GROBID Camp
Spring 2018
Patrice Lopez

ISTEX & bibliographical references
- GROBID 2014-2015: references perfectly recognized, f-score 60-65%
- Training completed for bib.ref. GROBID by ISTEX team based on 10k native publisher XML
➡ Perfectly recognized references: f-score up to 75%
➡ Estimated 90% for recent articles
➡ 1 millionPDF processed in 24h (Xeon 10 CPU, 10
GB RAM, 3GB used in average, 9 threads) - so 11,5 PDF/s
➡ very large Open citation dataset (at least ~10 times Open Citation corpus)

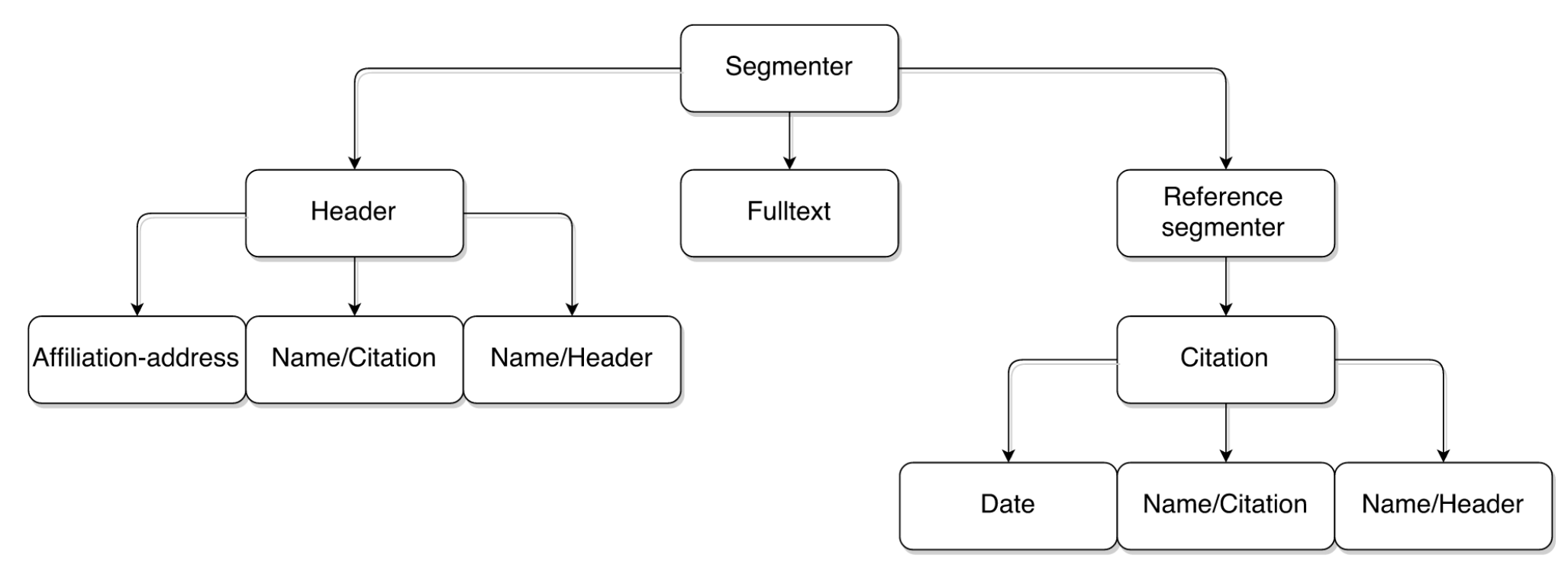
The legendary GROBID CRF cascade
Segmentation model
133 documents PMC/HAL
Addition of 91 documents ISTEX
Segmentation model
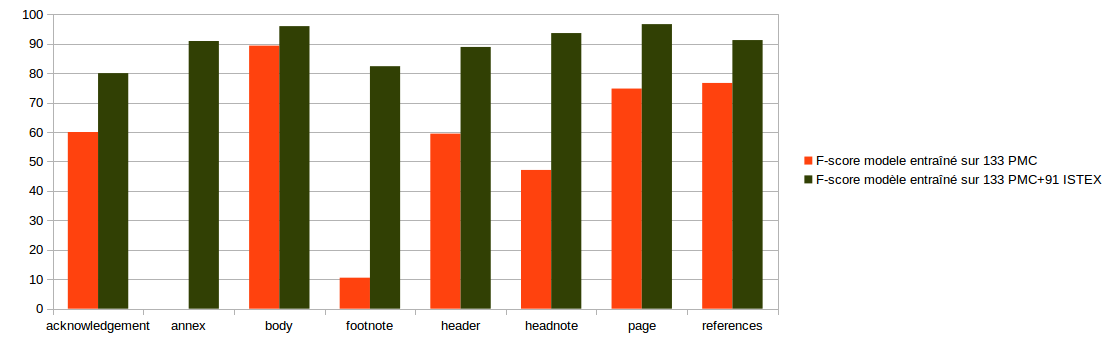
Evaluation on 19 ISTEX documents
Segmentation model
split 80-20% on 97 documents
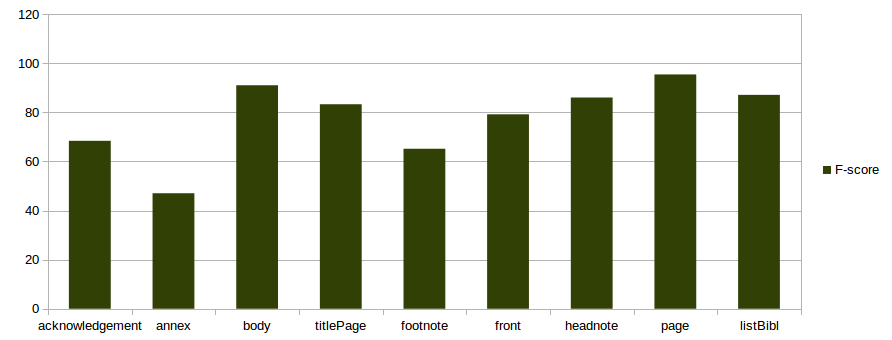
Full text model
20 documents de PubMed Central et HAL
Addition of 59 ISTEX full texts
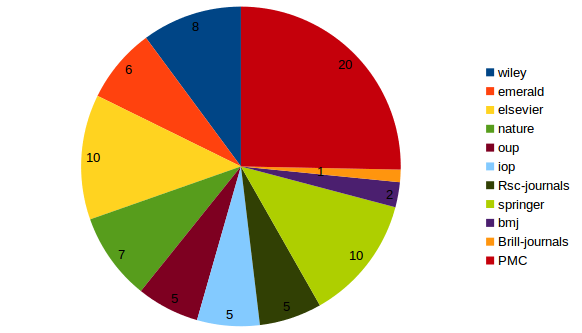
Full text model
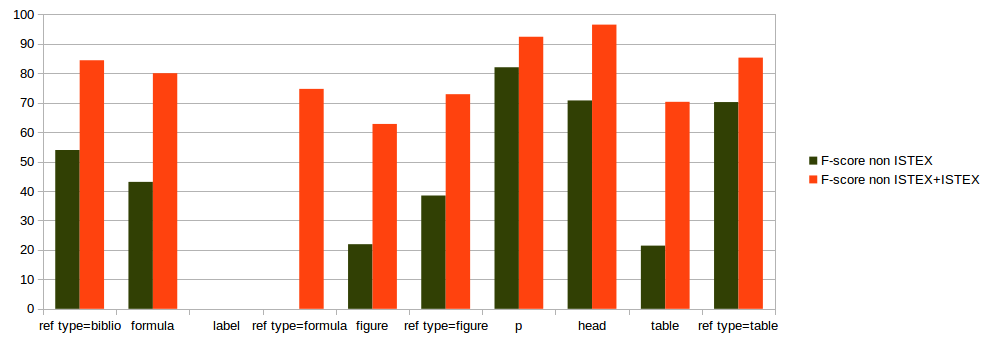
evaluation on 18 ISTEX documents
Full text model
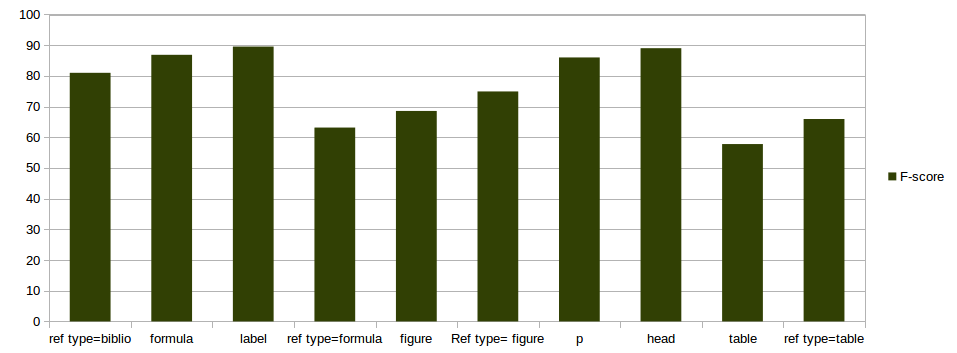
split 80-20% on 97 documents
Improvement on Pubmed Central API - Header model
Ratcliff/Obershelp Matching, similarity at 0.95)
===== Field-level results ===== end 2015
label accuracy precision recall f1
all fields 95.2 78.03 71.45 74.59 (micro average)
95.2 77.98 70.86 74.17 (macro average)
===== Field-level results ===== version 0.4.1
all fields 95.69 80.31 74.71 77.41 (micro average)
95.69 80.57 74.32 77.24 (macro average)
===== Field-level results ===== with current with new CrossRef API - v0.5.1
all fields 96.56 87.39 82.98 85.13 (micro average)
96.56 86.94 82.08 84.32 (macro average)
Support of CrossRef REST API
- Solution for title/author look-up in particular header model, not supporting the usual journal name/volume/first page query so weak for bib. references
- A "slow" consolidation based on old CrossRef OpenURL web API would still be necessary for bibliographical reference
- ... but we might simply implement our own resolver service by acquiring the complete CrossRef repo, subscribing to the new CrossRef Metadata APIs Plus Service (proposed since January 2018, with a SLA)
TEI-based full-text benchmarking
- We routinely evaluate GROBID with the PubMed Central sample, 1942 PDF + nlm files, in particular for each release to monitor accuracy, runtime, etc.
- Following an evaluation exercice of ISTEX R&D on automatic full text structuring with GROBID, we extended the benchmarking to PDF + TEI files, with TEI generated by Pub2TEI from the native publisher XML file
Current work in progress
-
Better PDF parsing: pdfalto (composed and special characters, reading order, spacing, etc.)
-
Structuring ebook (pdf/ALTO): training based on embedded "outline" (project Opaline)
-
Long due new header model: regenerate and reformat training data, new features, etc. targeting 0.90 f1 instance-based
-
New DL models for sequence labelling, and for text classification (Keras, efficient java embeddings)
-
it is challenging to make it production ready (loading of resources, native integration, memory usage)
-
Thanks !
Patrice Lopez
