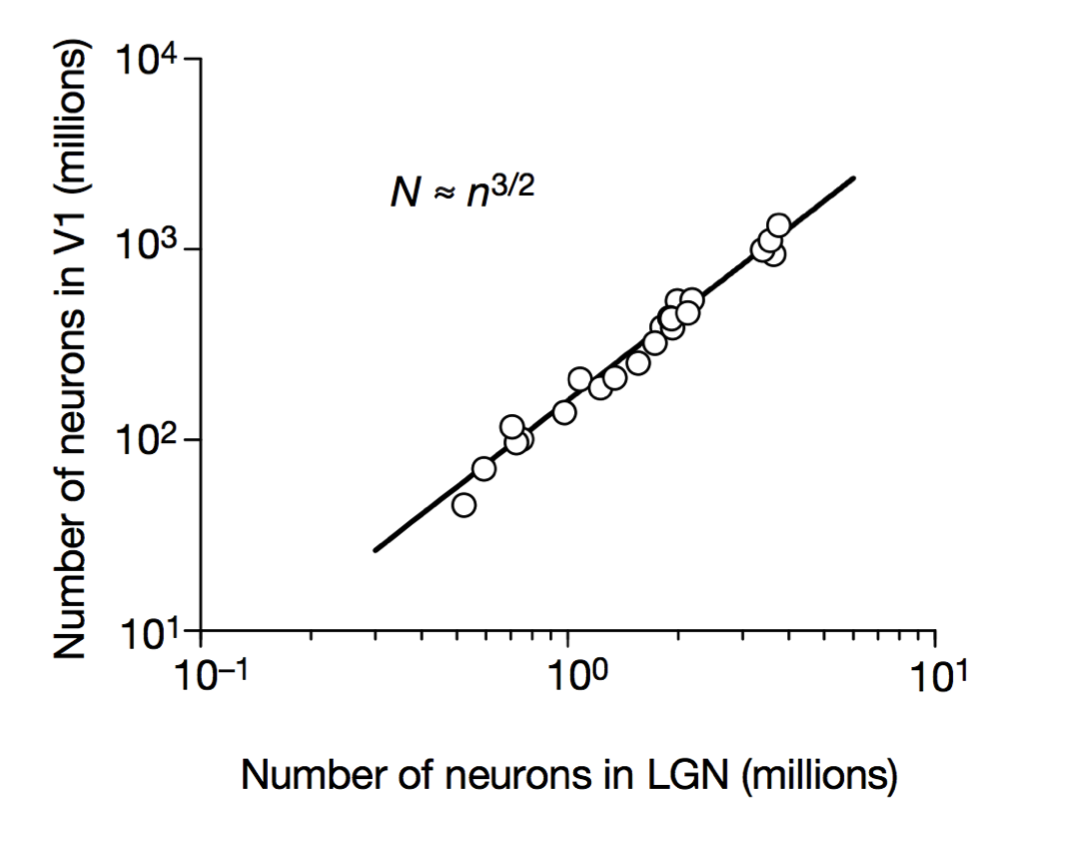
Stevens (1999)
Scalable monosynaptic architecture
Stevens (1999)
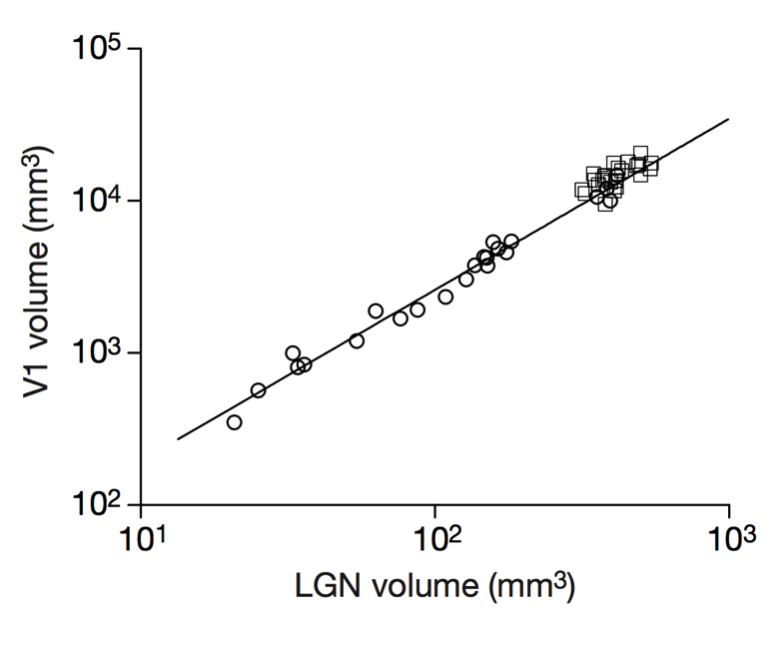
Scalable monosynaptic architecture
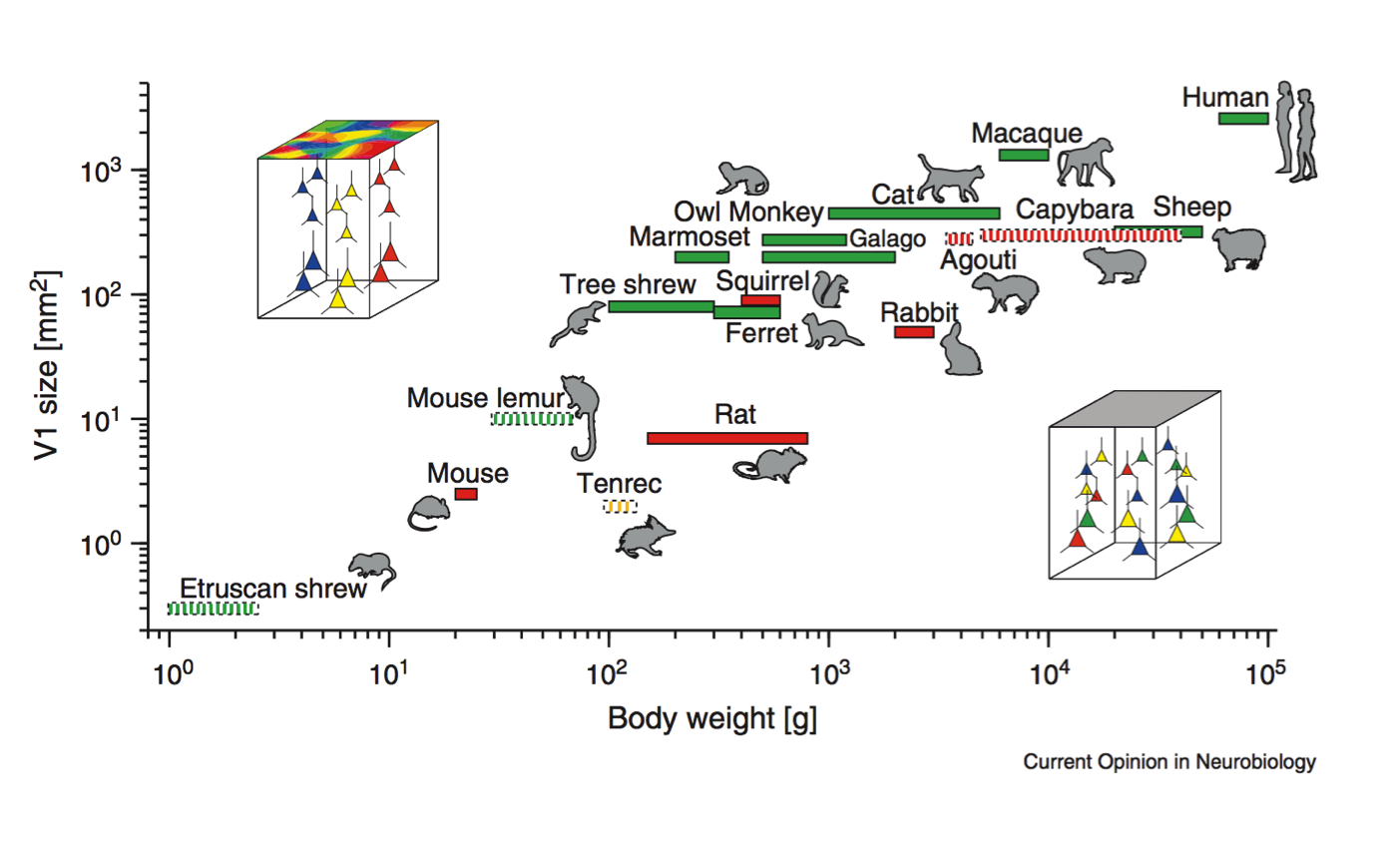
Scalable monosynaptic architecture
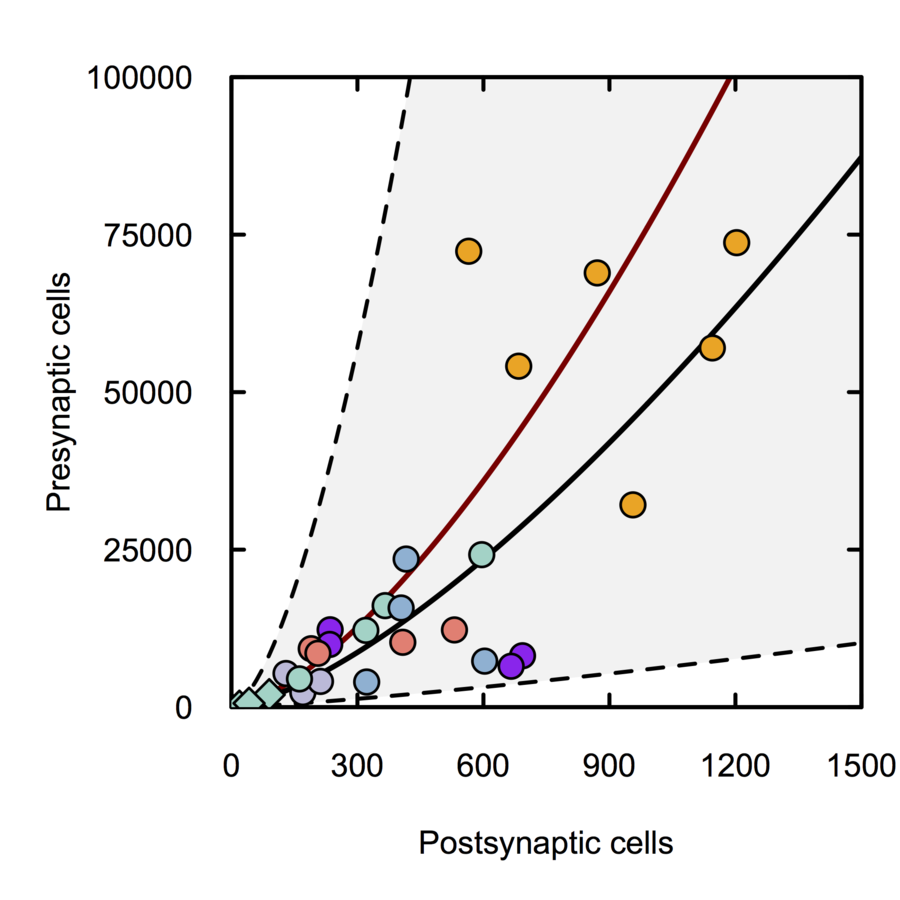
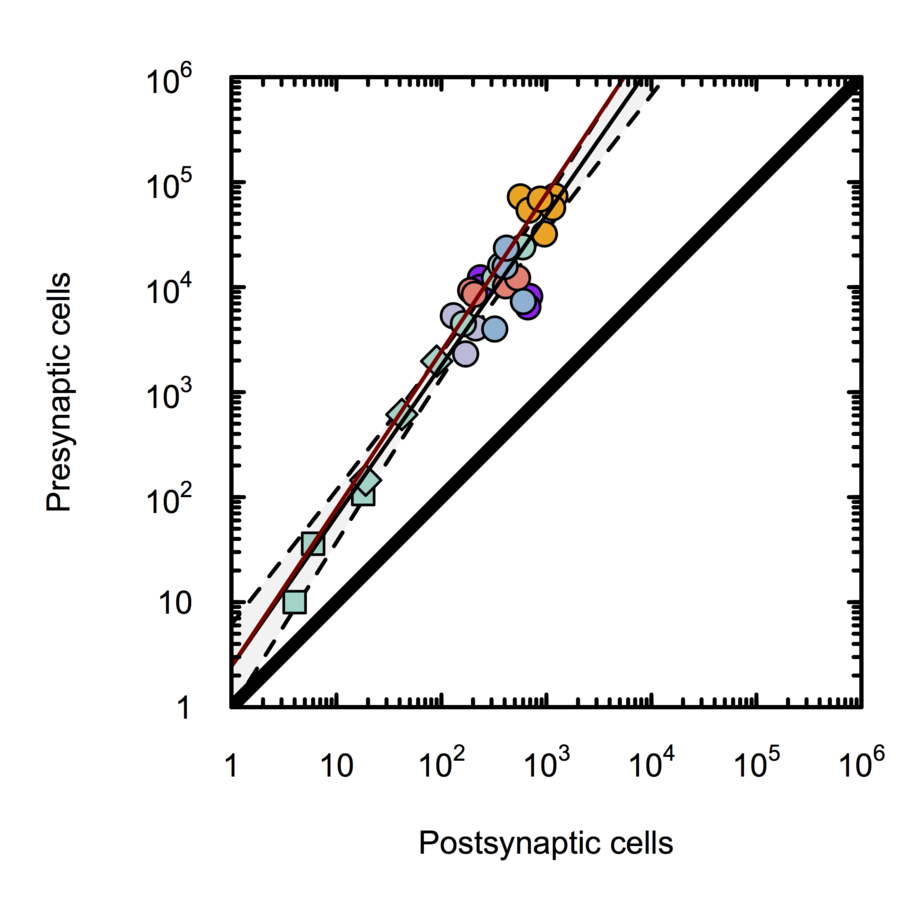
Scalable monosynaptic architecture
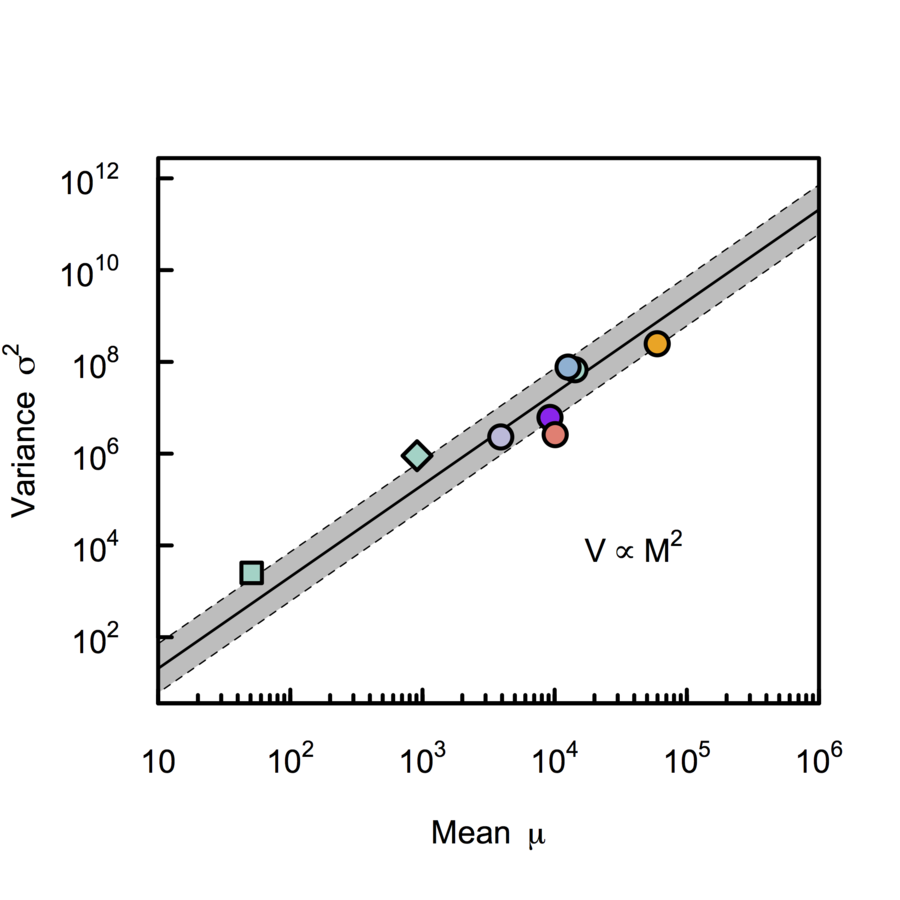

scRNA-seq
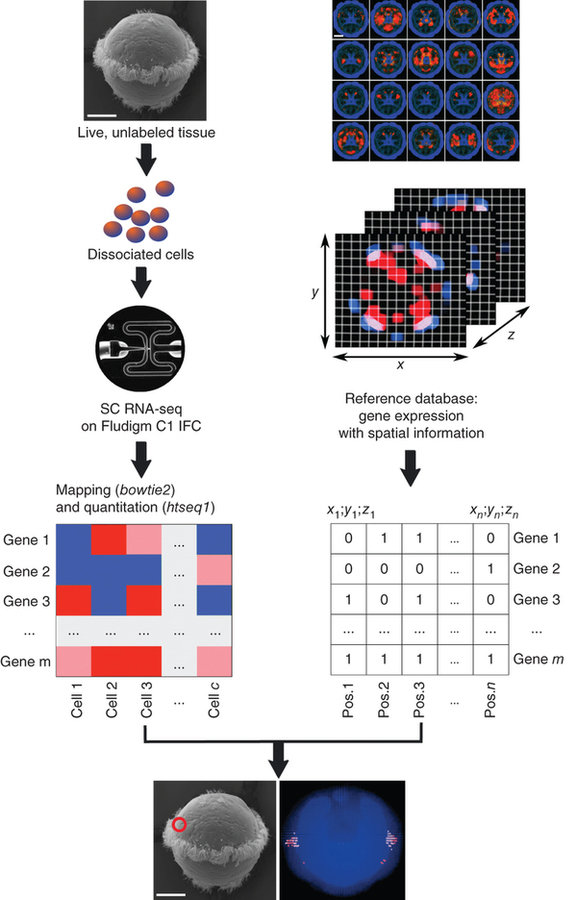
NATURE BIOTECHNOLOGY | COMPUTATIONAL BIOLOGY | ANALYSIS
High-throughput spatial mapping of single-cell RNA-seq data to tissue of origin
Kaia Achim, Jean-Baptiste Pettit, Luis R Saraiva, Daria Gavriouchkina, Tomas Larsson, Detlev Arendt & John C Marioni
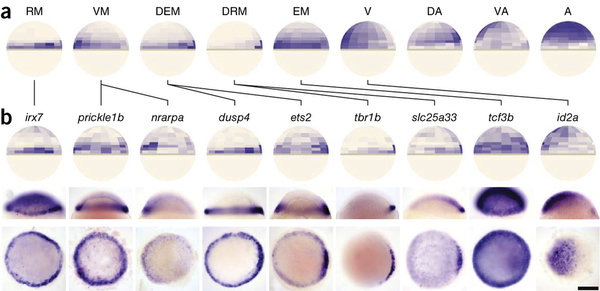
scRNA-seq
Allen Brain Reference Atlas

-
Atlas 2007 (manually drawn Nissl):
- 200 μm thick coronal sections.
-
Atlas 2011:
- 100 μm both coronal and sagital
- Atlas 2014 (connectivity avrg template)
-
Atlas 2015 (beginning of june):
- 10 x 50 μm
-
Registration atlas:
- 25 x 25 μm
-
Grid expression ISH:
- 200 x 200 μm MetaIOimage (.raw, .mhd)
scRNA-seq
scRNA-seq
Allen Brain Reference Atlas
scRNA-seq
Anatomic Gene Expression Atlas
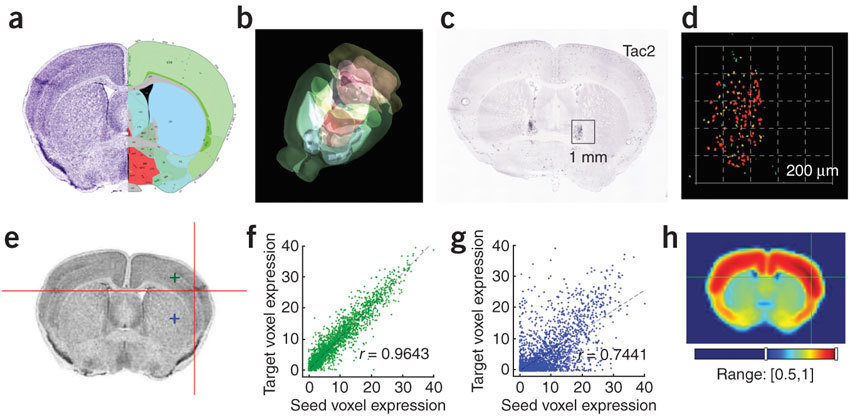
Lydia Ng, et al. (2009) Nat. Neuro.
http://mouse.brain-map.org/agea
scRNA-seq
324 cells from cortico-striatal section
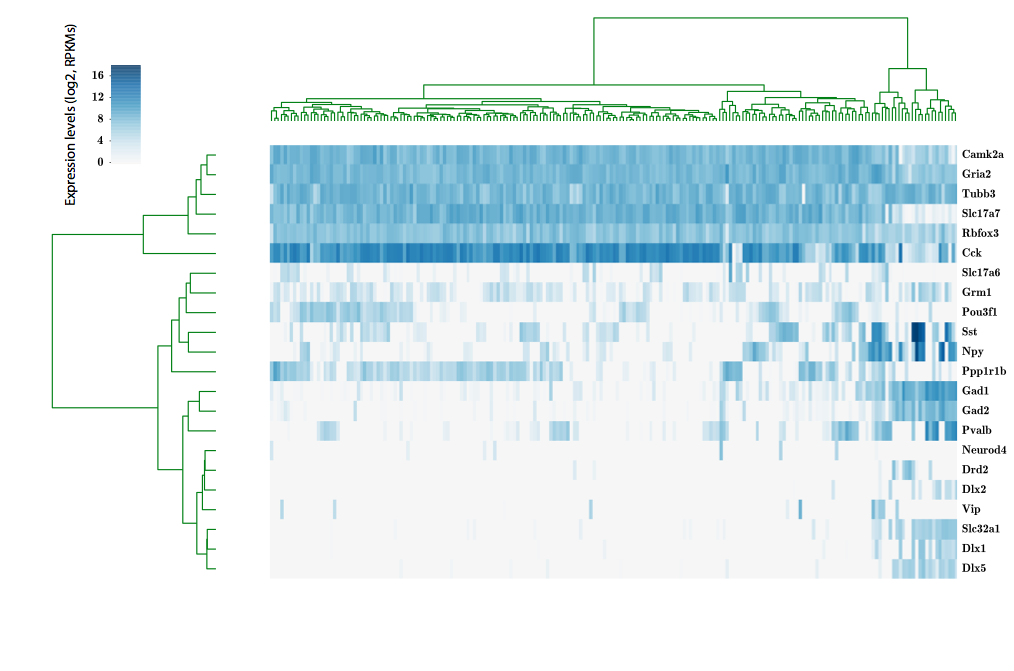
scRNA-seq
Our approach
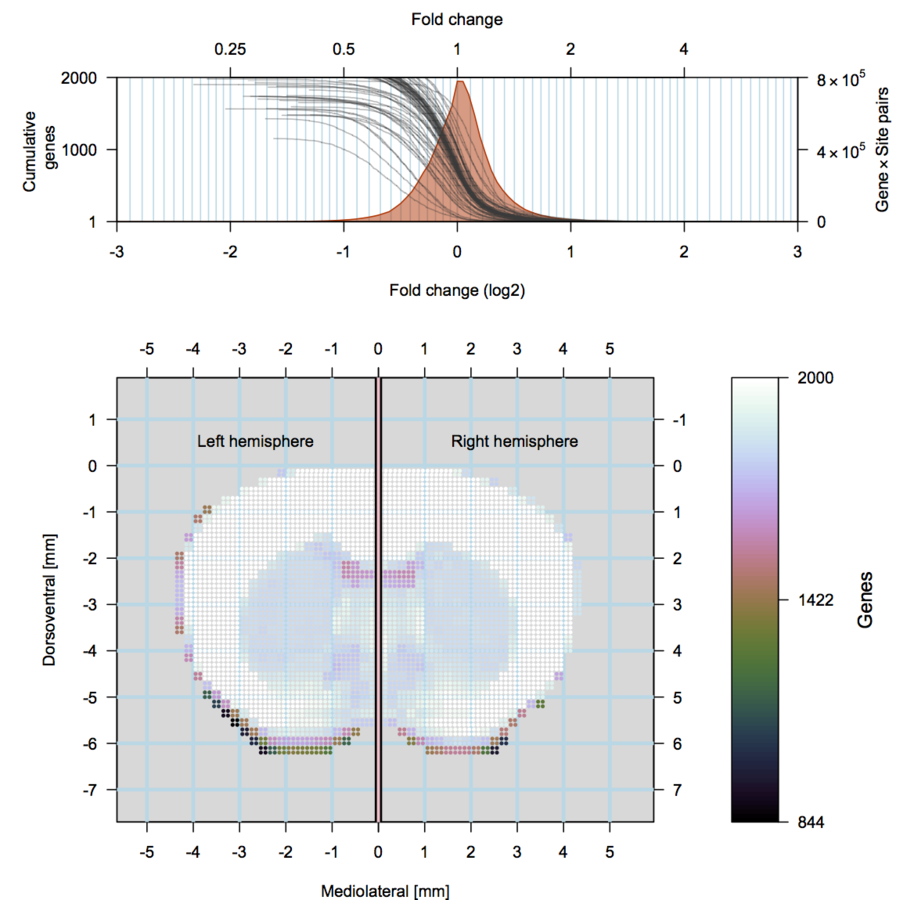
scRNA-seq
Our approach

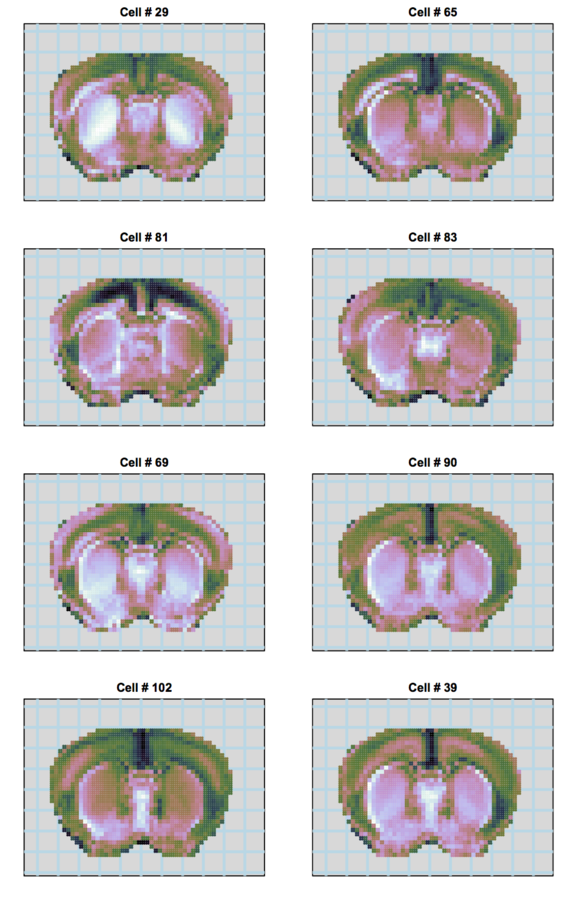
Multiplexed padlock probes
Mats Nilsson
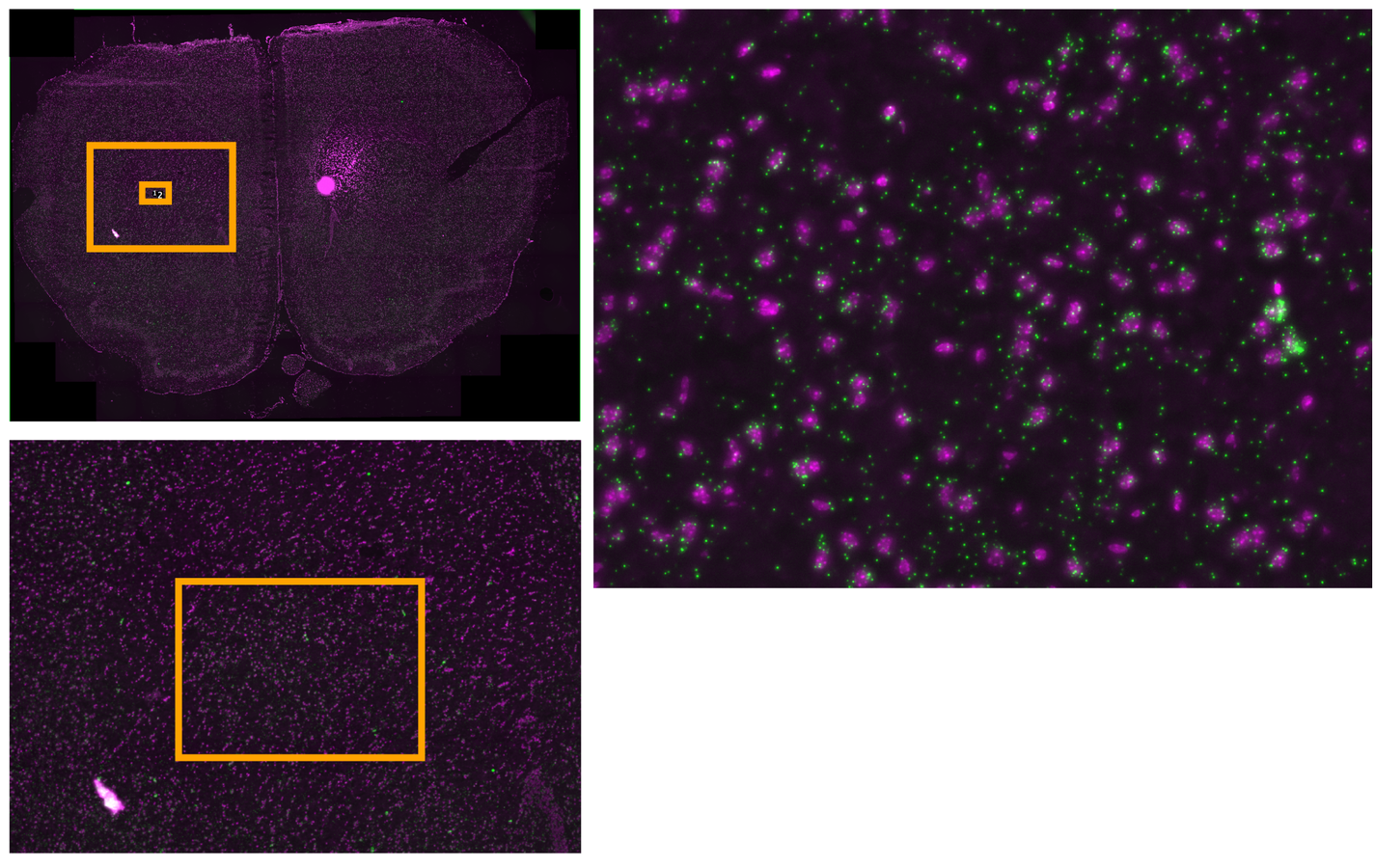
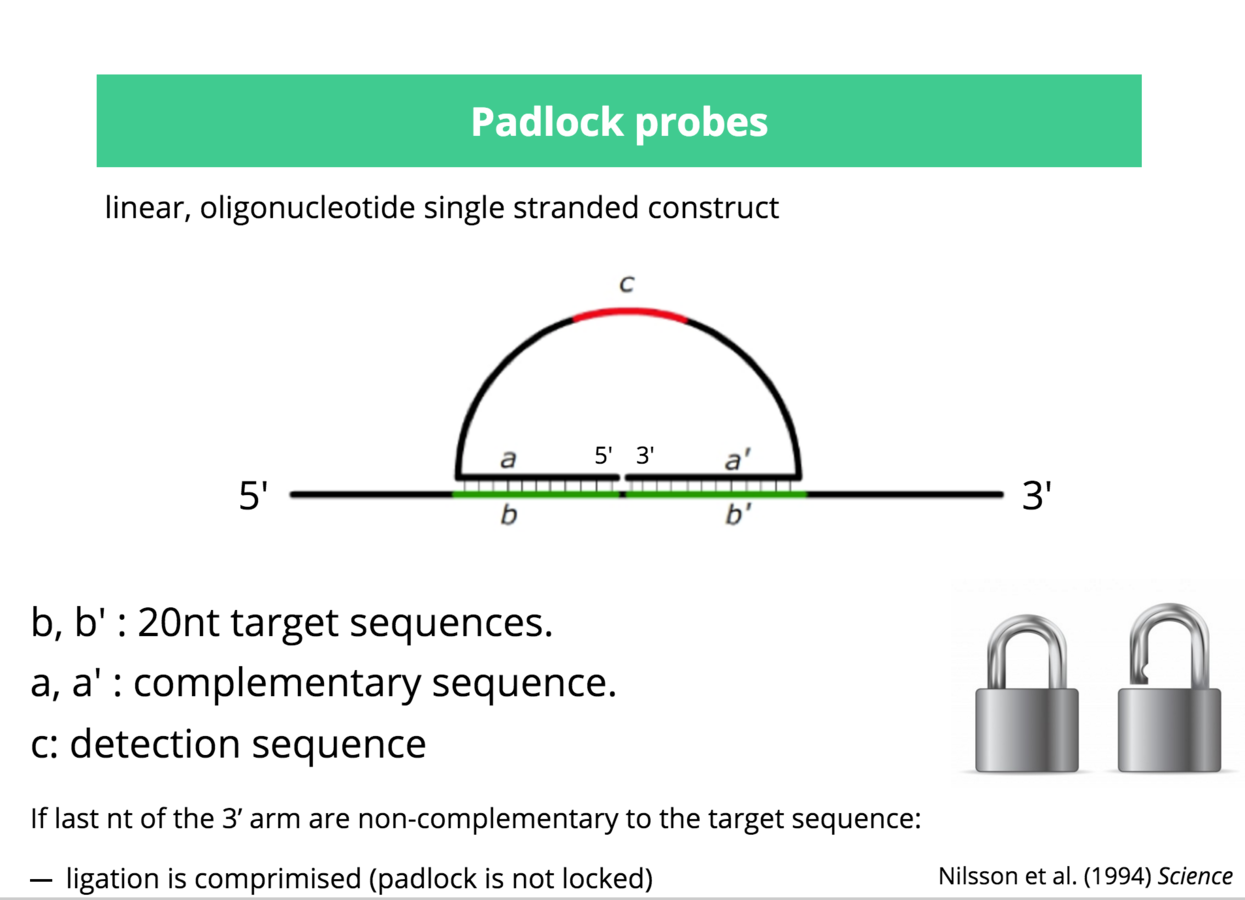
Multiplexed padlock probes
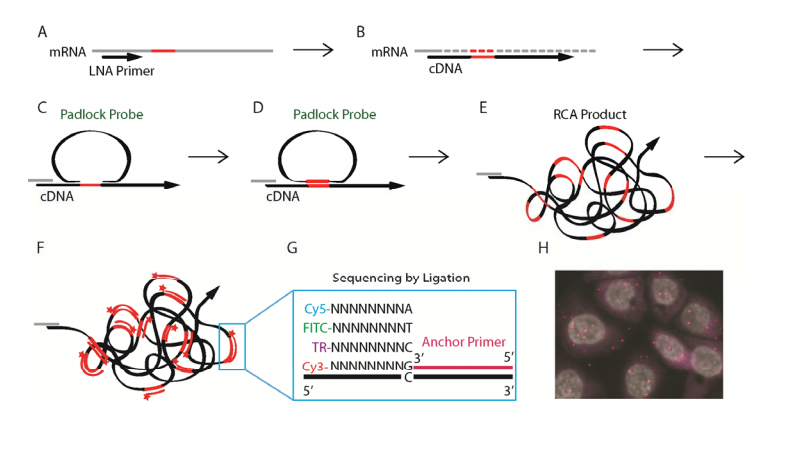
Reverse transcription
RNase H digestion
Hybridization
Ligation
Rolling Circle Amplification
mFISH
Base calling
Multiplexed padlock probes
Mats Nilsson
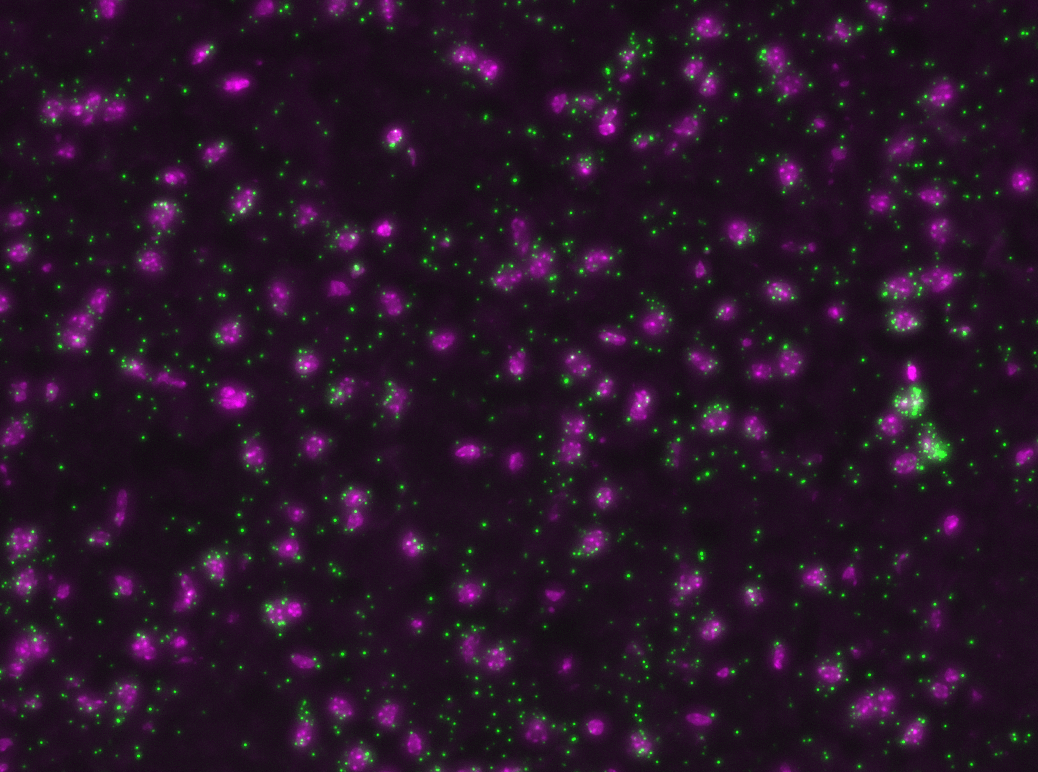
Multiplexed padlock probes
Mats Nilsson

Multiresolution decomposition

Thank you!
Concurrency and parallel programming
- Multi-threaded applications through .

#include <string>
#include <iostream>
#include <thread>
using namespace std;
//The functions we want to make the thread run.
void task1(string msg)
{
cout << "task1 says: " << msg;
}
void task2(string msg)
{
cout << "task1 says: " << msg;
}
//Main loop.
int main()
{
thread t1(task1, "Task 1 executed");
thread t2(task2, "Task 1 executed");
//let main wait for t1 and t2 to finish.
t1.join();
t2.join();
}Rcpp

Dual core
Concurrency and parallel programming
- Multi threaded applications through .

#include <string>
#include <iostream>
#include <thread>
using namespace std;
//The functions we want to make the thread run.
void task1(string msg)
{
cout << "task1 says: " << msg;
}
void task2(string msg)
{
cout << "task1 says: " << msg;
}
//Main loop.
int main()
{
thread t1(task1, "Task 1 executed");
thread t2(task2, "Task 1 executed");
t1.join();
t2.join();
}Rcpp
Parallel computing
- Parallel computing is extremely simple to implement from R.
# install.packages('foreach'); install.packages('doSNOW')
library(foreach)
library(doSNOW)
cl <- makeCluster(2, type = "SOCK")
registerDoSNOW(cl)
getDoParName()
#matrix operators
x <- foreach(i=1:8, .combine='rbind', .packages='wholebrain' ) %:%
foreach(j=1:2, .combine='c', .packages='wholebrain' ) %dopar% {
l <- runif(1, i, 100)
i + j + l
}R package
- Why R?
- Standard data analysis:
- load some data
- estimate the density distribution.
- plot it
- Standard data analysis:
xx <- faithful$eruptions
fit <- density(xx)
plot(fit)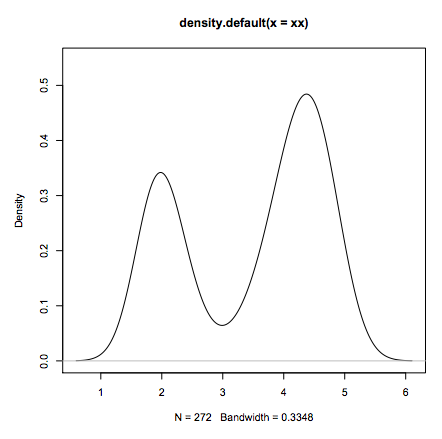
R package
- Why R?
#Line 1: loading
xx <- faithful$eruptions
#Line 2: estimate density
fit1 <- density(xx)
#Line 2: draw 10'000 bootstraps
fit2 <- replicate(10000, {
x <- sample(xx,replace=TRUE);
density(x, from=min(fit1$x), to=max(fit1$x))$y
})
#Line 3: compute 95% error "bars"
fit3 <- apply(fit2, 1, quantile,c(0.025,0.975))
#Line 4: plot the estimate
plot(fit1, ylim=range(fit3))
#Line 5: add estimation error as shaded region
polygon(c(fit1$x,rev(fit1$x)), c(fit3[1,], rev(fit3[2,])), col=’grey’, border=F)
#Line 6: add the line again since the polygon overshadows it.
lines(fit1)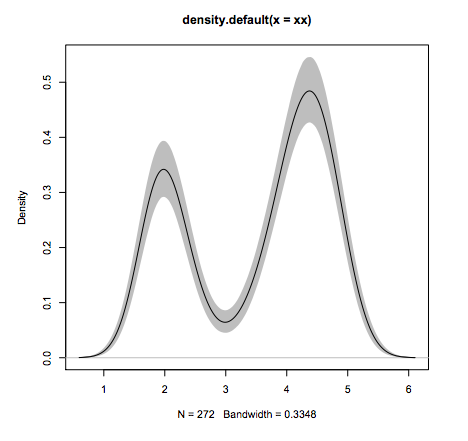
What other language can do this in 6 lines of code?
scRNA-seq
Gene specificity
about ~24'000 genes expressed in the brain.
