Butte Lab Meeting
July 8, 2015
Daniel Himmelstein
Sergio Baranzini Lab

Founding Insight
-
context
bioinformatics → data explosion
-
goal
mine the data to advance human health
-
problem
high-throughput data tends to predict weakly
-
remedy
combine diverse datasets into a strong predictor
-
method
heterogeneous network edge prediction
Predicting Disease-Associated Genes with Heterogeneous Network Edge Prediciton
In publishing delay at PLOS Computational Biology
wealth of GWAS associations
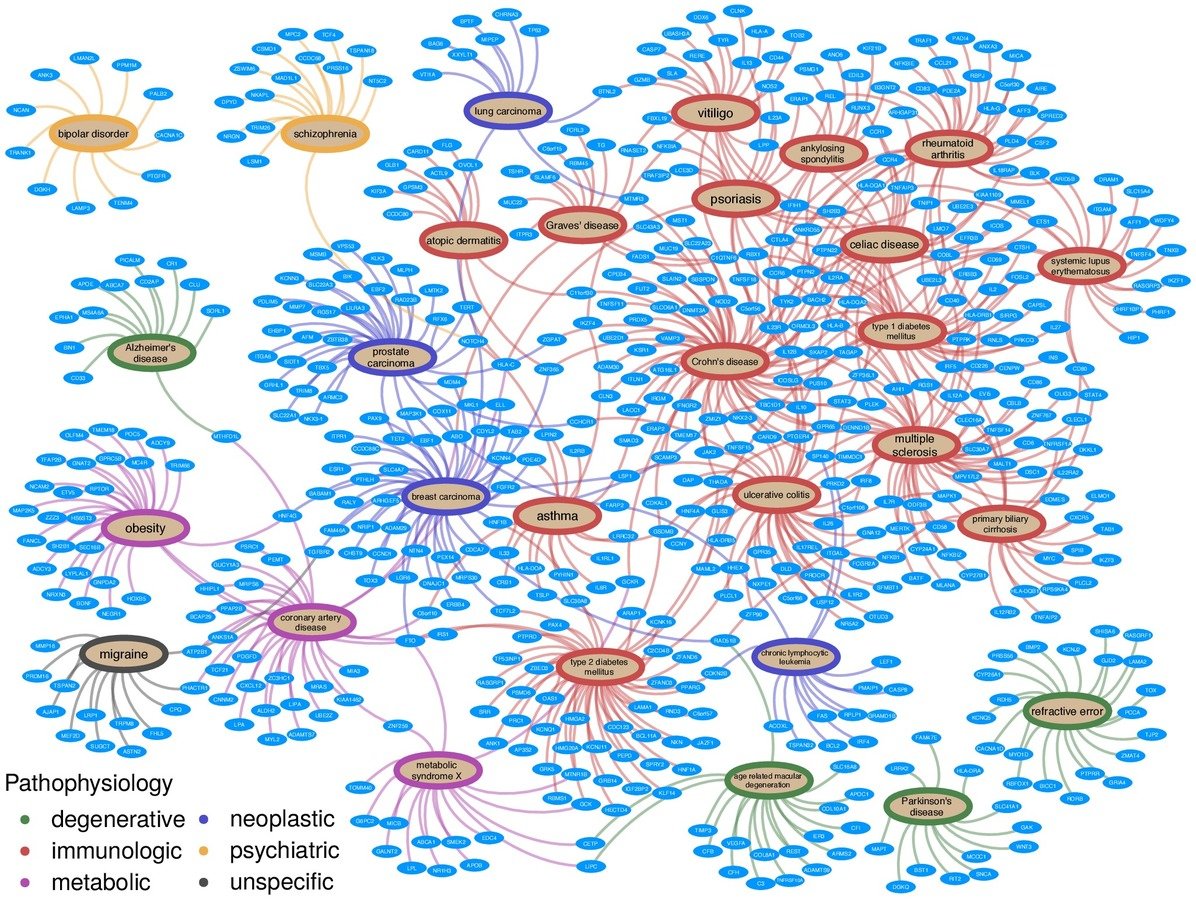
integrate diverse data to provide context
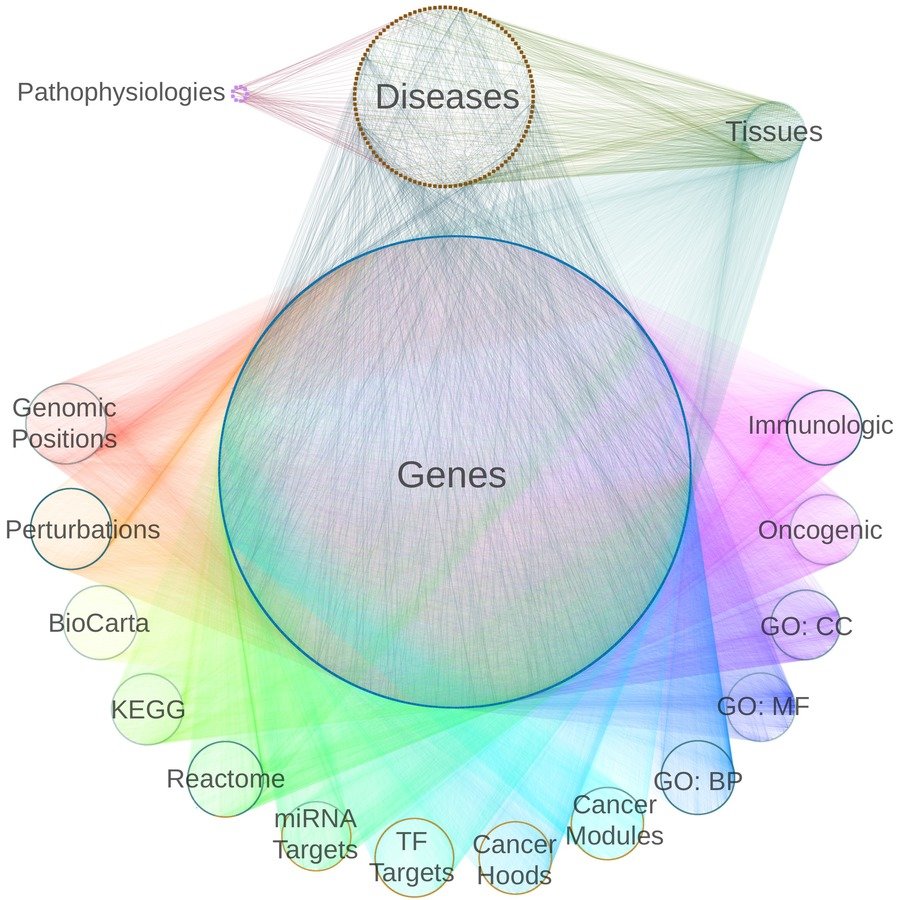
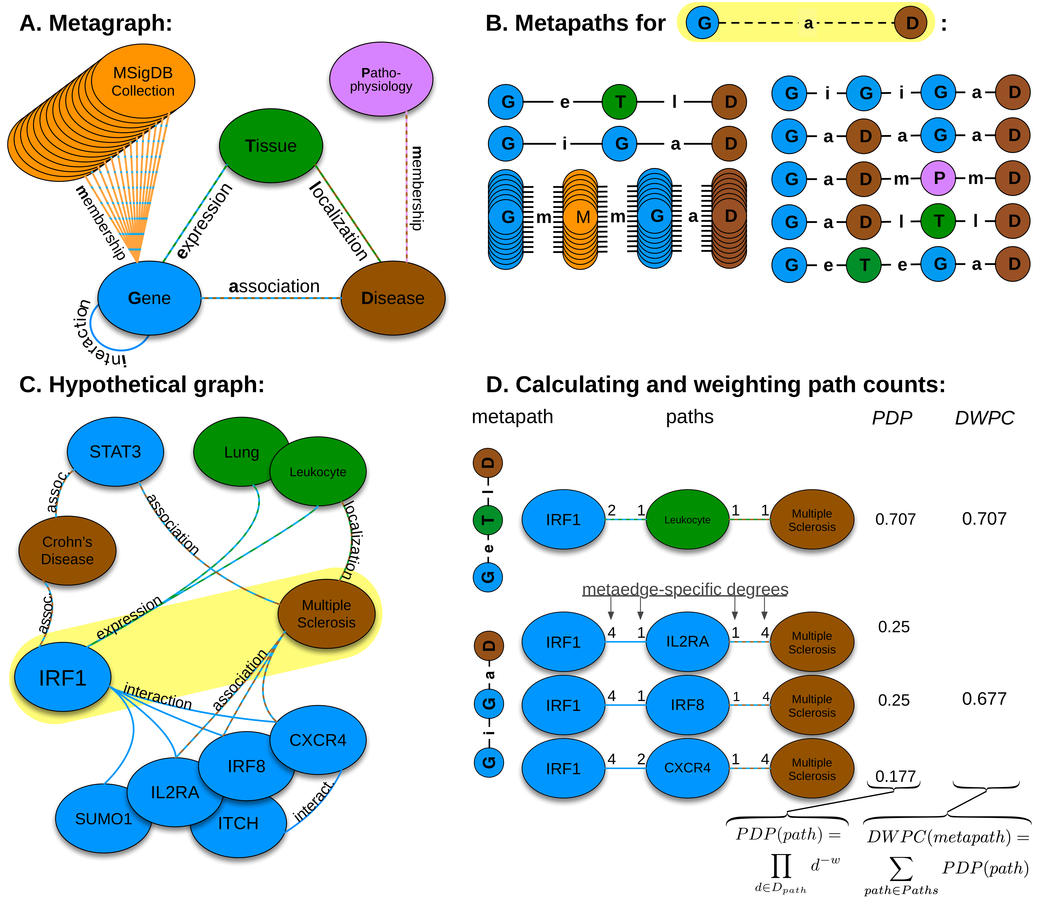
network of pathogenesis:
- 18 metanodes
- 40,343 nodes
- 19 metaedges
- 1,608,168 edges
HNEP
method
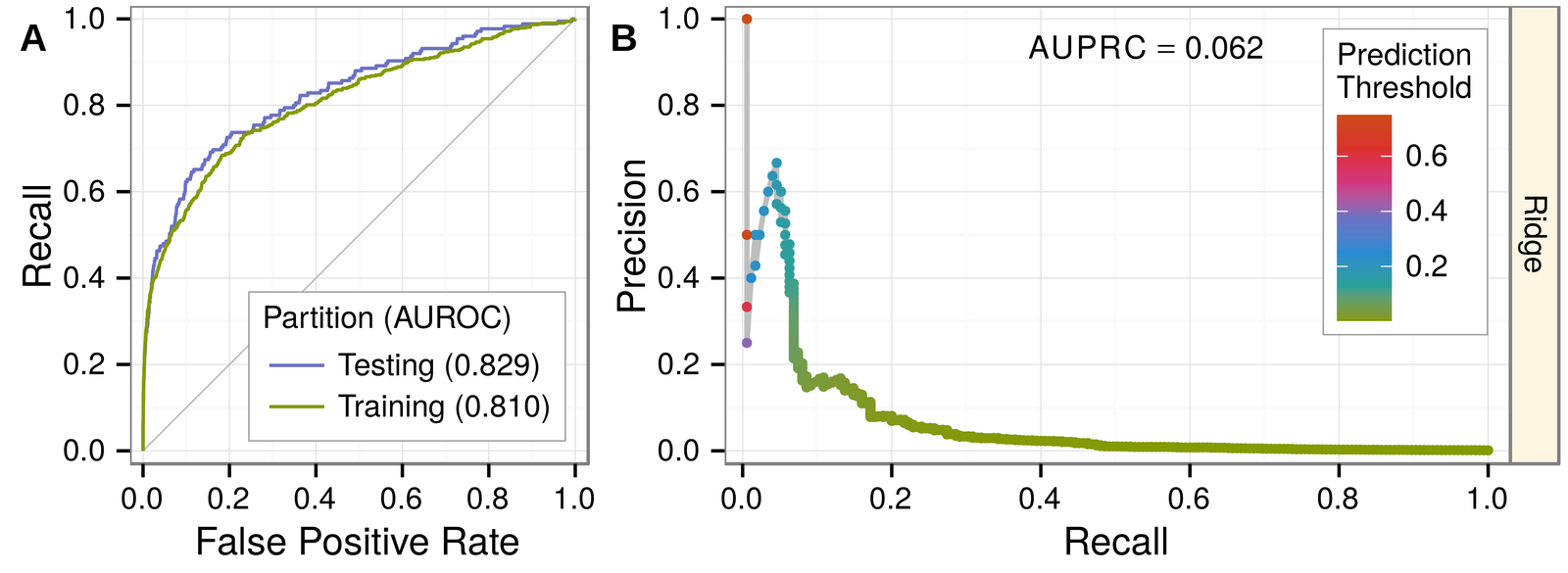
classifier:
- regularized logistic regression
- glmnet in R
- mixing parameter
- top predictions have high lift
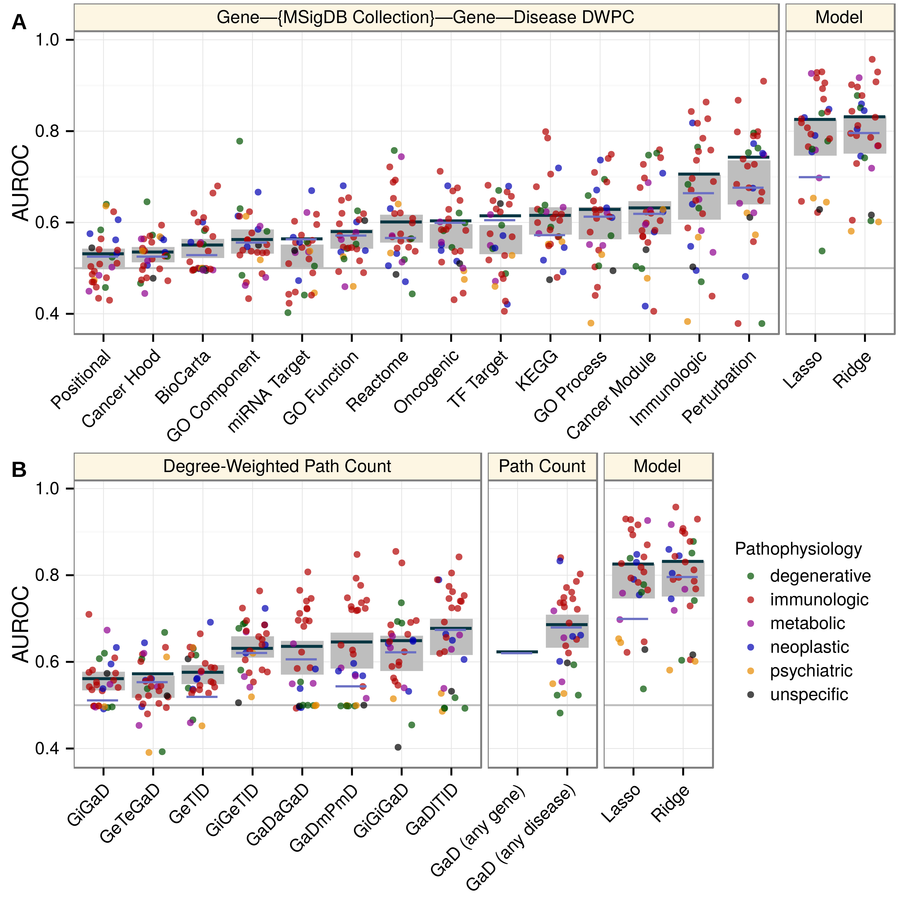
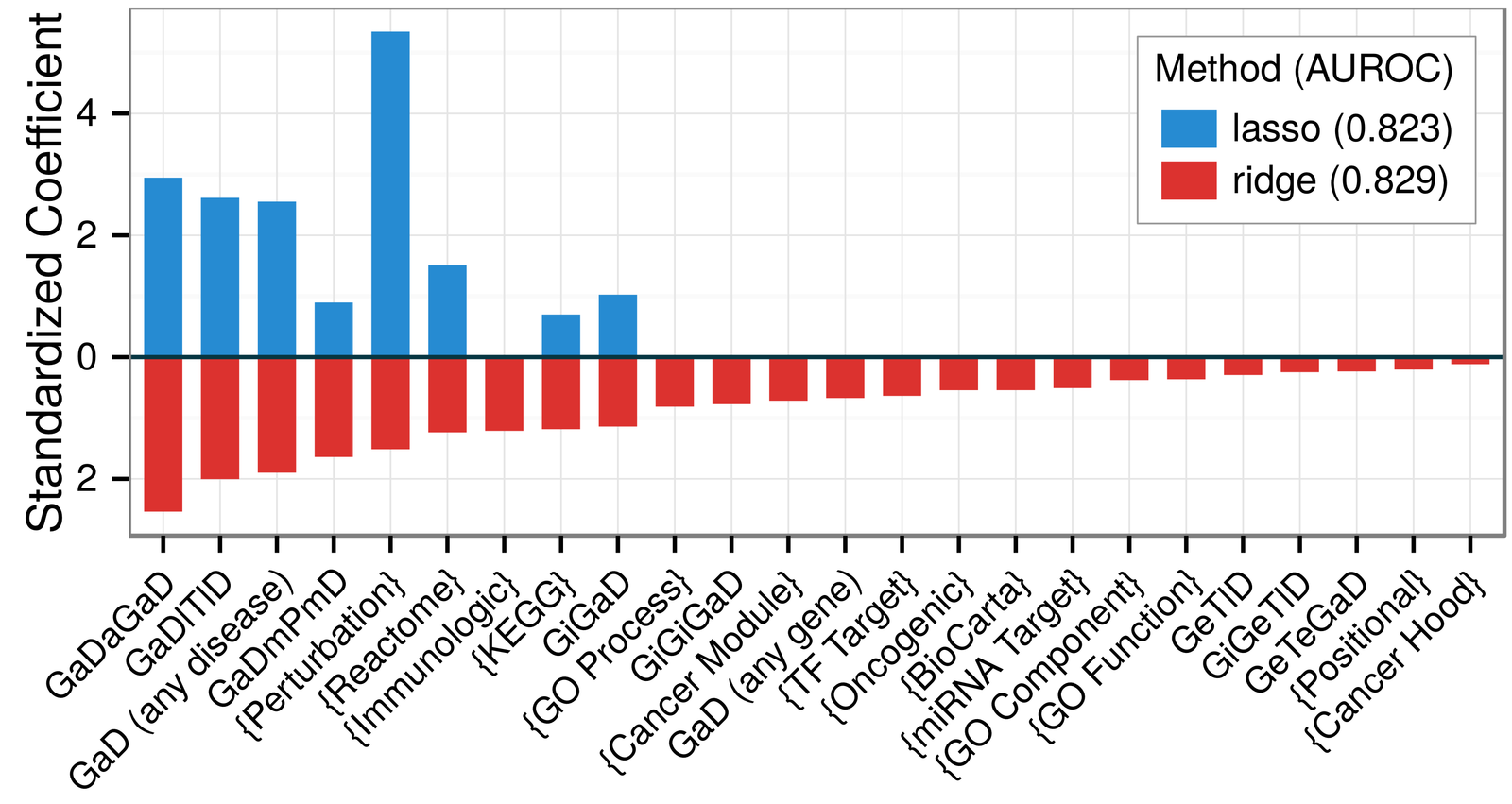
mechanisms of pathogenesis:
- integrative approach is best
- pleiotropy
- transcriptional signatures of perturbations
- pathways
- protein interactions
supplements
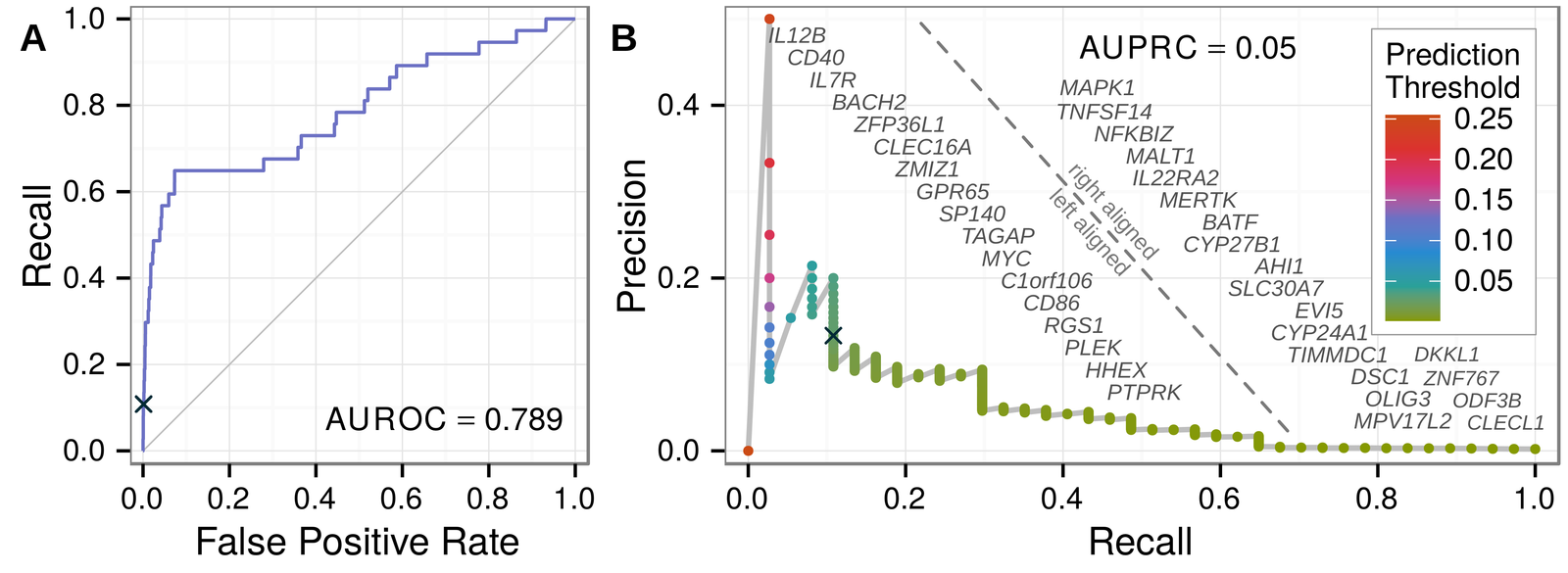
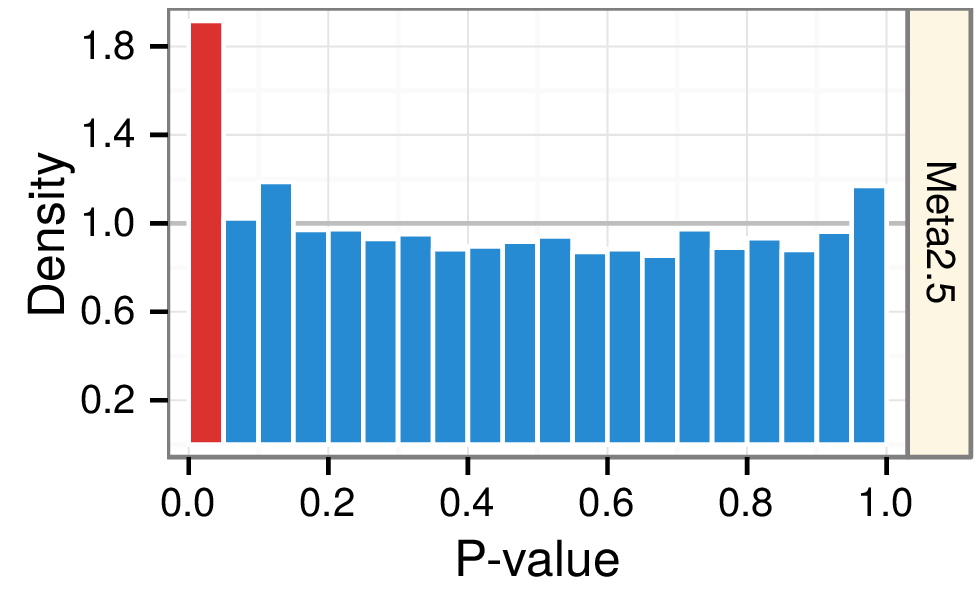

Predicting withheld MS associations
Novel MS associations
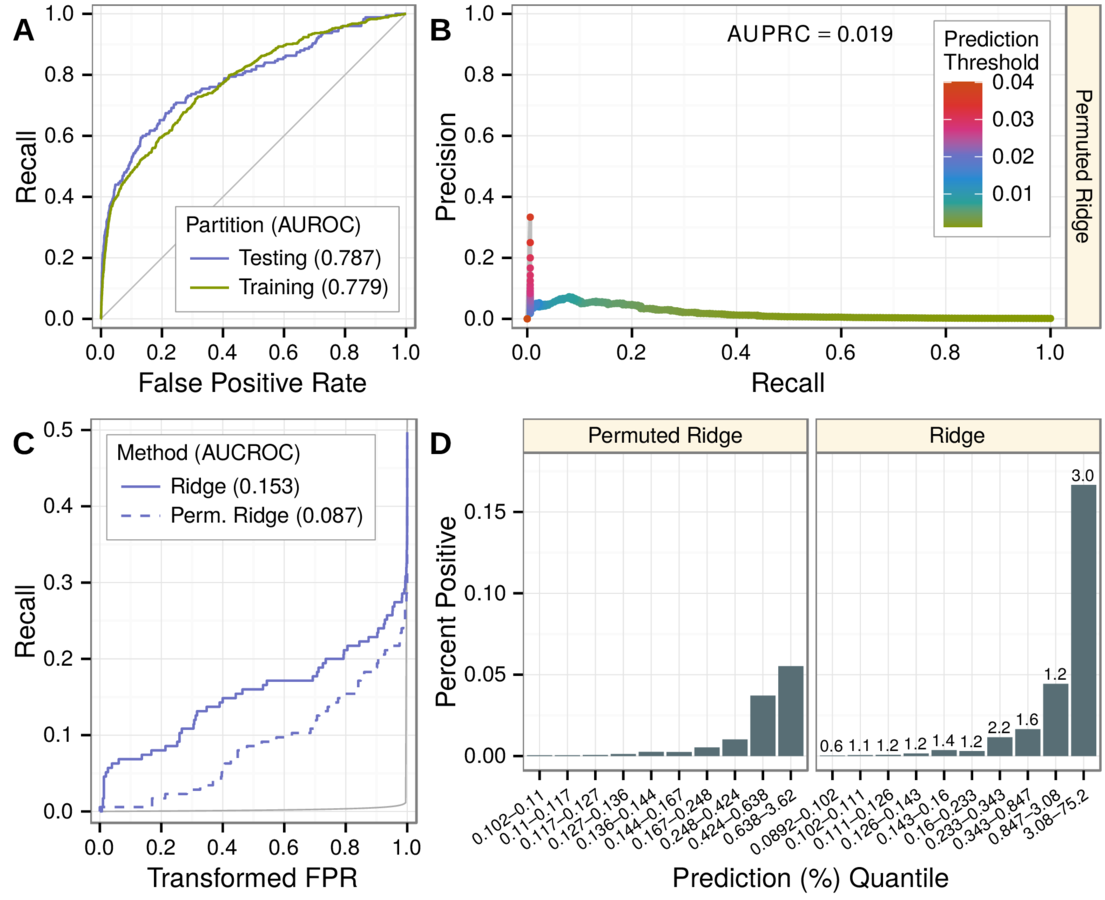
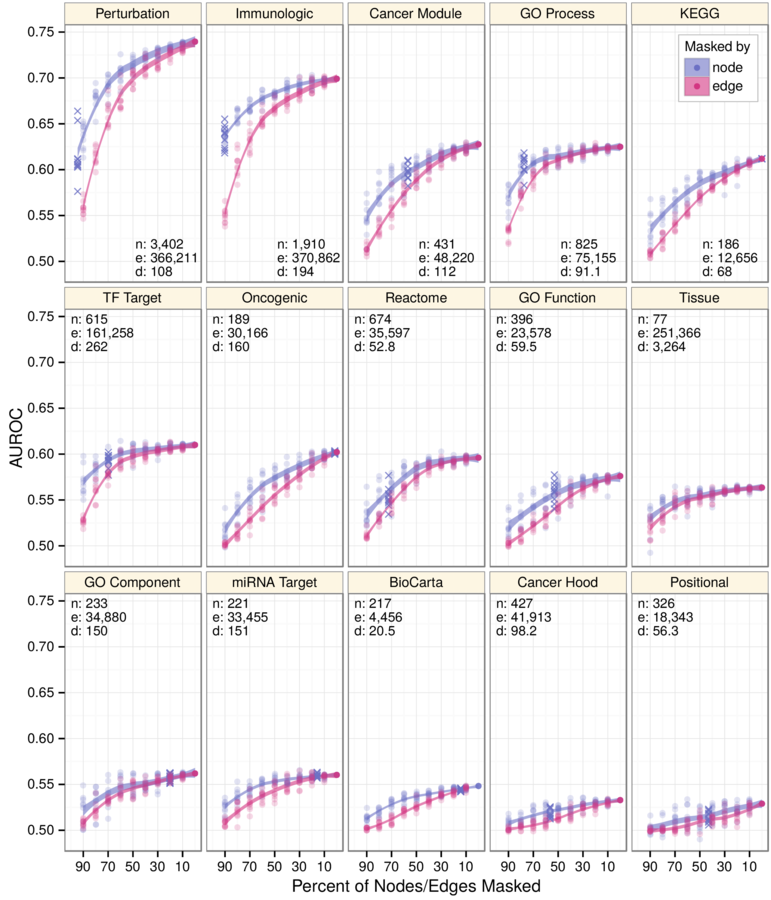
- robustness
Network subsampling
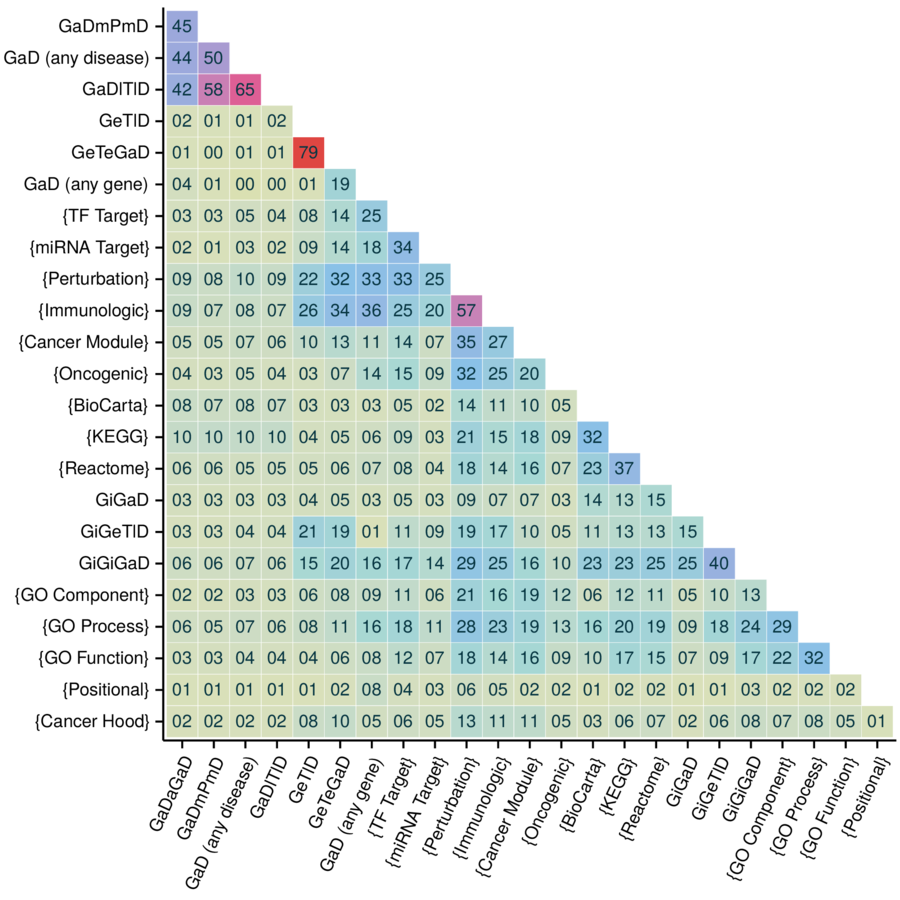
Feature redundancy
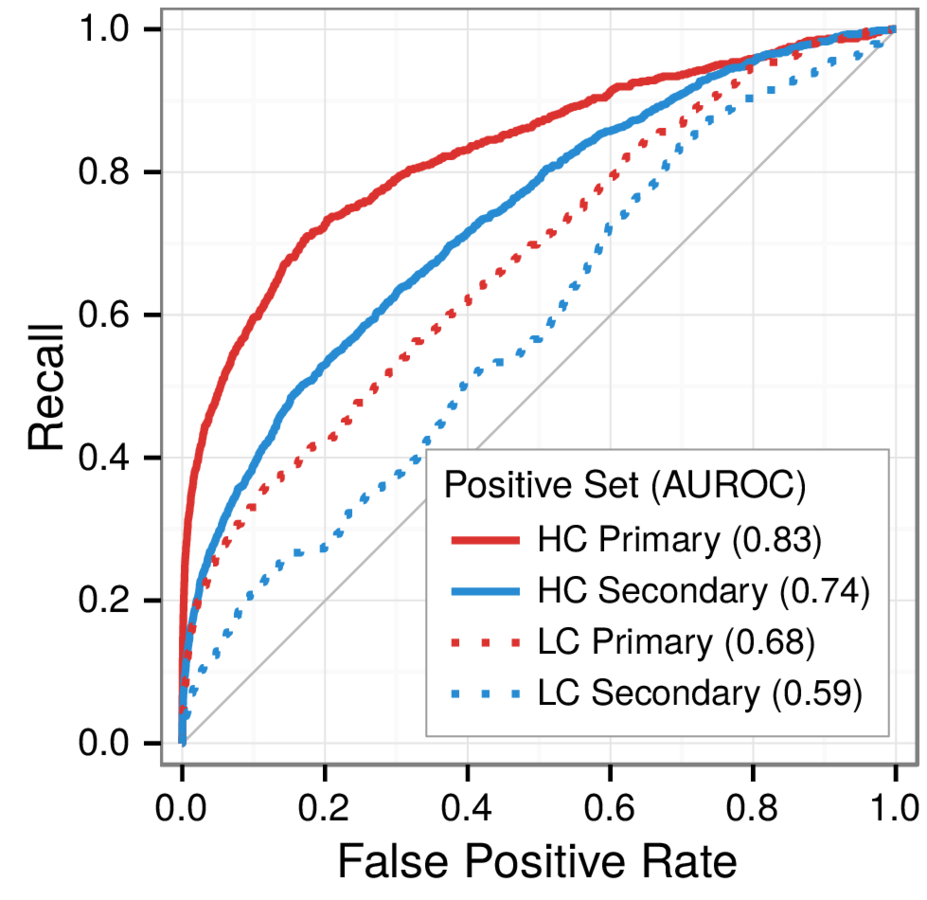
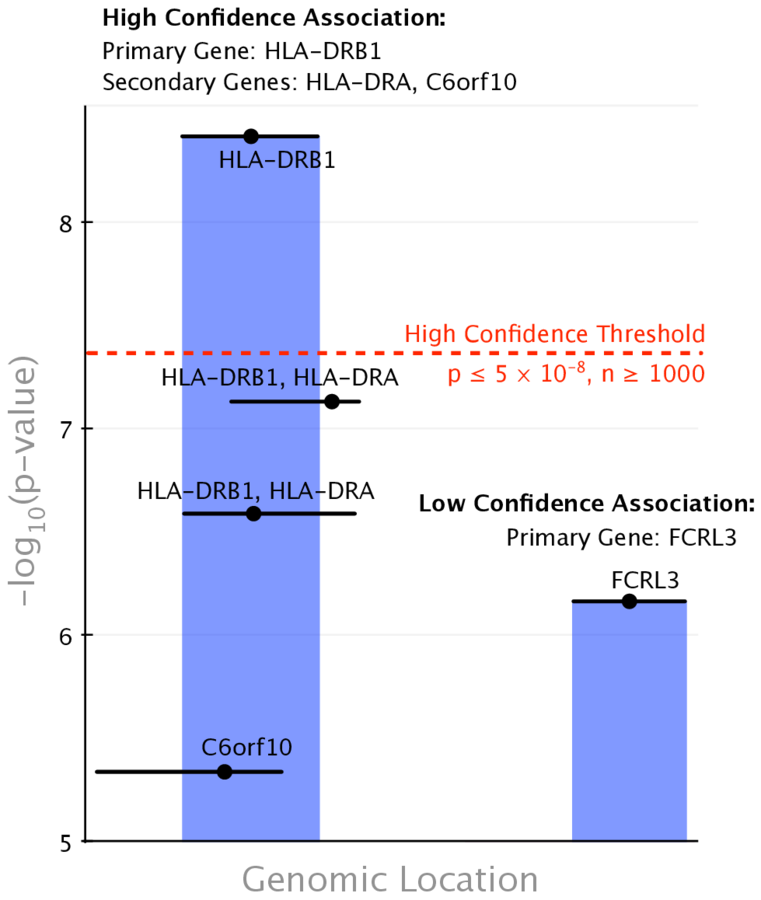
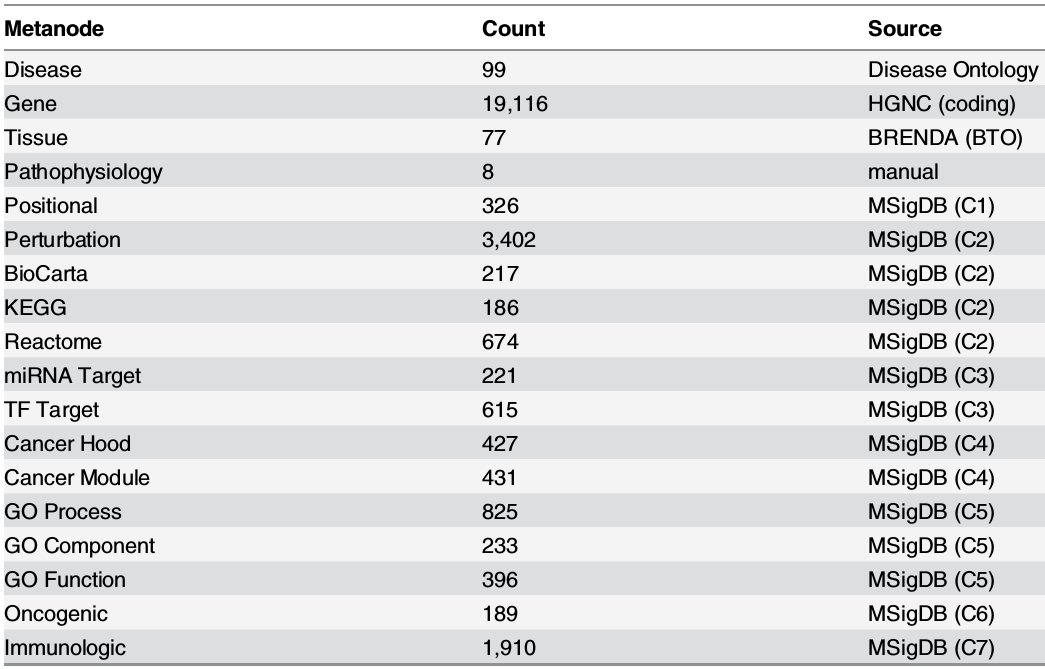
Metanodes (node types)
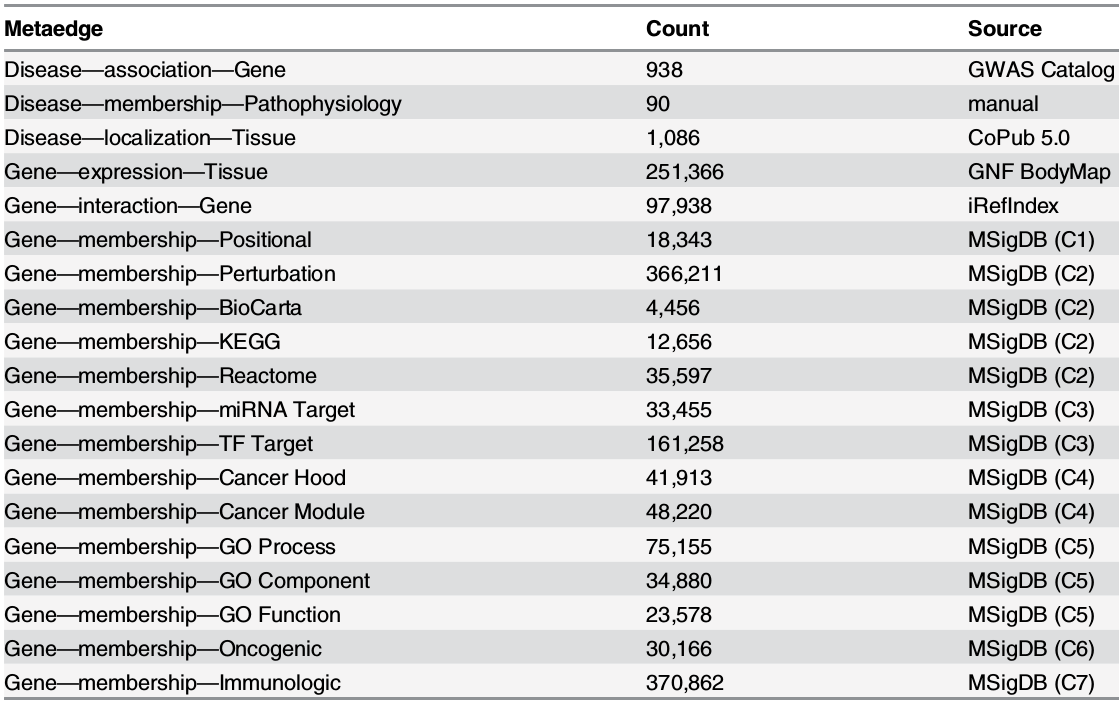
Metaedges (edge types)
Drug Repurposing
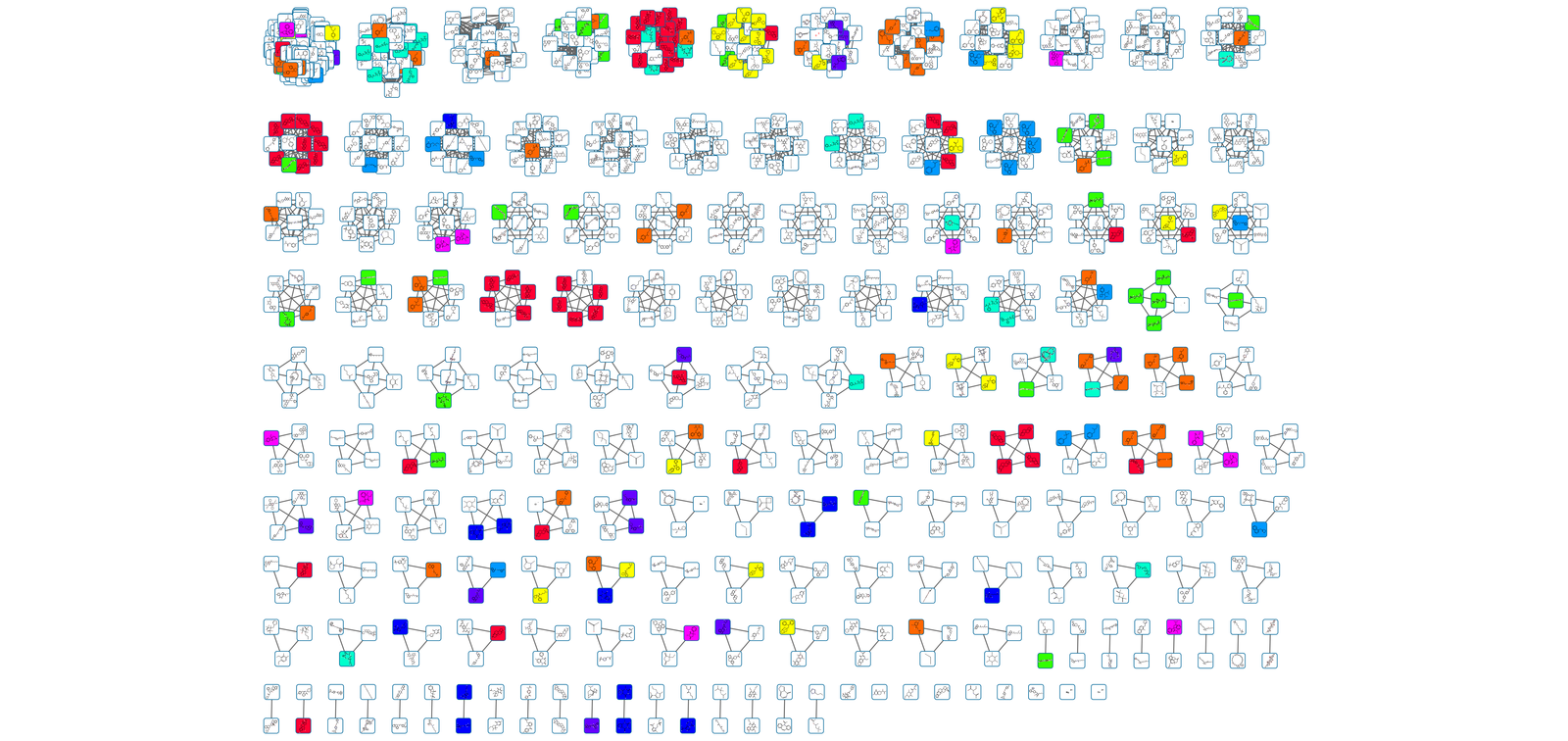
- expand our heterogeneous network into the realm of systems pharmacology
- predict indications between
- approved small molecules
- complex diseases
- drug repurposing
- learn the mechanisms of efficacy
17 contributors from 13 institutions
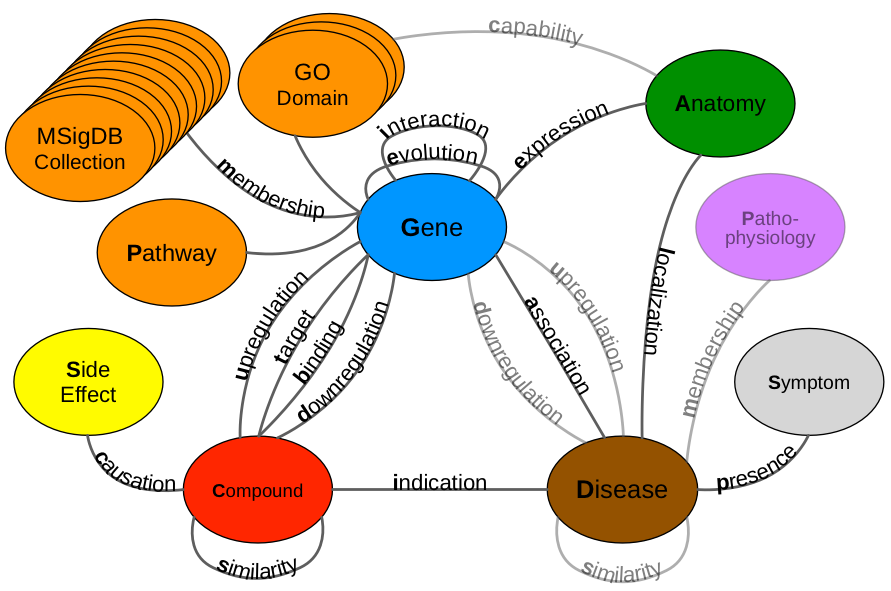
drug repurposing metagraph
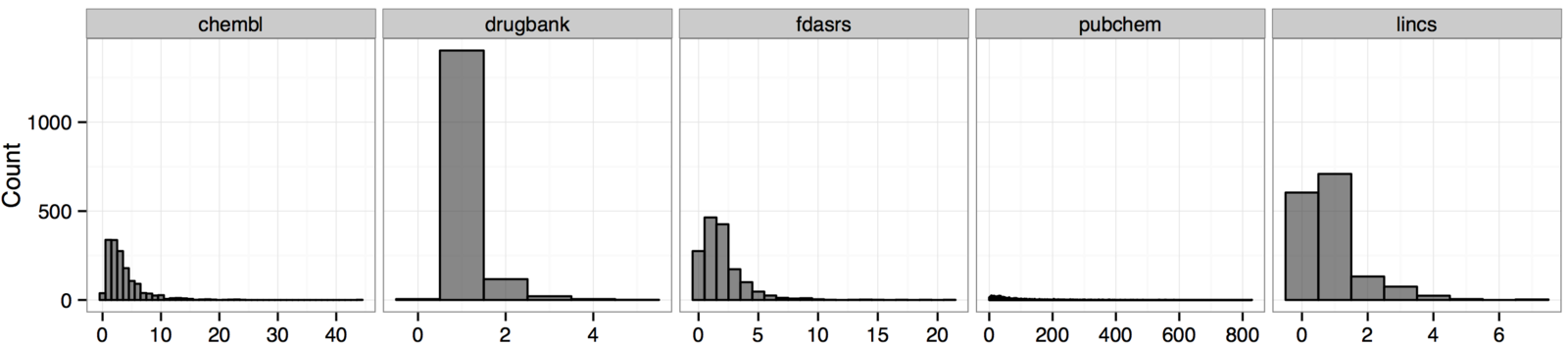
Unifying compound vocabularies
- compounds = DrugBank approved small molecules with structures
- Compound cannot be reliably matched by names
- Structures offer a solution
- Exact matching is limited -- small structural variations
- UniChem - matches on FIKHB components
- ThinkLab discussion
Unifying disease vocabularies
- Disease Ontology (DO)
- Mappings to other disease terminologies
- We require terms that are
- complex diseases
- distinct
- of clinically-relevant specificity
- well annotated in our resources
- DO Slim
- DOCancer Slim
- Non-cancers with GWAS
- 138 diseases
- ThinkLab discussion
building a catalog of indications
-
gold standard for our supervised approach
-
aggregated 4 resources:
-
MEDI-HPS – combines RxNorm, MedlinePlus, SIDER 2, and Wikipedia
-
ehrlink – linked data from health records
-
LabeledIn – expert and MTurk curated drug labels
-
PREDICT – UMLS links, drugs.com, drug labels
-
-
1,388 high and 1,114 low-confidence indications
Side Effects
- drug targets are poorly characterized
- embrace polypharmacology
- side effects may indicate unknown off-targets
- high-throughput methods
- drug labels (SIDER 2)
- FDA AERS (OFFSIDES)
- text mining
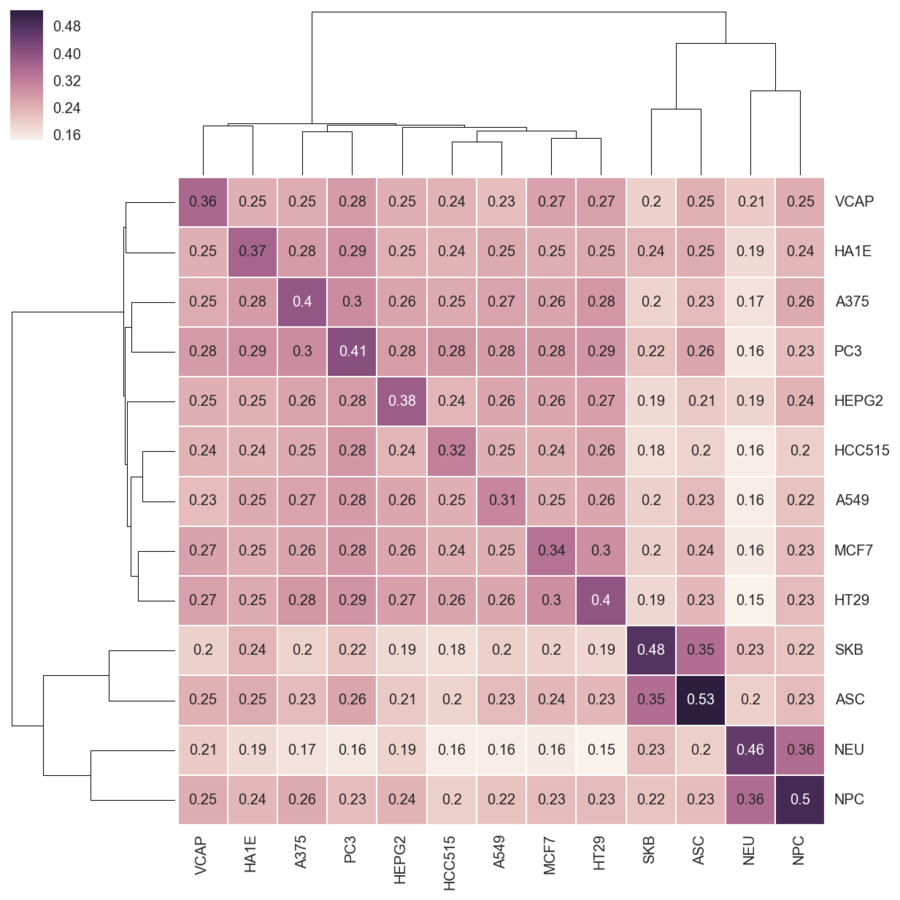
LINCS L1000
- Library of Integrated Cellular Signatures
- L1000 - Transcriptional profiles
- ~20,000 small molecules
- 978 measured genes
- Leo Brueggeman
- Resources
- http://slides.com/leoo/lincs#/
- https://dx.doi.org/10.6084/m9.figshare.1476293
- http://thinklab.com/d/43

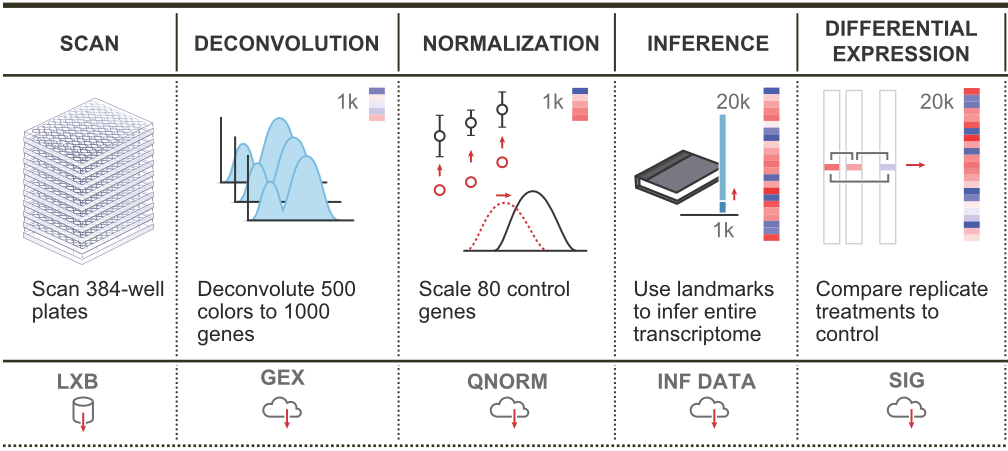
L1000 workflow
We combine all signatures for each DrugBank compound to get a consensus signature
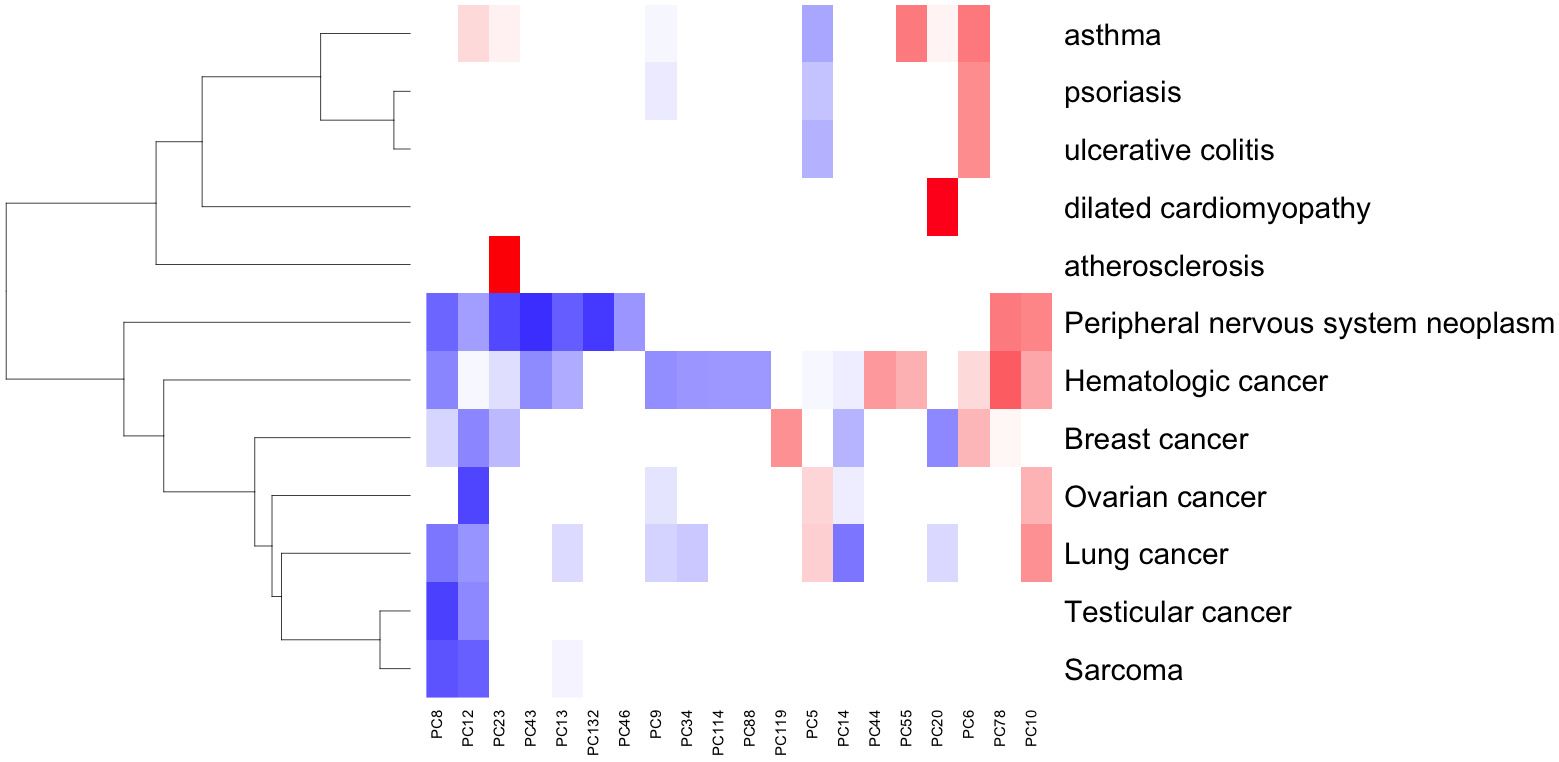
Disease-specific models uncover therapeutic signatures
Mining MEDLINE
http://thinklab.com/d/67
- curators read abstracts and annotate topics
- 21 million articles
-
5,594 journals
- cooccurence of two topics indicates a relation
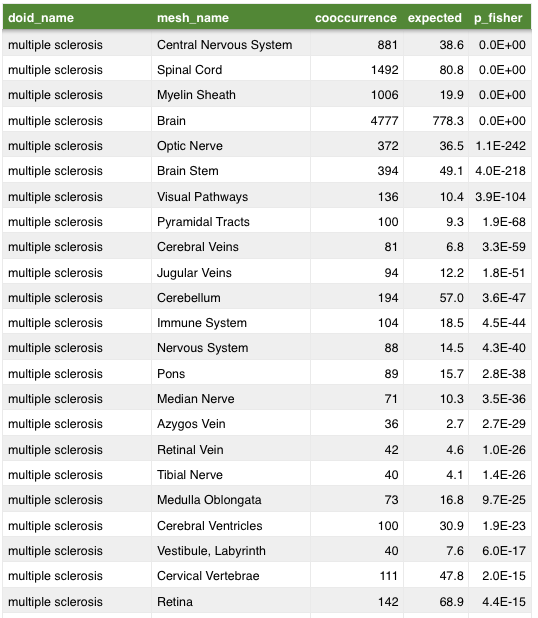
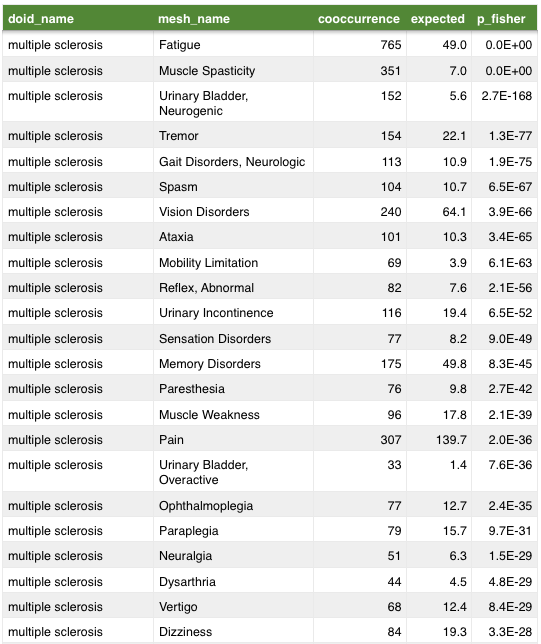
anatomy
symptom
Mining MEDLINE for disease context
Tissue-specific expression
Bgee
curates microarray and RNA-Seq experiments
http://bgee.unil.ch/TISSUES
transcriptomics, proteomics, text mining, UniProtKB
http://tissues.jensenlab.org/About
http://thinklab.com/d/81

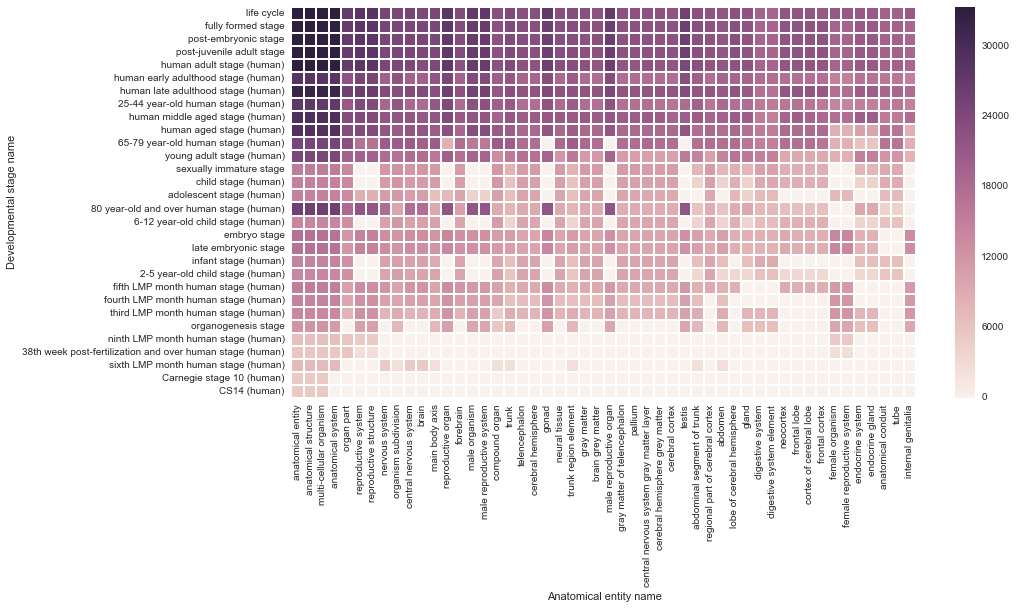
number of transcribed genes: developmental stage × anatomy
http://thinklab.com/d/81#293
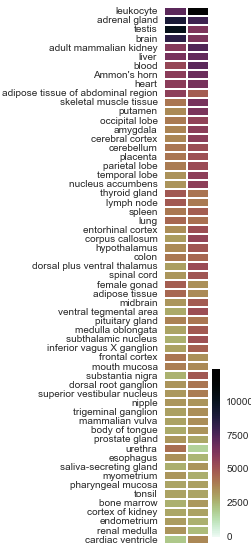
Number of enriched and depleted genes by anatomy
http://thinklab.com/d/81#293
-
depletion is more common than enrichment
-
highly-specialized tissues have the greatest differential expression
Next steps
- transcriptional signatures of disease (STAR-GEO)
- improved side effect resource
- removal of symptomatic indications
- discoveries!