Terrell Russell, Ph.D.
@terrellrussell
Executive Director, iRODS Consortium

March 30, 2022
ABRF 2022
Palm Springs, CA
Open Source Storage Abstraction, Image Management, and Data Curation with iRODS and OMERO
Open Source Storage Abstraction, Image Management, and Data Curation with iRODS and OMERO




Our Membership

















Consortium
Member
Consortium
Member
Consortium
Member






Consortium
Member


Today's Talk

- An overview of the iRODS platform, its capabilities and existing integrations
- A recent use case of implementing a read-only local analysis staging policy for the UNC Neuroscience Microscopy Core (NMC) at the UNC-Chapel Hill School of Medicine
- The ongoing efforts to integrate with and provide storage abstraction for OMERO, the Open Microscopy Environment's open source image management software
Why use iRODS?

People need a solution for:
- Managing large amounts of data across various storage technologies
- Controlling access to data
- Searching their data quickly and efficiently
- Automation
The larger the organization, the more they need software like iRODS.
iRODS as the Integration Layer
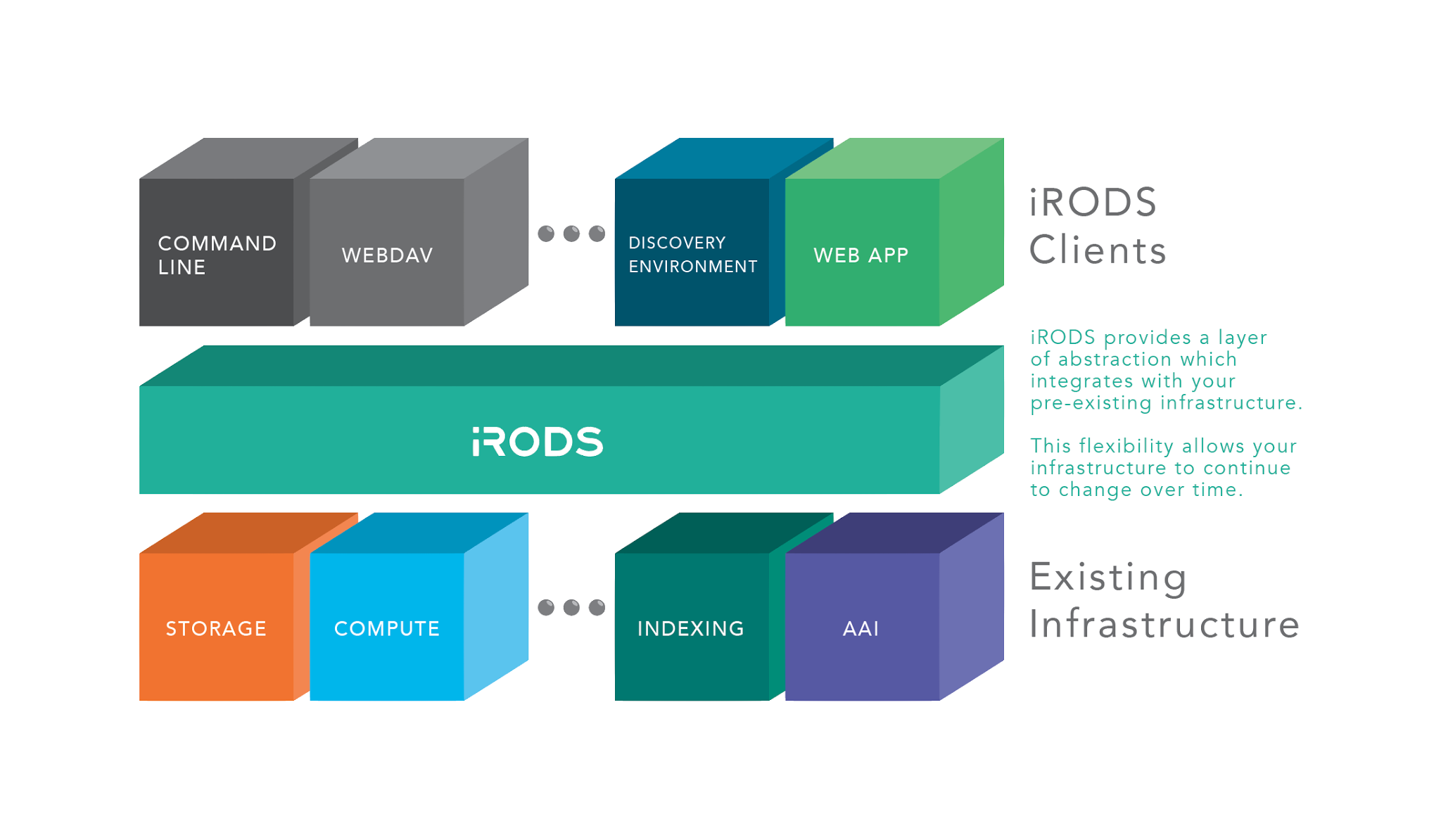

Philosophical Drivers
-
100-year view
-
Plugin Architecture
-
core is generic - protocol, api, bookkeeping
-
plugins are specific
-
policy composition
-
-
Modern core libraries
-
standardized interfaces
-
refactored iRODS internals
-
ease of (re)use
-
fewer bugs
-
-
- Configuration, Not Code

iRODS Core Competencies





- Packaged and supported solutions
- Require configuration not code
- Derived from the majority of use cases observed in the user community








iRODS Capabilities


The Data Management Model



Today's Talk

- An overview of the iRODS platform, its capabilities and existing integrations
- A recent use case of implementing a read-only local analysis staging policy for the UNC Neuroscience Microscopy Core (NMC) at the UNC-Chapel Hill School of Medicine
- The ongoing efforts to integrate with and provide storage abstraction for OMERO, the Open Microscopy Environment's open source image management software
BRAIN-I Project - Architecture

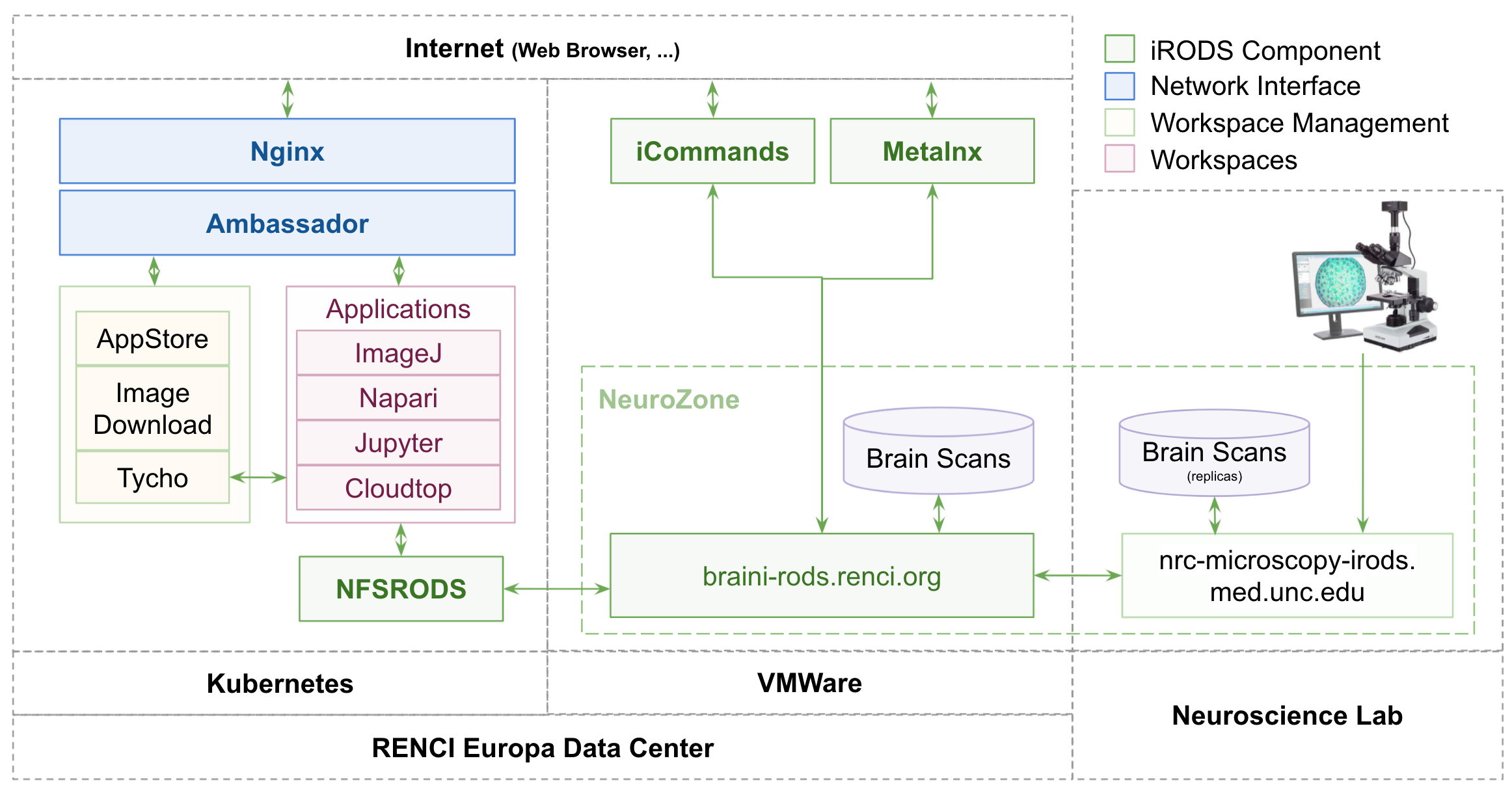
BRAIN-I Project - Design Goals

- Automatic Storage Tiering to primary storage at RENCI
- Manual targeting of files to NMC for local analysis
- Manual targeting of files to future location as published
BRAIN-I Project - Implemented Policy

As part of the BRAIN-I project, this iRODS policy set defines the policies for data analysis, replication, and retention in the NMC.
There are two parts of the policy managing the data flow within the iRODS Zone:
-
Automatic
The iRODS Storage Tiering Framework is handling newly ingested data and moving it into the long-term storage housed at RENCI. RENCI is providing storage and visualization tooling that prioritizes that local, long-term storage. -
Manual
When NMC staff want to run local analysis on data already in the iRODS namespace, they can 'tag' the data of interest, and this policy will manage the replication to their local machine, set permissions, and prevent removal of that data from the system until it has been 'untagged'. Once 'untagged', the data will be trimmed from the researchers' local storage and remain housed only in long-term storage at RENCI.

$ git clone https://github.com/bats-core/bats-core $ time bash bats-core/bin/bats test_nmc_analysis.bats ✓ tag a collection ✓ tag a data object ✓ untag a collection ✓ untag a data object ✓ overwrite a tagged data object ✓ overwrite a data object under a tagged collection ✓ trim a tagged data object - DISALLOWED ✓ trim a data object under a tagged collection - DISALLOWED ✓ remove a tagged data object - DISALLOWED ✓ remove a tagged collection - DISALLOWED ✓ remove a data object under a tagged collection - DISALLOWED ✓ remove a collection under a tagged collection - DISALLOWED ✓ remove a collection containing a tagged data object - DISALLOWED ✓ remove a collection containing a tagged collection - DISALLOWED ✓ untag an enqueued data object - DISALLOWED ✓ untag a collection with an enqueued descendent data object - DISALLOWED 16 tests, 0 failures real 2m4.745s user 0m8.606s sys 0m2.172s
BRAIN-I Project - Testing
Today's Talk

- An overview of the iRODS platform, its capabilities and existing integrations
- A recent use case of implementing a read-only local analysis staging policy for the UNC Neuroscience Microscopy Core (NMC) at the UNC-Chapel Hill School of Medicine
- The ongoing efforts to integrate with and provide storage abstraction for OMERO, the Open Microscopy Environment's open source image management software
Working Groups

Imaging Working Group
- Goal: To provide a standardized suite of imaging policies and practices for integration with existing tools and pipelines
- Open Microscopy Environment (and OMERO)
- Neuroscience Microscopy Core at UNC School of Medicine
- New York University
- Santa Clara University
- UC San Diego
- UC Santa Cruz
- UMass
- Harvard
- Maastricht University
- Wellcome Sanger Institute
- CyVerse
- NIEHS
- Netherlands Cancer Institute (NKI)
- Francis Crick Institute
- Fritz Lipmann Institute
- Osnabrück University
- RIKEN
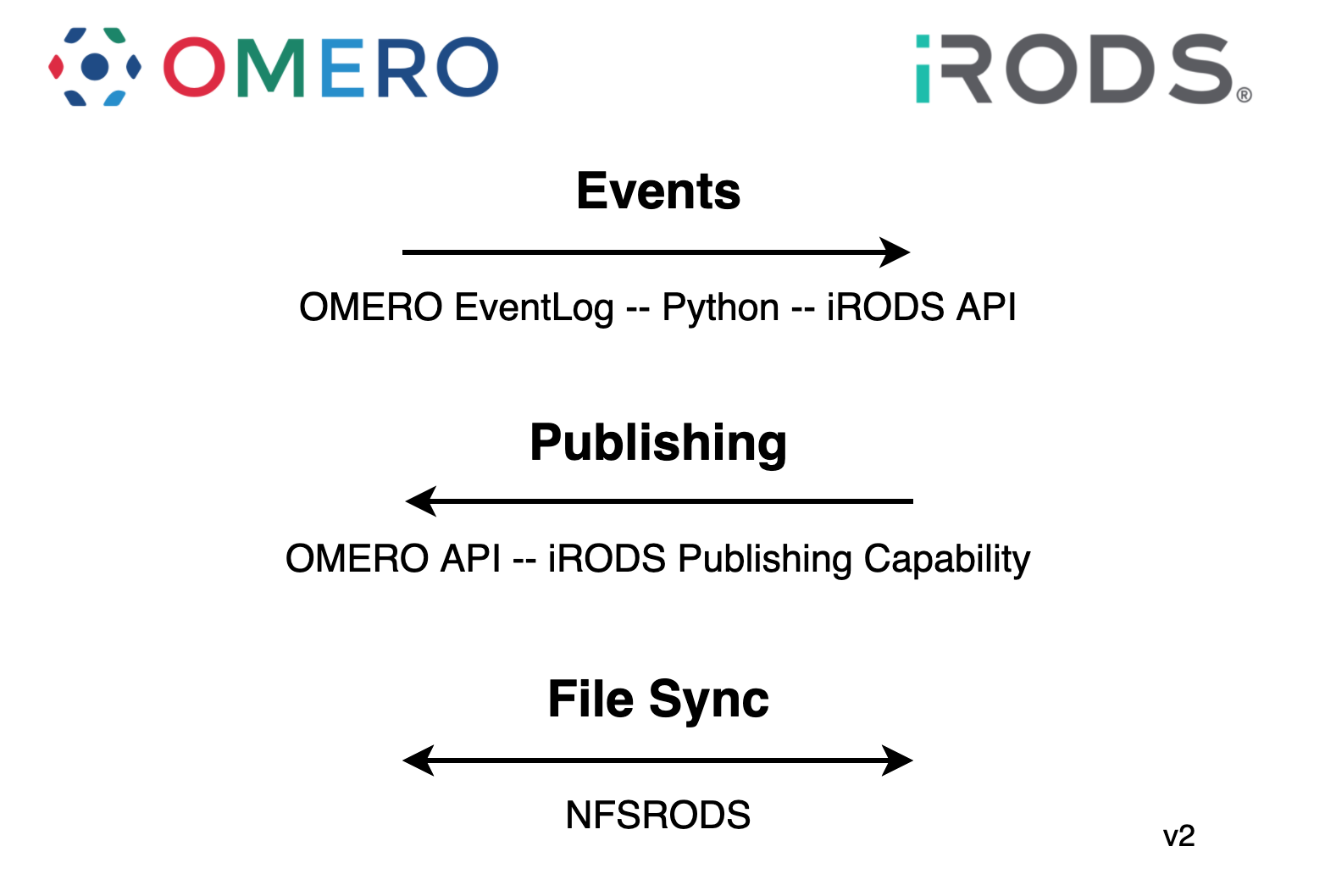
OMERO <-> iRODS

- The current plan is to design a sync agent to serve as the arbiter of truth
- New edits / annotations / events in OMERO will trigger equivalent and necessary updates in iRODS
- New edits / annotations / events in iRODS will trigger equivalent and necessary updates in OMERO
Questions?

Proper data management requires policy enforcement.
These policies will change over time.
Open source is the best practice for a 100-year view.
Terrell Russell
@terrellrussell
iRODS Consortium
Thank you.
irods.org
ABRF 2022 - Data Curation with iRODS and OMERO
By iRODS Consortium
ABRF 2022 - Data Curation with iRODS and OMERO
- 1,011

