From the wet lab
to the web lab
Anisha Keshavan
University of Washington
eScience Institute & Institute for Neuroengineering
Motivation
In Medicine
- Even though we are all unique individuals with unique biology
- We are treated the same way

Precision Medicine
What if we tailor treatment
to your unique biology?


Meet my mom
- Oncologist
- Colon cancer
- No risk factors
- Side effects of treatment
- Not a "mistake"

Treatment = f(Disease)
Treatment = f(Disease,Genotype,Phenotype,Environment)
So, where is precision medicine?
We must bridge the domain divide, first
A single domain

A domain divide

We need a web

We need the web
Goals for this talk
- Web-based tools for MRI neuroimaging
- Bridge the domain divide
- Not a neuroimager?
- what are your bottlenecks?
- can the web help?
Background
PhD: Multiple Sclerosis
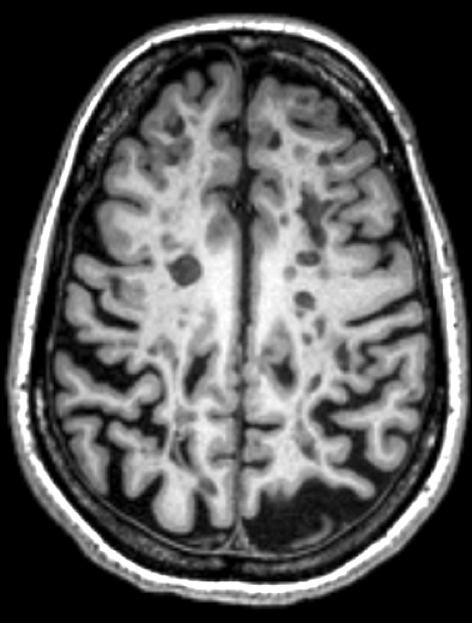
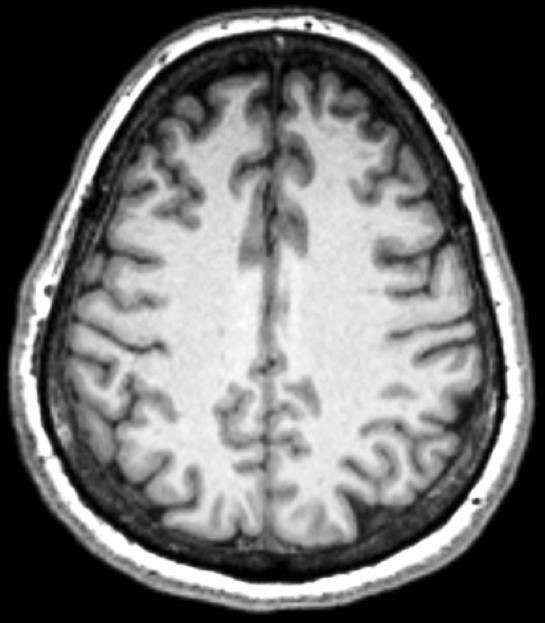
Disease Progression
Relapse Remitting vs Progressive
Treatment = f(Disease progression)
But we don't know Dp early on
A clue to Dp ?


Big Data Collection



500 patients, 12 years
= 6,000 MRI scans
Data Analysis Workflow









MRI
Algorithm
Image Features
Treatment
M.D
Reality








MRI
Algorithm
Image Features



Treatment
??
??
Expertise does not scale




MRI
Algorithm



And the domain divide makes it hard
Computer Science
Radiology / Neuroanatomy
High dimensionality = lost intuition




Image Features x 1000

??
??


Statistics
Neurology / Neuroscience / Biology
We need data visualization & exploration
Web Interfaces
to cross the divide
Postdoctoral Research


Data Science
Domain Science
Scaling Expertise


Data Science
Domain Science
Data Scientist
- Needs image annotations
- In a machine-readable format
Domain Expert
- Know UNIX to annotate image
- Save annotations consistently


Mindcontrol
A dashboard for collaborative brain quality control and annotation
Mindcontrol
Mindcontrol
A fruitful collaboration @ UCSF
(under review)
-
Also deployed at:
-
UT Austin Medical School
-
NIMH
-
University of Cambridge, Turing Institute
-
Now, as postdoc:
studying
Mental Health
-
Public dataset, 10k images
-
Motion → bad quality images
-
Visual inspection before & during analysis
-
Too time consuming in the “wet lab”
Let’s try the web lab
Healthy Brain Network
Scaling expertise in the web lab
- Start with small, expertly labelled training set
- Train citizen scientists to amplify training set
- Train a model on amplified labels
Expert Feedback
Mindcontrol is nice
but I want to play on my phone"
Dr. Satra Ghosh, MIT
Scaling Expertise in the web lab
- Start with small, expertly labelled training set
- Train citizen scientists to amplify training set
- Train a model on amplified labels
Braindr
Brain Data Review
Keshavan, Anisha, Jason Yeatman, and Ariel Rokem. "Combining citizen science and deep learning to amplify expertise in neuroimaging." bioRxiv (2018): 363382. (under review)
Instructions


Swipe Left
Swipe Right
Win Prizes

Reactions

Many Ratings

But often, no agreement

How to aggregate unreliable ratings?
Need to "down-weight" some raters
| image id | rater 0 | rater 1 |
|---|---|---|
| 1 | 0 | 1 |
| 2 | 1 | - |
| Expert |
|---|
| 1 |
| 0 |
each rater is a feature
each image
is a sample
with a model that handles missing data

Rater importance vs # ratings

Corrected Ratings

Improvement over average
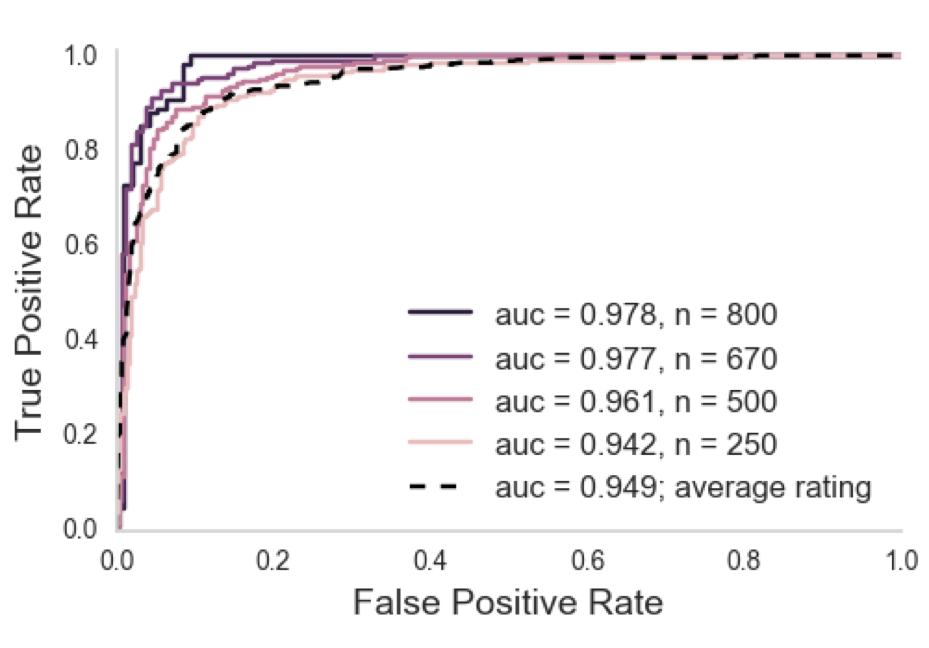
held out AUC
Scaling expertise in the web lab
- Start with small, expertly labelled training set
- Train citizen scientists to amplify training set
- Train a model on amplified labels
Predict image quality

> 100,000 ratings
> 400 users
0.99 AUC = near perfect
How do I make my own web lab?
maybe you're asking


Web Technology
Domain Science
Swipes for Science





Thanks eLife Innovation!
Data Visualization & Exploration


Data Science
Domain Science
Linked Visualizations
Rule #1: always create
Brain Morphometry
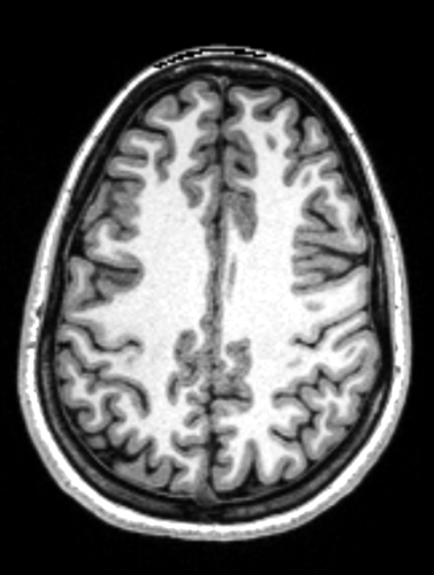

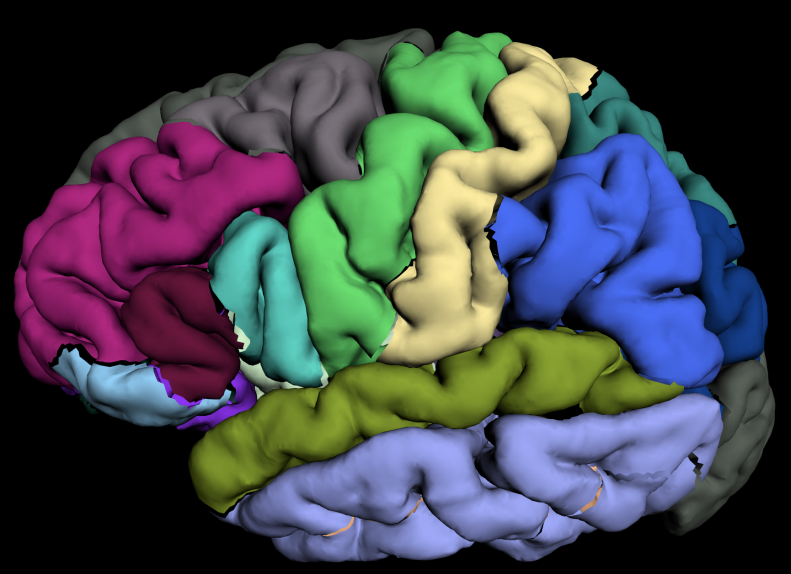
??
Tables.
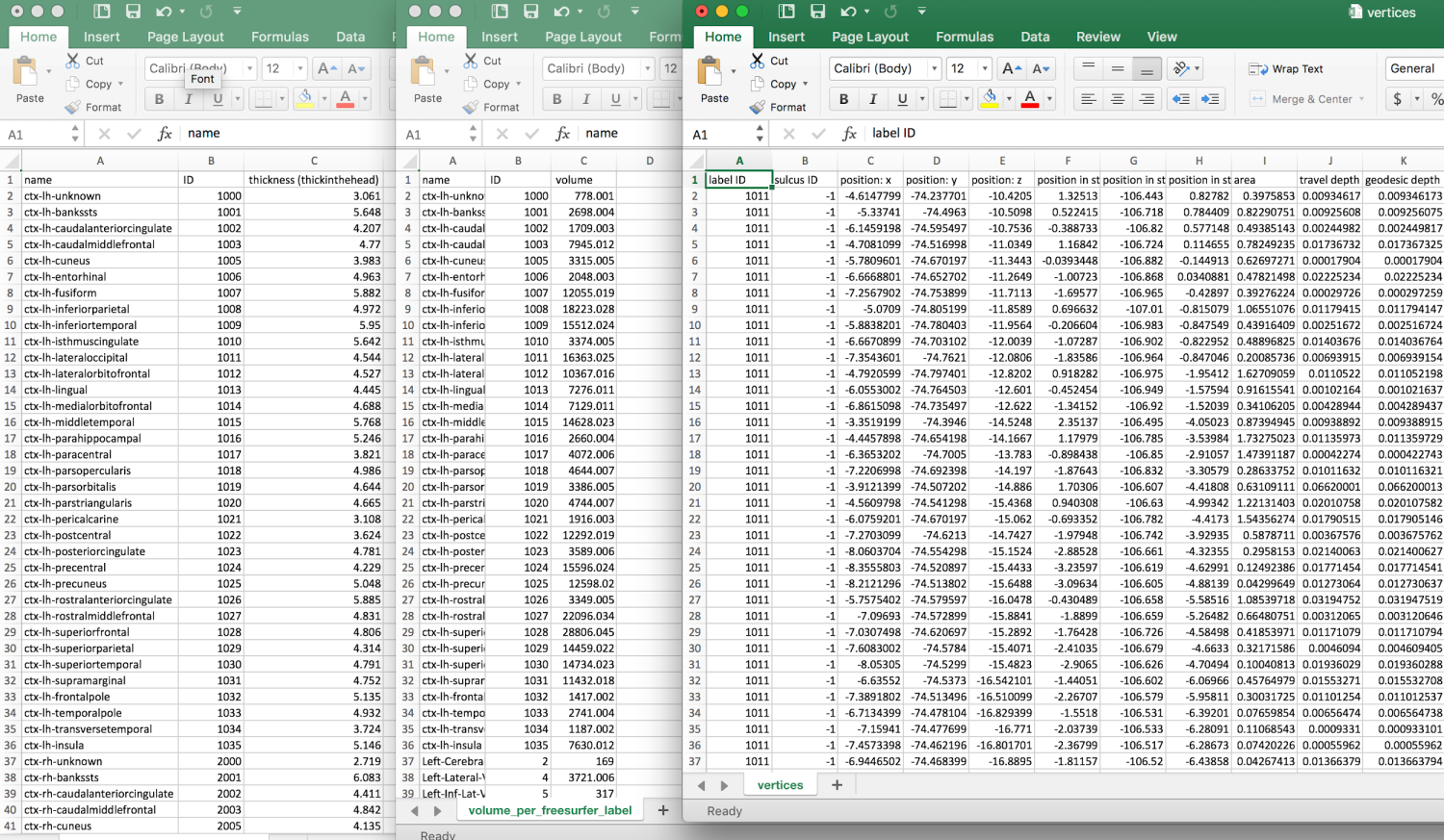
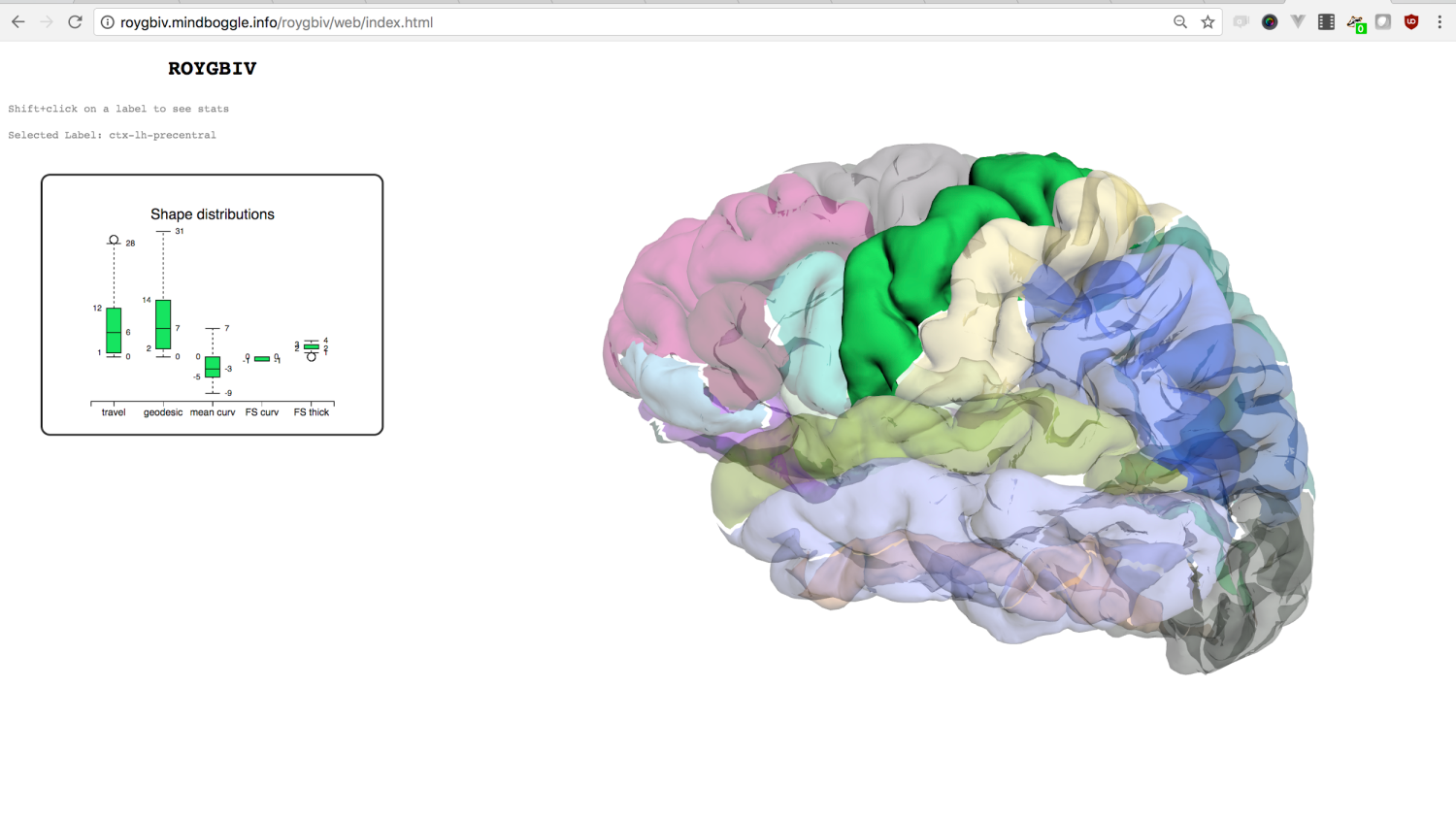



Klein A, Ghosh SS, Bao FS, Giard J, Häme Y, Stavsky E, Lee N, Rossa B, Reuter M, Neto EC, Keshavan A. Mindboggling morphometry of human brains. PLoS computational biology. 2017 Feb 23;13(2):e1005350.
Keshavan, Anisha, Arno Klein, and Ben Cipollini. "Interactive online brain shape visualization." Research Ideas and Outcomes 3 (2017): e12358.
Information Visualization
Scientific Data Visualization
Link
Share
Rule #2: make it easy to
Diffusion Imaging
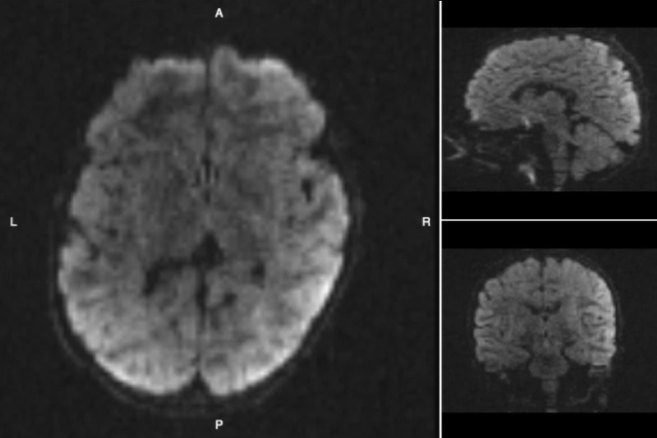
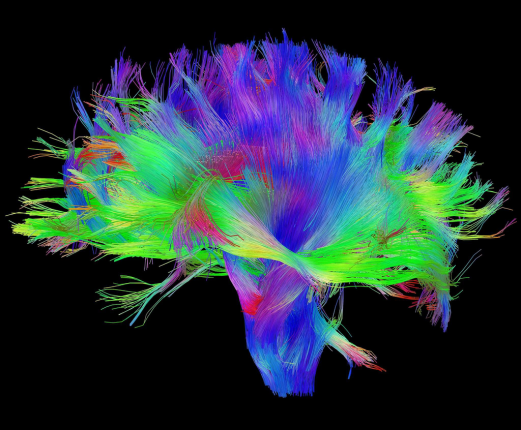
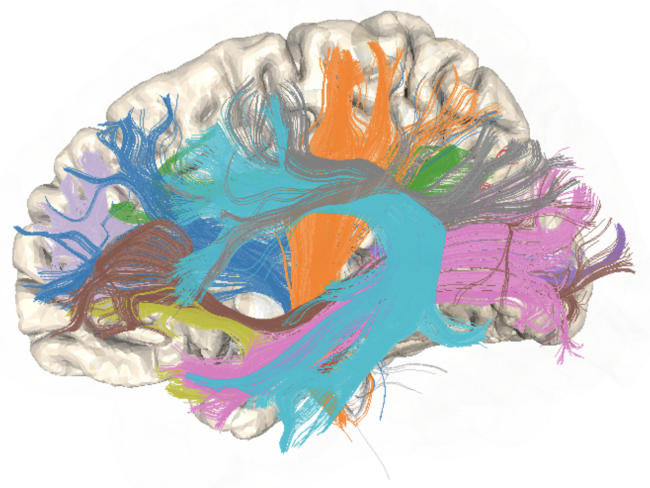
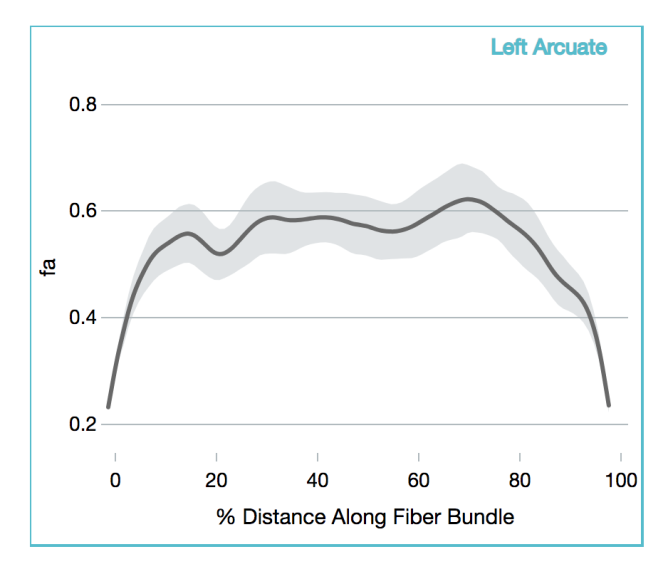
Tract Profile

AFQ-Browser
Yeatman, J. D., Richie-Halford, A., Smith, J. K., Keshavan, A., & Rokem, A. (2018). A browser-based tool for visualization and analysis of diffusion MRI data. Nature communications, 9(1), 940.
- We need to easily explore data
- We need to easily share data
- tabular data


Web Technology
Domain Science
pip install AFQ-Browser
afqbrowser-assemble /path/to/afq.mat
afqbrowser-publish /path/to/target/ reponame
Easy to Share
Conclusion
Big Data has bottlenecks
they delay us from developing precision medicine
they delay us from scientific discovery
We need to bridge the domain divide

The web browser can get us there

If our Big Data science is accessible
People will help

If our Big Data science is intuitive
People will understand

Thanks to my Web Lab!
Ariel Rokem
Jason Yeatman
Satra Ghosh
Dylan Nielson
Sharabesh Ramesh
Dave Kennedy
JB Poline
Neel Somani
Sharabesh Ramesh
Katie Bottenhorn
Arno Klein
Amanda Easson
Adam Richie-Halford
Josh Smith
Greg Kiar
Sook-Lei Liew
Chris Madan
Chris Markiewicz
Elizabeth Dupre
Kesshi Jordan
Rob Fatland
Shima Abadi
Naomi Penfold
Andreea Hrincu
Adam Thomas
NVIDIA
AWS
Microsoft Azure
eLife Innovation
UW Institute for Neuroengineering
eScience Institute
All braindr raters
Questions?
From the wet lab to the web lab
By Anisha Keshavan
From the wet lab to the web lab
- 1,507



