Some updates on what I've been up to
+ fMRI preprocessing with fMRIPrep
+ some tools I like to use
Lab meeting
2026-04-01
Just look at these.



Now look at these.






And one last time.



Something changed. But what?




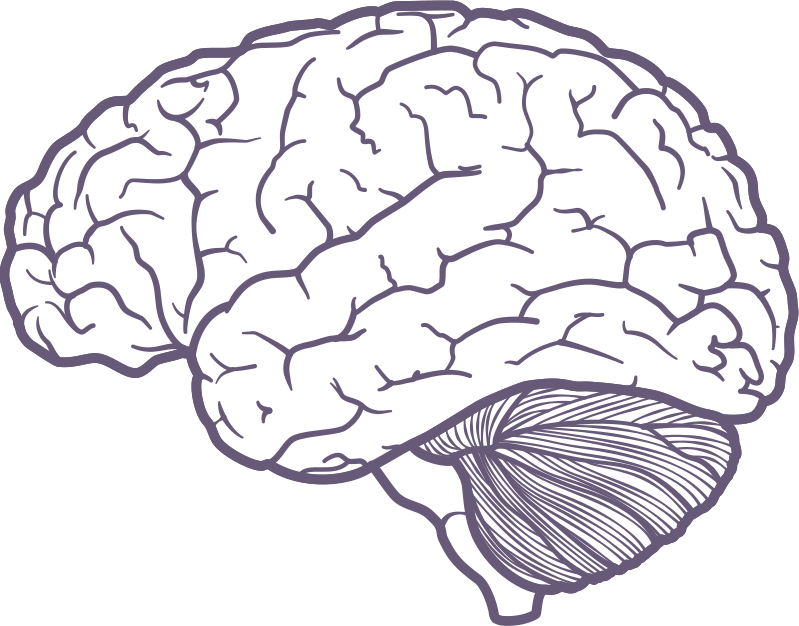


Top-down feedback?
Persistent change in early representations?








COLLECT ALL THE DATA!







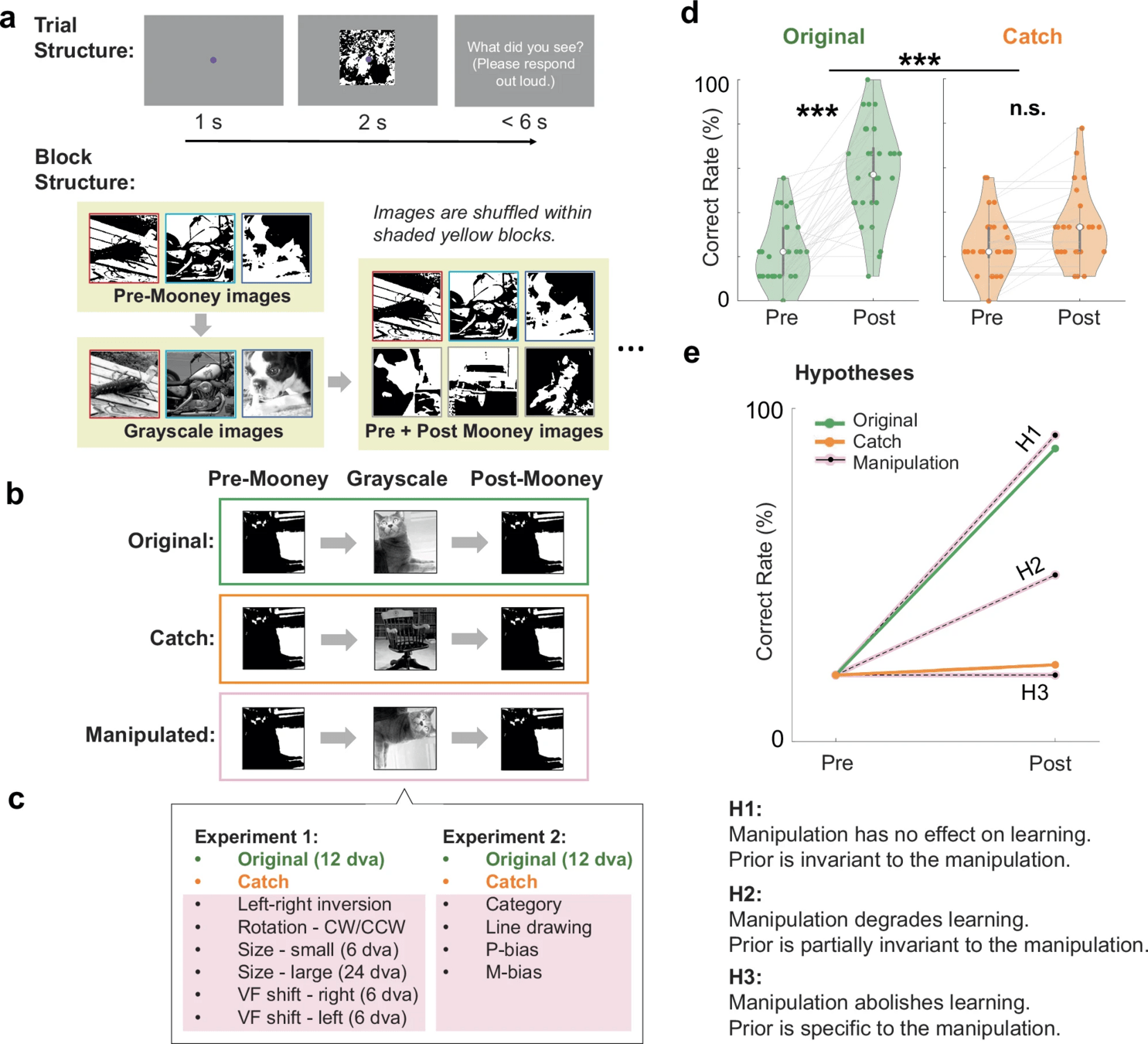
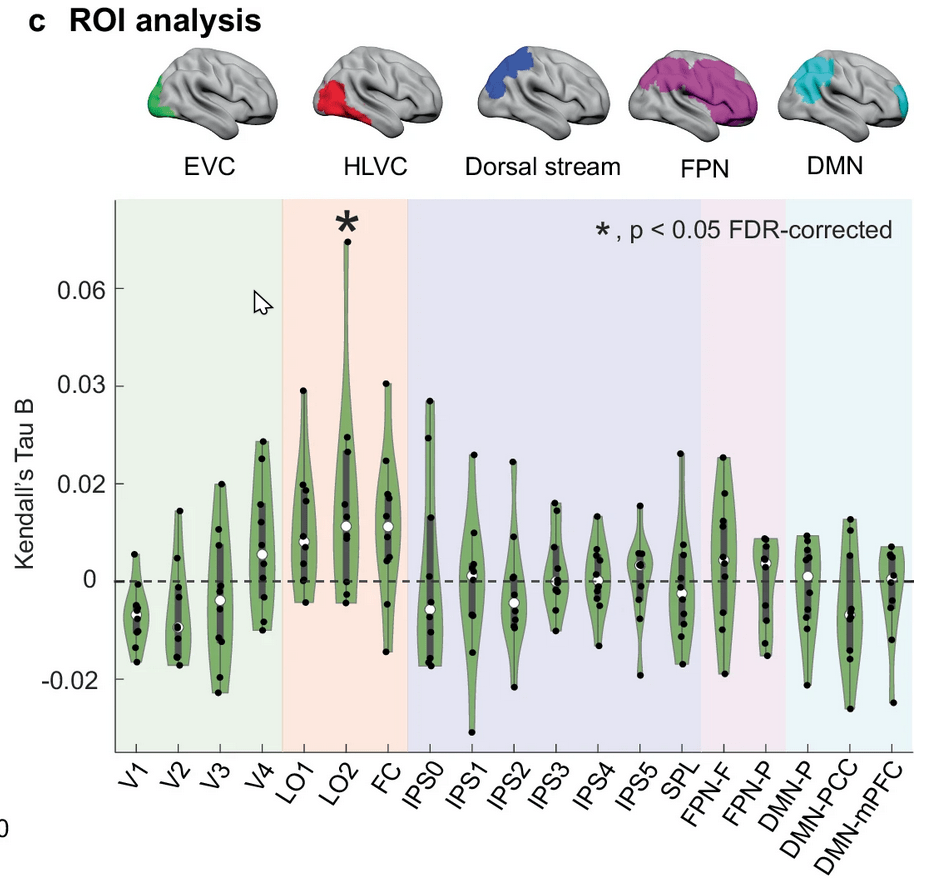
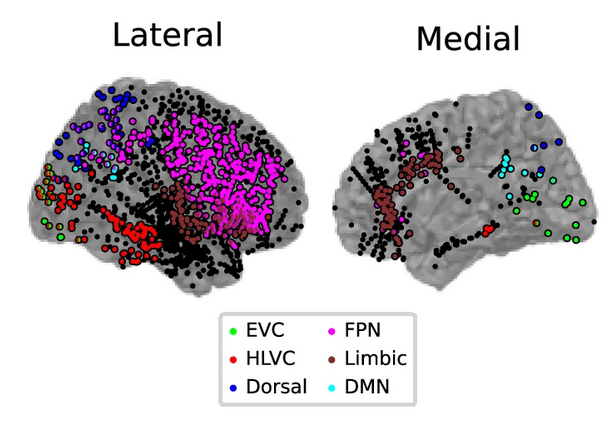
... fMRI ...
... sEEG !?
Similar design ...
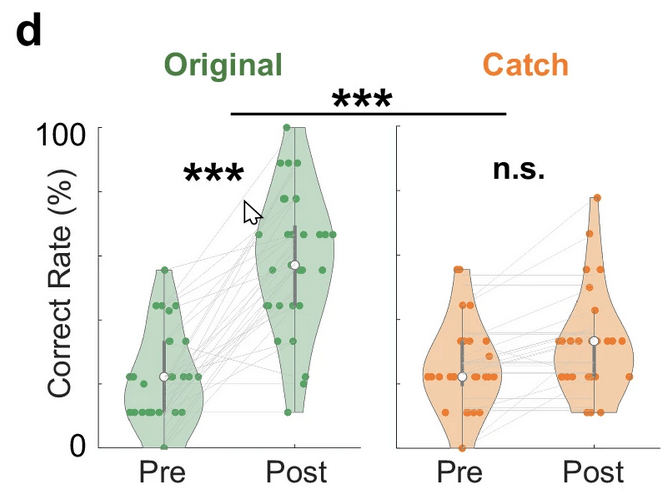
... behavior ...
😞


The neural developmental trajectory is unclear.



MTL lesions preserve 1-shot perceptual memory

1-shot perceptual memory involves widespread cortical changes
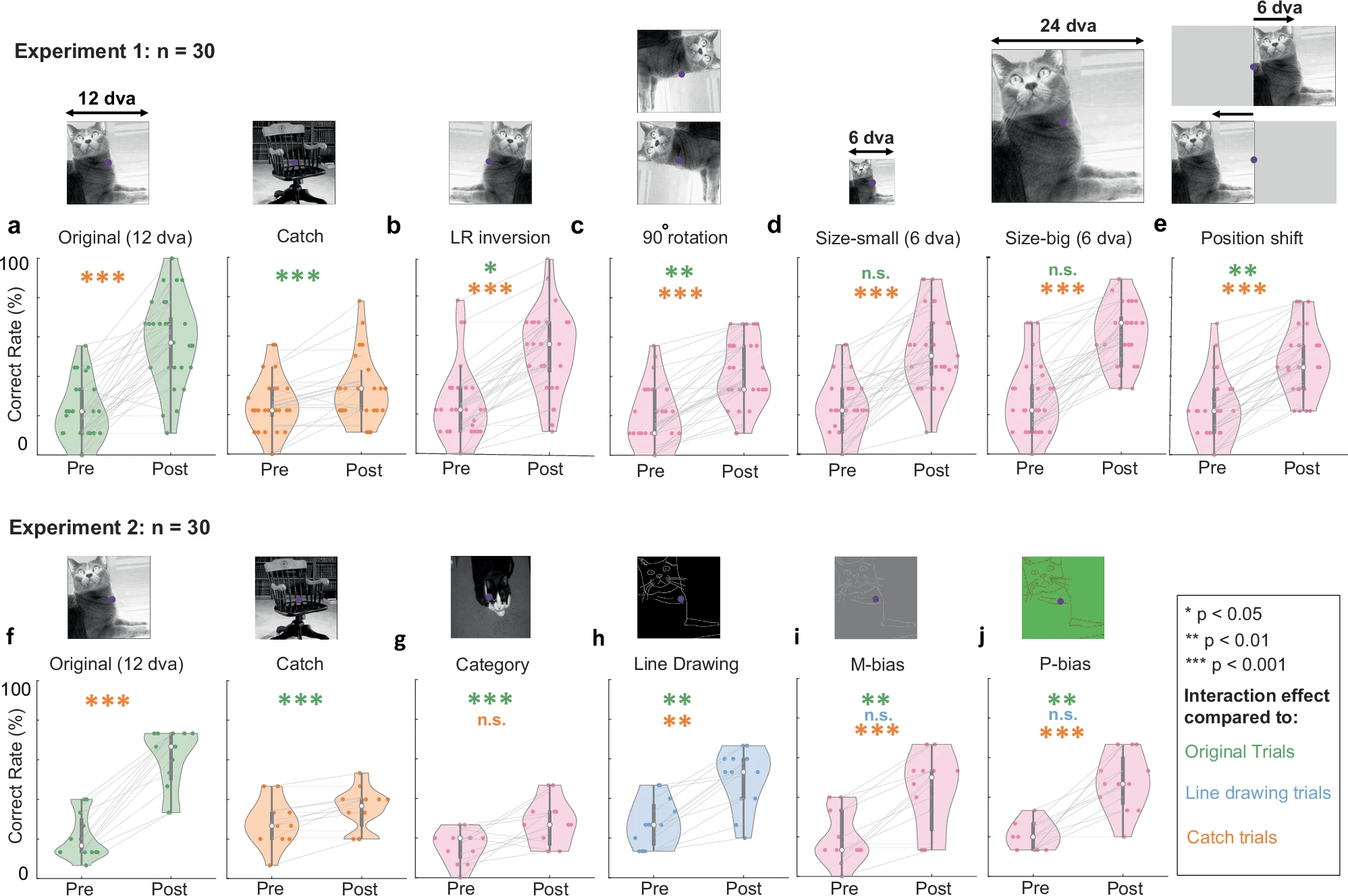
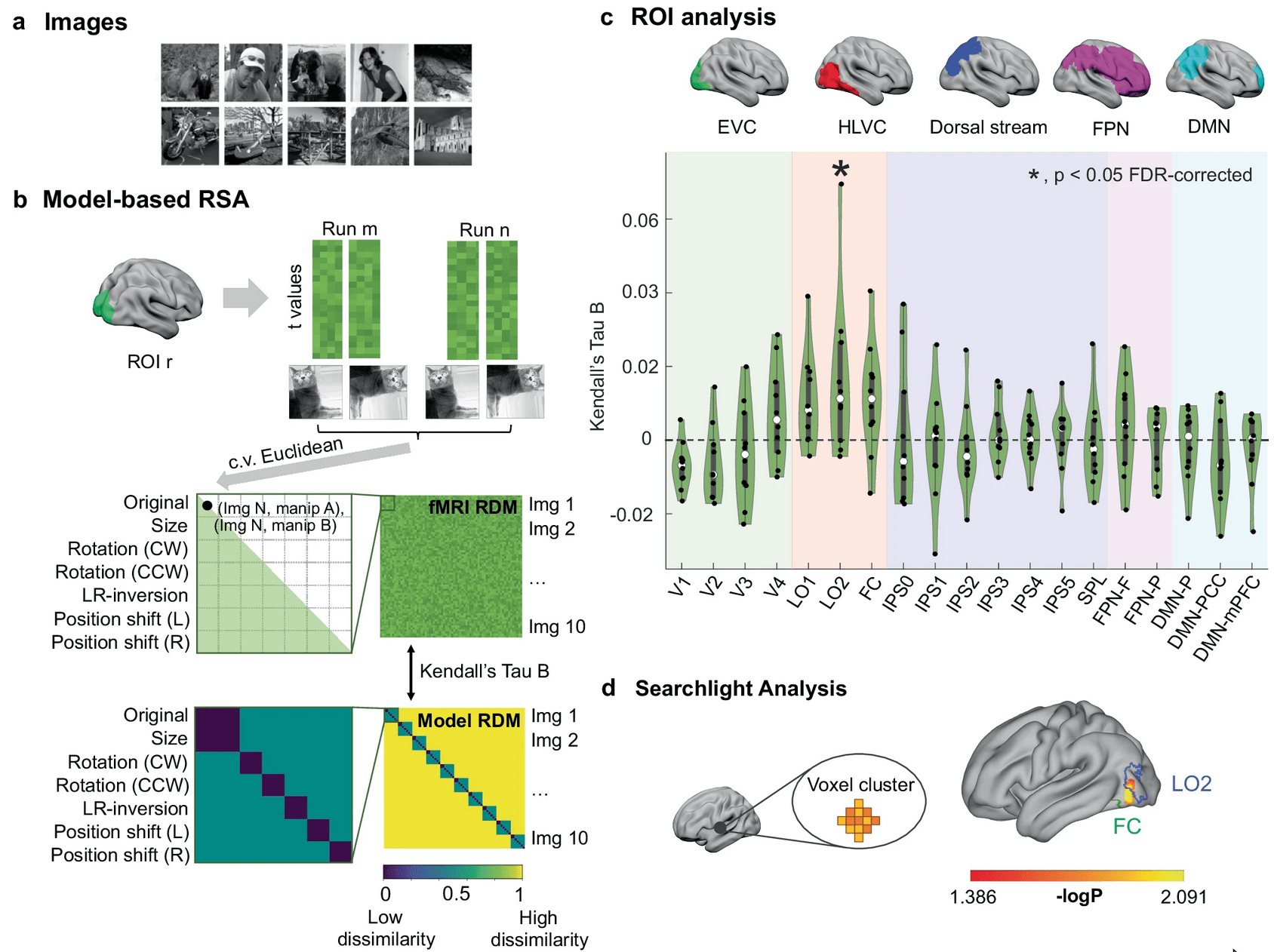
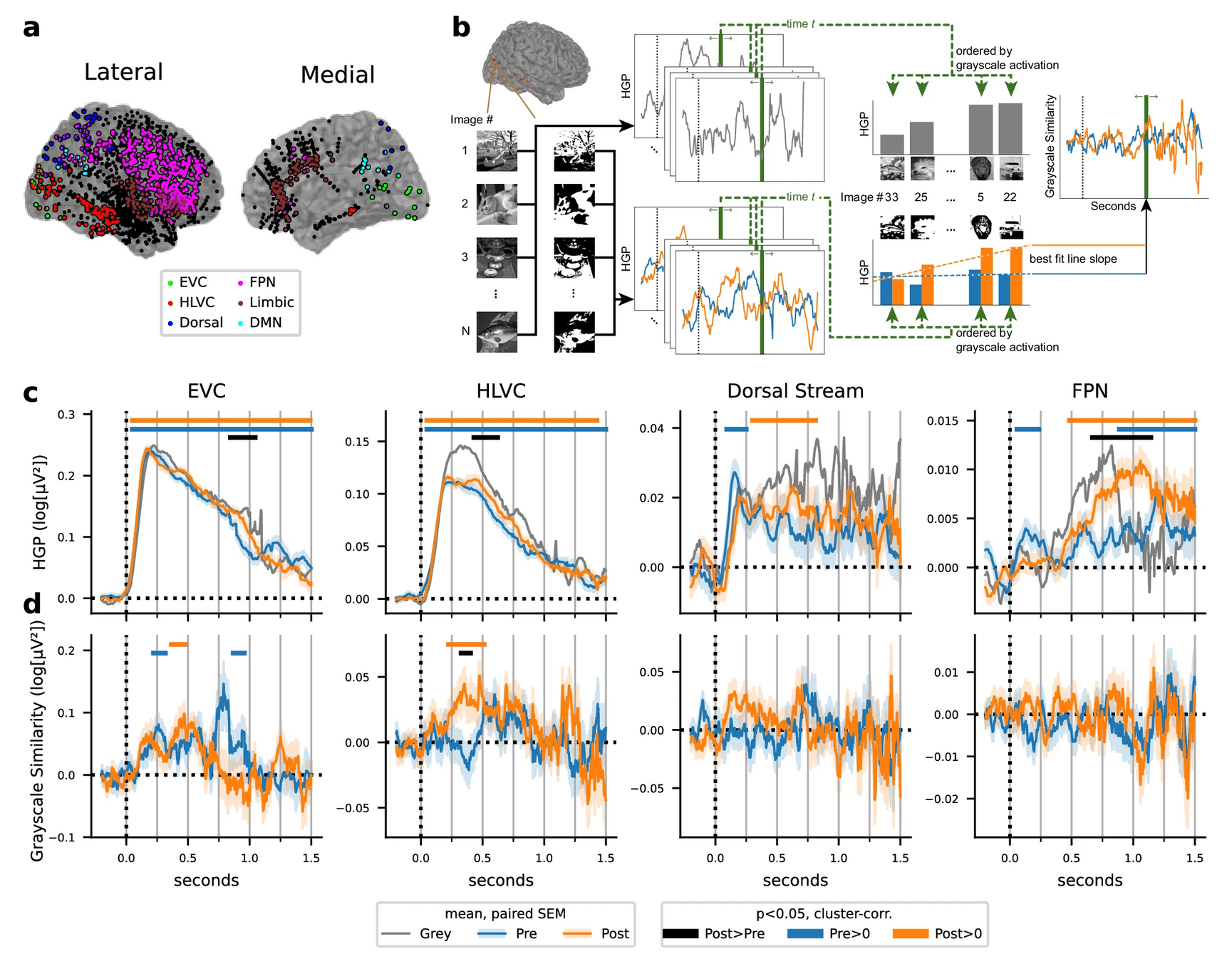

COLLECT A LITTLE BIT OF DATA!
(1 participant)





Interim progress
Preprocessing!
Simple contrasts!
Learning a lot of methods!
Note to self:
Show sample results


Why bother with fMRIPrep?
Standardization is great.
(Decisions are hard. Let other people make them!)
- Enforces BIDS: easy to release data.
- Reproducible.
- Pipeline is less likely to have errors/bugs (likely much better code than I can write).
- Automation reduces a lot of manual effort.
- Excellent quality-control reports.
- Uses standard tools: easy to peek under the hood.
- Well-documented; large community.
- Commonly (increasingly) used in the field.
- Language-agnostic (if you don't peek inside).
- (Mostly) free and open-source.
- Can (supposedly) be interrupted and resumed.


*
A super-quick recap of (f)MRI
- Most things are made of atoms, which have nuclei
- Some nuclei have something called "spin", which
- has a magnitude
- has a direction,
- can add up with spins of other nuclei, and
- can be affected by a "magnetic field" (!)

Aligned spins add up a lot
Random spins tend to cancel out

- Brains are made of water (H2O)
- Water is a thing, and so has nuclei
- Turns out, the hydrogen nuclei (protons!) have spin
- But they're randomly oriented.
🌊
🌊
🌊
🌊
🌊
H
H
O




When you put the brain in a magnetic field, these spins tend to align with it.
And so the total spin increases.
("net polarization")
this is the main magnet in the MRI machine (+"gradient coils")



If you shine just the right color of light on the brain for a tiny bit*,
you can flip these spins!
*this is a "radio frequency (RF) pulse" created by the "RF coil"






If you wait, this net spin will gradually disappear.
This "relaxation" time* varies across the brain, depending on what tissues are nearby.
And as this happens, the brain emits some light!
*e.g. T1, T2









If you collect this light, using fancy mathemagics, you can plot relaxation time across the brain!
high spatial resolution: ~1 mm
this is measured using the receiver head coil
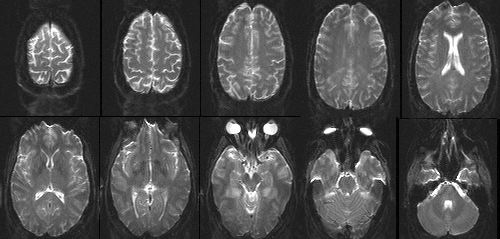
The presence of nearby oxygenated blood also changes the relaxation times of protons.
So we can measure a "blood oxygen level dependent" signal too.
worse spatial resolution (>2 mm), but temporal resolution of ~1 s 🤷
"echo-planar imaging (EPI)"
🩸
🩸

vs


Typical fMRI experiment
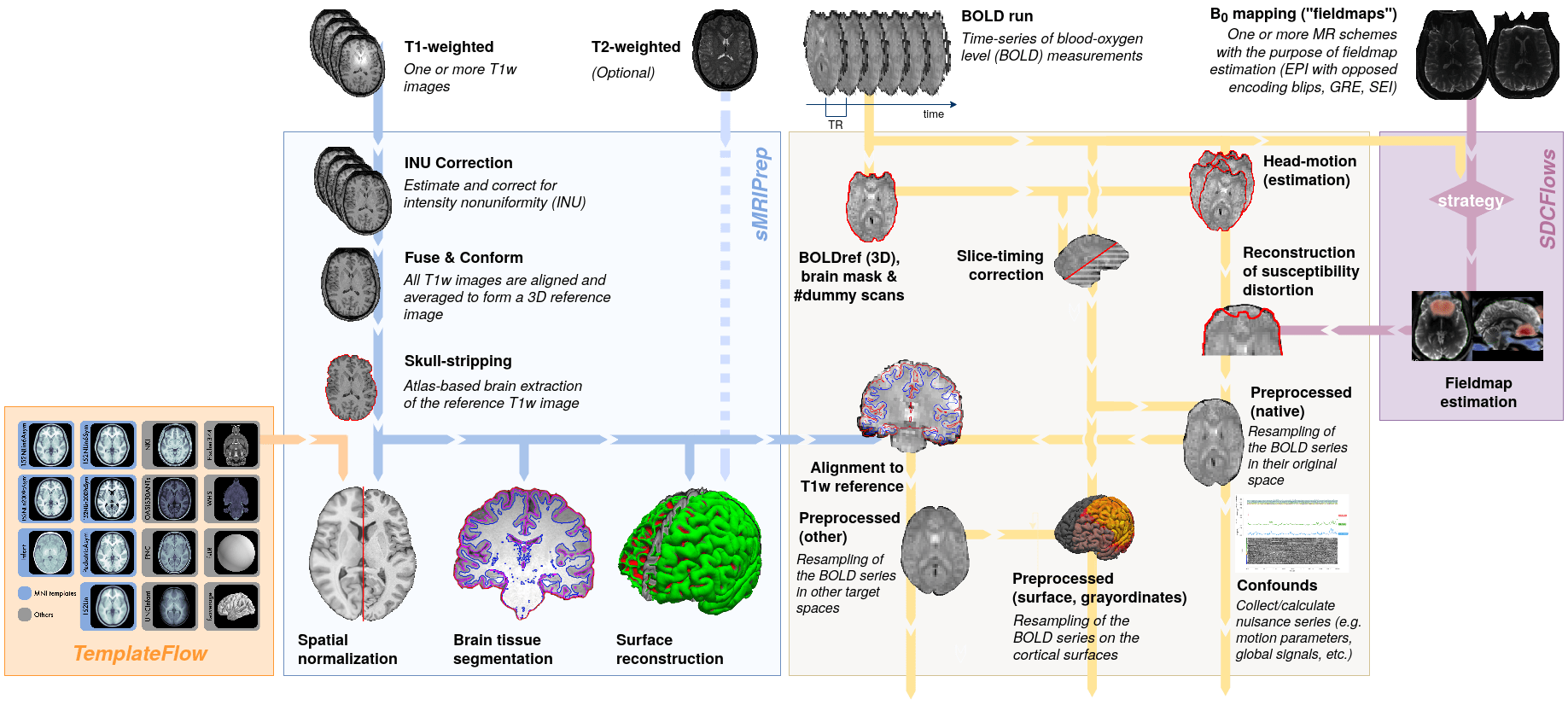


- high-resolution (1 mm)
- "anatomical scan"
- T1-weighted image
[x, y, z]
- low-resolution (1 mm)
- "functional scans"
- EPI images
- measured over time
[x, y, z, t]
- measures inhomogeneity in the magnetic field
- used to correct EPI data

What fMRIPrep does for you
Note to self:
Show example output.
How to use fMRIPrep?

Deface your data.
(I use mri_deface from the container's FreeSurfer)
Do preliminary QC.
(LPT: giving up early saves time!)

Build the container with all the dependencies. (or use the shared one on Mind)
apptainer build /path/to/fmriprep-container.simg docker://nipreps/fmriprep:{version}Get a FreeSurfer license.
--fs-license-file
/freesurfer/license.txtHow to use fMRIPrep?
Specify the output spaces you want.
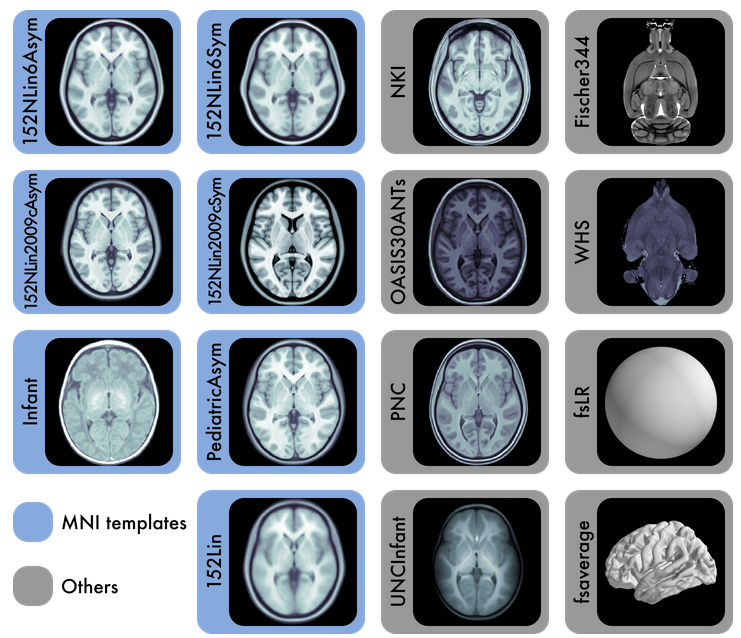
any template from TemplateFlow
(default MNI152NLin2009cAsym)
[they have kid templates too]

or non-standard spaces
How to use fMRIPrep?


Run the fMRIPrep command
likely no need to adjust other defaults
(Optionally) Add a lesion mask, e.g. for hemispherectomy
How to use fMRIPrep?
Wait for a long time.
(several hours, ~3 h?, mostly FreeSurfer's surface registration)
If you're lucky, everything ran without errors.
Inspect the quality assessment report.
(sub-XX.html)
How to use fMRIPrep?
What next?
fMRIPrep only does preprocessing, not any data analysis.
(e.g. GLMs, contrasts, thresholding, MVPA, etc.)
But it gives you useful confound regressors.
(e.g. motion, noise components from WM/CSF/outside the brain)
Use whatever tools/code/languages you like to analyze the (standardized) outputs of fMRIPrep!
sub-{subject}_task-{task}_run-{run}_space-{space}_desc-preproc_bold.nii.gz
My take
It's a lot of work upfront to organize the data well enough for fMRIPrep to run.
But it's worth it for the peace of mind and ease of releasing data afterward.
And honestly, it's easier than running individual steps manually and stressing about setting hyperparameters.
References (and resources)
Highly recommended:

Code review?
(Credit: Maria!)
