Predicting Alzheimer’s Disease Progression
with Deep Generative Models
from MRI Data
presented by:
[Hamid Salehian]
[Prof. Babak Majidi]
Supervisor:
Academic Year
1396 - 1397

Introduction
AlZheimer's Disease: What
- Alzheimer’s is an irreversible, progressive brain disease that slowly destroys memory and disorder cognitive function.
- Clinical studies can indicate disease first symptoms of cognitive impairment while the disease is already in an dvanced stage
- Sixth leading cause of death in the United States.
- One out of 85 persons will have Alzheimers disease (AD) by 2050

AlZheimer's Disease: Why
- Plaques. These clumps of a protein called beta-amyloid may damage and destroy brain cells in several ways, including interfering with inter-cell communication.
-
Tangles. Twisted fibers of a protein called tau. disintegrate the transport system so nutrients and other essential supplies can no longer move through the cells, which eventually die.
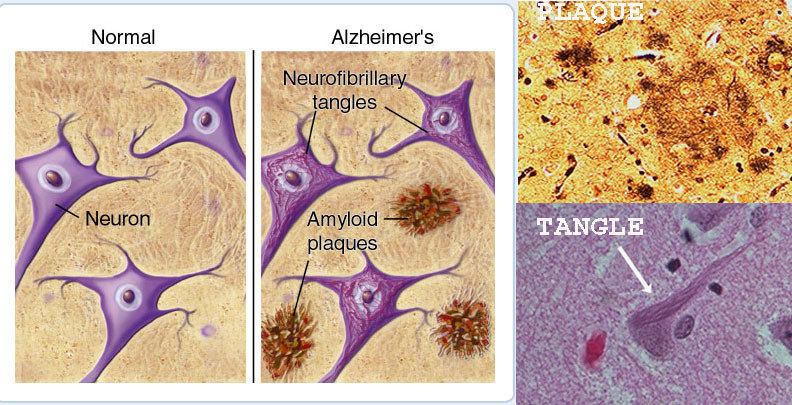
AlZheimer's Disease: Causes
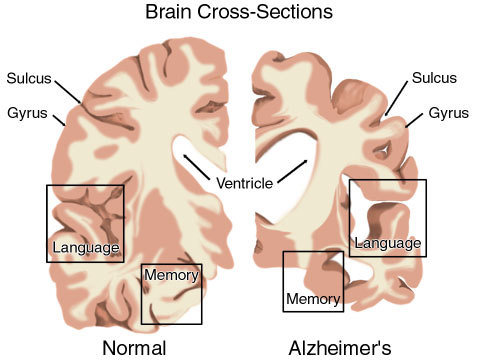
Dying cells cause brain shrinkage
AlZheimer’s Diagnostic
- Review medical history
- Mini Mental Status Exam (MMSE)
- Physical Exam
- Neurological Exam
- Brain Image: Structural(MRI,CT), Functional(fMRI), PET (Positron Emission Tomography)
- Cerebrospinal Fluid (CSF)
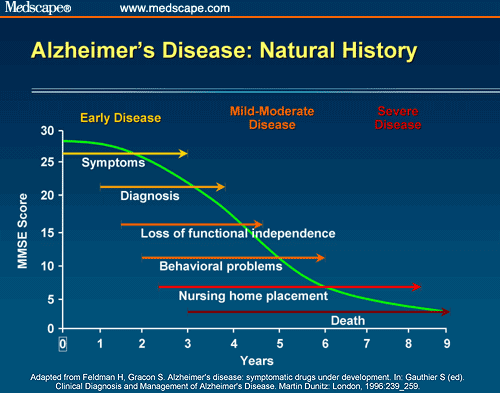
Allows for measurement of the 3-dimensional (3D) volume of brain structures, especially the size of the hippocampus and ventricles enlargement
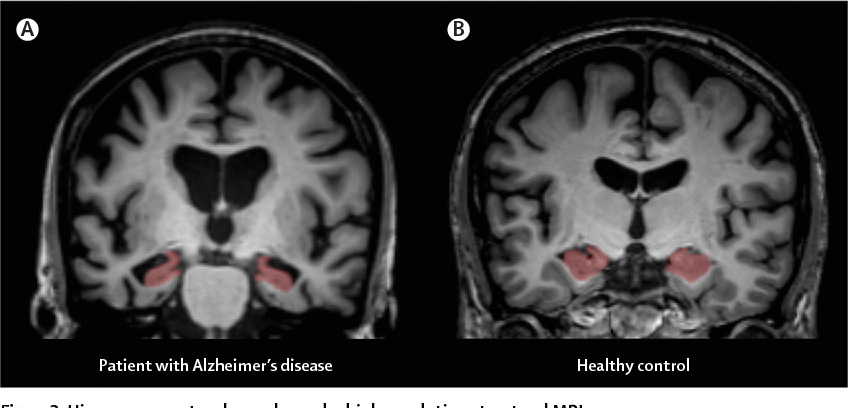
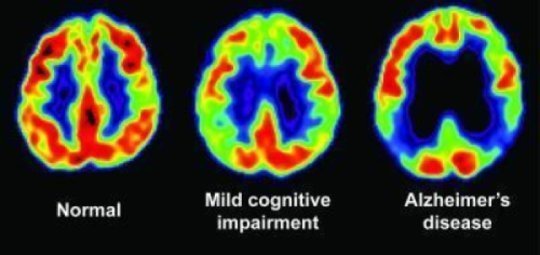
Previous Works
Diagnosis Methods
Automatic methods:
volumetric and shape features together with PCA and SVM were used to
classify MRI images as having the disease or not (Lee, et al., 2009).
Effect of PCA on SVM classfier and k-means Clustering for Alzheimer's Disease Diagnosis
Deep Learning: using 3D Convolutional Neural Networks for classification. (Korelev 2017).
| Reference | Year | Modality | Method | AD/MCI/NC | AD/NC |
|---|---|---|---|---|---|
| Suk et al | 2013 | PET+MRI+CSF | SAE+SVM | N/A | 95.9 |
| Suk et al | 2014 | PET+MRI | SAE+SVM | N/A | 95.4 |
| Zhu et al | 2014 | PET+MRI+CSF | MSLF+SVM | N/A | 95.9 |
| Zu et al | 2015 | PET+MRI | MTFS+SVM | N/A | 96 |
| Liu et al. | 2015 | PET+MRI | SAE+SVM | 53.8 | 91.4 |
| Liu et al. | 2015 | MRI | MFE+SVM | N/A | 93.8 |
| Li et al. | 2015 | PET+MRI+CSF | PCA+SVM | N/A | 91.4 |
| Payan et al. | 2016 | MRI | 2D-SAE | 89.4 | 95.4 |
| Hossein-Asl | 2016 | MRI | 3D-CAE | 89.1 | 97.6 |
| Sarraf et al. | 2016 | rs-fMRI | CNN | N/A | 99.9 |
| Sarraf et al. | 2016 | MRI | CNN | N/A | 98.84 |
| Sarraf et al. | 2017 | rs-fMRI | CNN | N/A | 97.77 |
| 0 | 2017 | PET+MRI | CNN+SAE | N/A | 90 |
CAE: Convolutional Autoencoder
SAE: Sparse Autoencoder
CNN: Convolutional Neural Networks
MSLF: Matrix-Similarity based Loss Function
MTFS: Multi-task feature selection
MFE: Multiview Feature Extraction
PCA: Principal Component Analysis
Diagnosis Methods
| Reference | Year | Modality | Method | AD/MCI/NC | AD/NC |
|---|---|---|---|---|---|
| Domínguez et al | 2016 | MRI | PCA+SVM | 83.48 | N/A |
| Khajehnejad | 2017 | PET+MRI | MBSL | 67.5 | N/A |
| Milana | 2018 | PET+MRI | CVAE+CAAE | 93.86 | N/A |
| Lu et al | 2018 | FDG-PET | MDL | 82.51 | N/A |
CVAE: Convolutional Variational Autoencoders
CAAE:Conditional Adversarial Autoencoders
SAE: Sparse Autoencoder
CNN: Convolutional Neural Networks
PCA: Principal Component Analysis
SVM: Support Vector Machine
MDL: Manifold Deep Learning
EarlY Diagnosis Methods
GOALS
Solutions
- Image reconstruction by domain-transform
- Deep Generative Model to improve MRI images
- Predict Disease progression by reconsrtructed Images
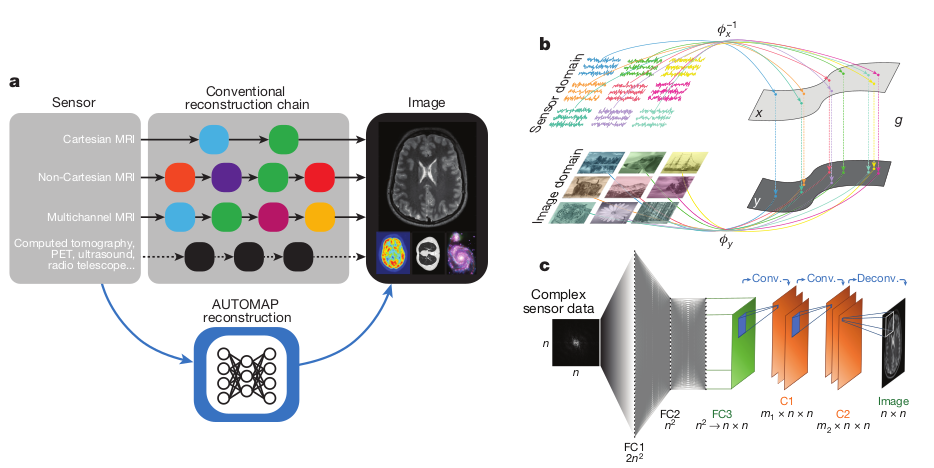
Zhu et al: Image reconstruction by domain-transform
manifold learning (2017) (AUTOMAP)
Khajehnejad et al: Alzheimer’s Disease Early Diagnosis Using
Manifold-Based Semi-Supervised Learning (2017)
Datasets
Online Datasets
HARP: European Alzheimer’s Disease Consortium to gether with the Alzheimer’s Disease Neuroimaging Initiative (ADNI). It was provided as HARmonized Protocol (HARP) for manual hippocampal segmentation from MRI and it consists of 131 volumes.
ADNI: Alzheimer’s Disease Neuroimaging Initiative
OASIS: the Open Access Series of Imaging Studies (OASIS) was created by the Washington University Alzheimer’s Disease Research Center, It consists of 233 volumes.

References
[1] B. Birur, N. V. Kraguljac, R. C. Shelton, and A. C. Lahti. Brain structure, function, and neurochemistry in schizophrenia and bipolardisorder a systematic review of the magnetic resonance neuroimaging literature. NPJ Schizophrenia, 3, 15, 2017.
[2] Y. LeCun, B. Boser, J. S. Denker, D. Henderson, R. E. Howard, W. Hubbard, and L. D. Jackel. Handwritten digit recognition with a back-propagation network.NIPS, 1989.
[3] A. Krizhevsky, I. Sutskever, and G. E. Hinton. Imagenet classificationwith deep convolutional neural networks. Advances in neural information processing systems (NIPS), 1097-1105, 2012.
[4] P. Bojanowski, A. Joulin, D. Lopez-Paz, and A. Szlam. Optimizing theLatent Space of Generative Networks. arXiv:1707.05776v1 [stat.ML], 2017.
[5] A. Radford, L. Metz, and S. Chintala. Unsupervised Representation Learning with Deep Convolutional Generative Adversarial Networks. arXiv:1511.06434v2 [cs.LG], 2015.
[6] T. D. Kulkarni, W.F . Whitney, P. Kohli, and J. B. Tenenbaum. Deep Convolutional Inverse Graphics Network. Advances in Neural Informa- tion Processing Systems 28 (NIPS ), 2015.
[7] A. Achlioptas, O. Diamanti, I. Mitliagkas, and L. Guibas. Representation Learning and Adversarial Generation of 3D Point Clouds. arXiv:1707.02392v1 [cs.CV], 2017.
[8] Z. Zhang, Y. Song, and H. Qi. Age progression/regression by con-ditional adversarial autoencoder. In IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 2017.
References
[9] M. R. Arbabshirani, S. Plis, J. Sui, and V. D. Calhoun. Single subject prediction of brain disorders in neuroimaging: Promises and pitfalls. Neuroimage; 145(Pt B):137-165, 2016.
[10] M. Liu, E. Zhang, D. Adeli-Mosabbeb, and D. Shen. Inherent Structure Based Multi-view Learning with Multi-template Feature Representation for Alzheimer’s Disease Diagnosis. IEEE Trans BiomedicalEngineering; 63(7): 14731482, 2016.
[11] N. Amoroso, M. La Rocca, s: Bruno, T. Maggipinto, A. Monaco, R. Bellotti, and S. Tangaro. Brain structural connectivity atrophy in Alzheimer’s disease.
[12] R. Brosch, T. andTam and Initiative for the Alzheimer’s Disease Neuroimaging. Manifold learning of brain MRIs by deep learning. MICCAI16 (Pt 2):633-40, 2013.
[13] E. Hosseini-Asl, R. Keynton, and A. El-Baz. Alzheimer’s disease diagnostics by adaptation of 3D convolutional network. IEEE International Conference on Image Processing (ICIP), 2016.
[14] L. G. Apostolova, A. E. Green, S. Babakchanian, K. S. Hwang, Y.-Y. Chou, A. W. Toga, and P. M. Thompson. Hippocampal atrophy and ventricular enlargement in normal aging, mild cognitive impairment and Alzheimer’s disease. Alzheimer’s Disease and Associated Disorders, 26(1), 17 - 27, 2012.
[15] A. G. Roy et al. Error Corrective Boosting for Learning Fully Con- volutional Networks with Limited Data. Proceedings Medical Image Computing and Computer-Assisted Intervention - MICCAI 2017 - 20th International Conference, 2017.
[16] 2017 Alzheimer’s Disease Facts and Figures. Alzheimer’s Disease As- sociation, 2017.
[17] L. G. Apostolova. Alzheimer’s Disease. Continuum: Lifelong Learning in Neurology 22. 2 Dementia, 419-434, 2016.
[18] D. P. Kingma and M. Welling. Auto-Encoding Variational Bayes. Pro- ceedings of the 2nd International Conference on Learning Representations (ICLR), 2014.
Question!?
Thank You For Your Attention
powered by: slides.com
see more: https://slides.com/hsarena
Predicting Alzheimer’s Disease Progression with Deep Generative Models from MRI Data
By Hamid Salehian
Predicting Alzheimer’s Disease Progression with Deep Generative Models from MRI Data
Predicting Alzheimer’s Disease Progression with Deep Generative Models from MRI Data
- 204



