Johannes Köster
2019
https://koesterlab.github.io

Reproducible data analysis with
Agenda
- Introduction to Conda
- Packages & channels
- Environments
- Writing Recipes
- Introduction to Snakemake
- Workflow definition
- Workflow execution
- Live demo
dataset
results
Data analysis
"Let me do that by hand..."

dataset
results
dataset
dataset
dataset
dataset
dataset


"Let me do that by hand..."
Data analysis
dataset
results
dataset
dataset
dataset
dataset
dataset
automation
From raw data to final figures:
- document parameters, tools, versions
- execute without manual intervention
Reproducible data analysis
dataset
results
dataset
dataset
dataset
dataset
dataset
scalability
Handle parallelization:
- execute for tens to thousands of datasets
- efficiently use any computing platform
automation
Reproducible data analysis
dataset
results
dataset
dataset
dataset
dataset
dataset
Handle deployment:
be able to easily execute analyses on a different system/platform/infrastructure
portability
scalability
automation
Reproducible data analysis
150k downloads since 2015
















Snakemake is a popular solution

dataset
results
dataset
dataset
dataset
dataset
dataset
scalability
automation
portability
dataset
results
dataset
dataset
dataset
dataset
dataset
Define workflows
in terms of rules
Define workflows
in terms of rules
rule mytask:
input:
"path/to/{dataset}.txt"
output:
"result/{dataset}.txt"
script:
"scripts/myscript.R"
rule myfiltration:
input:
"result/{dataset}.txt"
output:
"result/{dataset}.filtered.txt"
shell:
"mycommand {input} > {output}"
rule aggregate:
input:
"results/dataset1.filtered.txt",
"results/dataset2.filtered.txt"
output:
"plots/myplot.pdf"
script:
"scripts/myplot.R"Define workflows
in terms of rules
Define workflows
in terms of rules
rule mytask:
input:
"data/{sample}.txt"
output:
"result/{sample}.txt"
shell:
"some-tool {input} > {output}"rule name
how to create output from input
define
- input
- output
- log files
- parameters
- resources
rule mytask:
input:
"path/to/{dataset}.txt"
output:
"result/{dataset}.txt"
script:
"scripts/myscript.R"
rule myfiltration:
input:
"result/{dataset}.txt"
output:
"result/{dataset}.filtered.txt"
shell:
"mycommand {input} > {output}"
rule aggregate:
input:
"results/dataset1.filtered.txt",
"results/dataset2.filtered.txt"
output:
"plots/myplot.pdf"
script:
"scripts/myplot.R"Define workflows
in terms of rules

Define workflows
in terms of rules
rule mytask:
input:
"data/{sample}.txt"
output:
"result/{sample}.txt"
script:
"scripts/myscript.py"reusable
Python/R scripts
External scripts
import pandas as pd
data = pd.read_table(snakemake.input[0])
data = data.sort_values("id")
data.to_csv(snakemake.output[0], sep="\t")Python scripts:
External scripts
data <- read.table(snakemake@input[[1]])
data <- data[order(data$id),]
write.table(data, file = snakemake@output[[1]])R scripts:
Define workflows
in terms of rules
rule map_reads:
input:
"{sample}.bam"
output:
"{sample}.sorted.bam"
wrapper:
"0.22.0/bio/samtools/sort"reuseable wrappers from central repository
Define workflows
in terms of rules
use CWL tool
definitions
rule map_reads:
input:
"{sample}.bam"
output:
"{sample}.sorted.bam"
cwl:
"https://github.com/common-workflow-language/"
"workflows/blob/fb406c95/tools/samtools-sort.cwl"Output handling
rule mytask:
input:
"data/{sample}.txt"
output:
temp("result/{sample}.txt")
shell:
"some-tool {input} > {output}"Output handling
rule mytask:
input:
"data/{sample}.txt"
output:
protected("result/{sample}.txt")
shell:
"some-tool {input} > {output}"Output handling
rule mytask:
input:
"data/{sample}.txt"
output:
pipe("result/{sample}.txt")
shell:
"some-tool {input} > {output}"dataset
results
dataset
dataset
dataset
dataset
dataset
scalability
automation
portability
Scheduling
Paradigm:
Workflow definition shall be independent of computing platform and available resources
Rules:
define resource usage (threads, memory, ...)
Scheduler:
- solves multidimensional knapsack problem
- schedules independent jobs in parallel
- passes resource requirements to any backend
Scalable to any platform



workstation
compute server
cluster






grid computing
cloud computing



Command-line interface
# perfom dry-run
snakemake -n
# execute workflow locally with 16 CPU cores
snakemake --cores 16
# execute on cluster
snakemake --cluster qsub --jobs 100
# execute in the cloud
snakemake --kubernetes --jobs 1000 --default-remote-provider GS --default-remote-prefix mybucketConfiguration profiles
snakemake --profile slurm --jobs 1000$HOME/.config/snakemake/slurm
├── config.yaml
├── slurm-jobscript.sh
├── slurm-status.py
└── slurm-submit.pydataset
results
dataset
dataset
dataset
dataset
dataset
Full reproducibility:
install required software and all dependencies in exact versions
portability
scalability
automation
Software installation is a pain
source("https://bioconductor.org/biocLite.R")
biocLite("DESeq2")easy_install snakemake./configure --prefix=/usr/local
make
make installcp lib/amd64/jli/*.so lib
cp lib/amd64/*.so lib
cp * $PREFIXcpan -i bioperlcmake ../../my_project \
-DCMAKE_MODULE_PATH=~/devel/seqan/util/cmake \
-DSEQAN_INCLUDE_PATH=~/devel/seqan/include
make
make installapt-get install bwayum install python-h5pyinstall.packages("matrixpls")rule mytask:
input:
"path/to/{dataset}.txt"
output:
"result/{dataset}.txt"
conda:
"envs/some-tool.yaml"
shell:
"some-tool {input} > {output}"Conda integration
channels:
- conda-forge
dependencies:
- some-tool =2.3.1
- some-lib =1.1.2Singularity integration
rule mytask:
input:
"path/to/{dataset}.txt"
output:
"result/{dataset}.txt"
singularity:
"docker://biocontainers/some-tool#2.3.1"
shell:
"some-tool {input} > {output}"Singularity + Conda
singularity:
"docker://continuumio/miniconda3:4.4.1"
rule mytask:
input:
"path/to/{dataset}.txt"
output:
"result/{dataset}.txt"
conda:
"envs/some-tool.yaml"
shell:
"some-tool {input} > {output}"define OS
define tools/libs
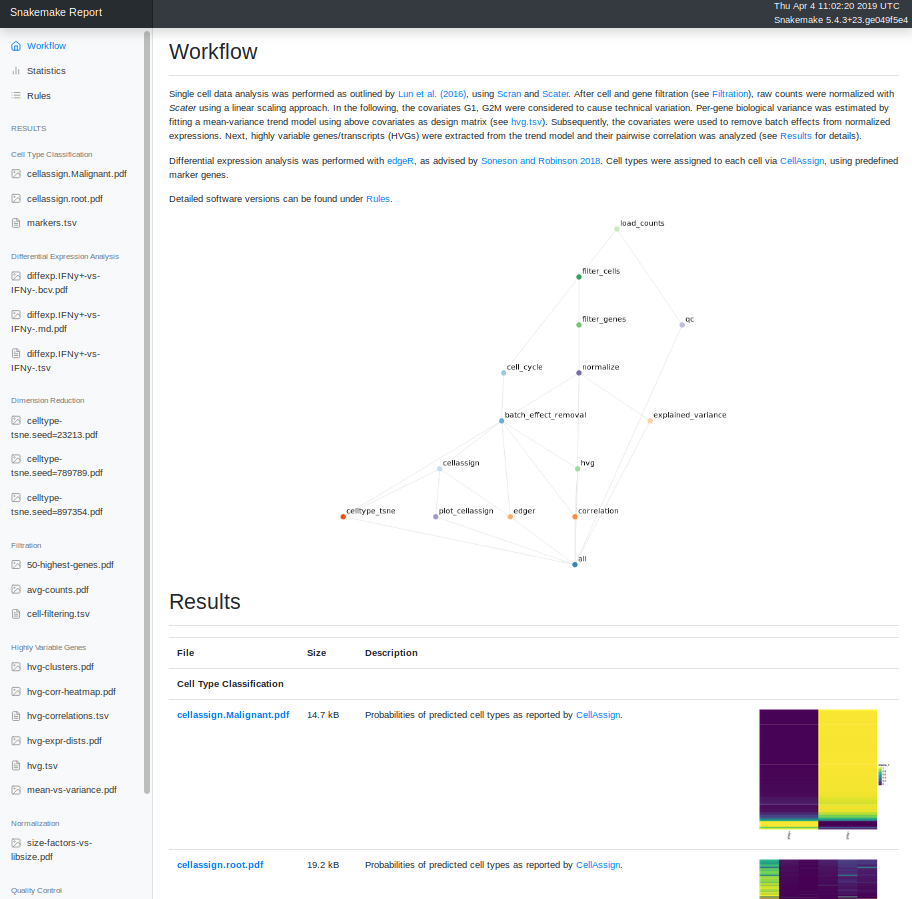
Self-contained HTML reports
Sustainable publishing
# archive workflow (including Conda packages)
snakemake --archive myworkflow.tar.gzAuthor:
- Upload to Zenodo and acquire DOI.
- Cite DOI in paper.
Reader:
- Download and unpack workflow archive from DOI.
# execute workflow (Conda packages are deployed automatically)
snakemake --use-conda --cores 16More features
Today:
- conditional DAG updates based on job output
- semi-automatic graph partitioning
- resource-constrained scheduling
- various ways to constrain or enforce job execution
- data provenance and log file handling
- CWL export
- ...
Future:
- jupyter notebook integration
- ML-based inference of resource requirements
- more backends (TES, GCP, AWS Batch)
Conclusion
With
- the human readable specification language
- reusable modularization capabilities
- seamless execution on all platforms without adaptation of the workflow definition
- integrated package management and containerization
Snakemake covers all three dimensions of fully reproducible data analysis.
portability
scalability
automation
Acknowledgements
Contributors:
Andreas Wilm
Anthony Underwood
Ryan Dale
David Alexander
Elias Kuthe
Elmar Pruesse
Hyeshik Chang
Jay Hesselberth
Jesper Foldager
John Huddleston
all users and supporters
Joona Lehtomäki
Justin Fear
Karel Brinda
Karl Gutwin
Kemal Eren
Kostis Anagnostopoulos
Kyle A. Beauchamp
Simon Ye
Tobias Marschall
Willem Ligtenberg
Development team:
Christopher Tomkins-Tinch
David Koppstein
Tim Booth
Manuel Holtgrewe
Christian Arnold
Wibowo Arindrarto
Rasmus Ågren
Kyle Meyer
Lance Parsons
Manuel Holtgrewe
Marcel Martin
Matthew Shirley
Mattias Franberg
Matt Shirley
Paul Moore
percyfal
Per Unneberg
Ryan C. Thompson
Ryan Dale
Sean Davis
Resources
Documentation and change log:
https://snakemake.readthedocs.io
Questions:
http://stackoverflow.com/questions/tagged/snakemake
Gold standard workflows:
https://github.com/snakemake-workflows/docs
Configuration profiles:
https://github.com/snakemake-profiles/doc
Command line help:
snakemake --help
Let us know what you think :-)
ISMB-Snakemake-Tutorial
By Johannes Köster
ISMB-Snakemake-Tutorial
Data analyses usually entail the application of many command line tools or scripts to transform, filter, aggregate or plot data and results. With ever increasing amounts of data being collected in science, reproducible and scalable automatic workflow management becomes increasingly important. Snakemake is a workflow management system, consisting of a text-based workflow specification language and a scalable execution environment, that allows the parallelized execution of workflows on workstations, compute servers, clusters and the cloud without modification of the workflow definition. Since its publication, Snakemake has been widely adopted and was used to build analysis workflows for a variety of high impact publications. With about thousands of homepage visits per month, it has a large and stable user community. This talk will show how Snakemake can be used to easily document, execute, and reproduce data analyses.
- 3,065