Towards a unified theory of variant calling
Johannes Köster, Louis Dijkstra, Tobias Marschall, Alexander Schönhuth
Genome Informatics 2020
small
structural
SNVs, Indels
Inversions, Duplications, Indels, ...
?
?
?
?
somatic
germline
Variant calling
small
structural
?
?
somatic
germline
Variant calling
tumor/normal
sample, cohort, pedigree
?
?
small
structural
germline
somatic
GATK
Freebayes
Platypus
Pindel
Mutect
Strelka
Octopus
Lancet
Variant calling
Delly
Manta
Gridss
##fileformat=VCFv4.3
##fileDate=20090805
##source=myImputationProgramV3.1
##reference=file:///seq/references/1000GenomesPilot-NCBI36.fasta
##contig=<ID=20,length=62435964,assembly=B36,md5=f126cdf8a6e0c7f379d618ff66beb2da,species="Homo sapiens",taxonomy=x>
##phasing=partial
##INFO=<ID=NS,Number=1,Type=Integer,Description="Number of Samples With Data">
##INFO=<ID=DP,Number=1,Type=Integer,Description="Total Depth">
##INFO=<ID=AF,Number=A,Type=Float,Description="Allele Frequency">
##INFO=<ID=AA,Number=1,Type=String,Description="Ancestral Allele">
##INFO=<ID=DB,Number=0,Type=Flag,Description="dbSNP membership, build 129">
##INFO=<ID=H2,Number=0,Type=Flag,Description="HapMap2 membership">
##FILTER=<ID=q10,Description="Quality below 10">
##FILTER=<ID=s50,Description="Less than 50% of samples have data">
##FORMAT=<ID=GT,Number=1,Type=String,Description="Genotype">
##FORMAT=<ID=GQ,Number=1,Type=Integer,Description="Genotype Quality">
##FORMAT=<ID=DP,Number=1,Type=Integer,Description="Read Depth">
##FORMAT=<ID=HQ,Number=2,Type=Integer,Description="Haplotype Quality">
#CHROM POS ID REF ALT QUAL FILTER INFO FORMAT NA00001
20 14370 rs6054257 G A 29 PASS NS=3;DP=14;AF=0.5;DB;H2 GT:GQ:DP:HQ 0|0:48:1:51,51
20 17330 . T A 3 q10 NS=3;DP=11;AF=0.017 GT:GQ:DP:HQ 0|0:49:3:58,50
20 1110696 rs6040355 A G,T 67 PASS NS=2;DP=10;AF=0.333,0.667;AA=T;DB GT:GQ:DP:HQ 1|2:21:6:23,27
20 1230237 . T . 47 PASS NS=3;DP=13;AA=T GT:GQ:DP:HQ 0|0:54:7:56,60
20 1234567 microsat1 GTC G,GTCT 50 PASS NS=3;DP=9;AA=G GT:GQ:DP 0/1:35:4 ##fileformat=VCFv4.3
##fileDate=20090805
##source=myImputationProgramV3.1
##reference=file:///seq/references/1000GenomesPilot-NCBI36.fasta
##contig=<ID=20,length=62435964,assembly=B36,md5=f126cdf8a6e0c7f379d618ff66beb2da,species="Homo sapiens",taxonomy=x>
##phasing=partial
##INFO=<ID=NS,Number=1,Type=Integer,Description="Number of Samples With Data">
##INFO=<ID=DP,Number=1,Type=Integer,Description="Total Depth">
##INFO=<ID=AF,Number=A,Type=Float,Description="Allele Frequency">
##INFO=<ID=AA,Number=1,Type=String,Description="Ancestral Allele">
##INFO=<ID=DB,Number=0,Type=Flag,Description="dbSNP membership, build 129">
##INFO=<ID=H2,Number=0,Type=Flag,Description="HapMap2 membership">
##FILTER=<ID=q10,Description="Quality below 10">
##FILTER=<ID=s50,Description="Less than 50% of samples have data">
##FORMAT=<ID=GT,Number=1,Type=String,Description="Genotype">
##FORMAT=<ID=GQ,Number=1,Type=Integer,Description="Genotype Quality">
##FORMAT=<ID=DP,Number=1,Type=Integer,Description="Read Depth">
##FORMAT=<ID=HQ,Number=2,Type=Integer,Description="Haplotype Quality">
#CHROM POS ID REF ALT QUAL FILTER INFO FORMAT NA00002 NA00003
20 14370 rs6054257 G A 29 PASS NS=3;DP=14;AF=0.5;DB;H2 GT:GQ:DP:HQ 1|0:48:8:51,51 1/1:43:5:.,.
20 17330 . T A 3 q10 NS=3;DP=11;AF=0.017 GT:GQ:DP:HQ 0|1:3:5:65,3 0/0:41:3
20 1110696 rs6040355 A G,T 67 PASS NS=2;DP=10;AF=0.333,0.667;AA=T;DB GT:GQ:DP:HQ 2|1:2:0:18,2 2/2:35:4
20 1230237 . T . 47 PASS NS=3;DP=13;AA=T GT:GQ:DP:HQ 0|0:48:4:51,51 0/0:61:2
20 1234567 microsat1 GTC G,GTCT 50 PASS NS=3;DP=9;AA=G GT:GQ:DP 0/2:17:2 1/1:40:3⊖
Problem:
accumulation of errors
More complex: so far...
small
structural
germline
somatic
Delly
GATK
Freebayes
Platypus
Pindel
Mutect
Strelka
Lancet
Manta
Octopus
arbitrary
Gridss
Approach
Given:
- aligned NGS reads
- candidate variants
Find:
- classification of variants into events (somatic, germline, ...)
- unbiased allele frequency estimates
- while controlling FDR

Model

sample a
sample b
...
contaminates
Model

- insert size
- read sequence
- strand
- read orientation
Latent variables:
- allele frequency
- fragment from variant allele
- fragment correctly mapped
- biases
Observed variables:
Model
\(\xi_i \sim \text{Bernoulli}(\alpha \theta_c \tau + (1-\alpha) \theta_h \tau)\)
\(\omega_i \sim Bernoulli(\pi_i)\)
\(Z_i \mid \xi_i, \omega_i=1,\beta,\delta \sim\)


\(\beta, \delta\)
- allele frequency
- contamination
- sampling bias
- typing uncertainty
- strand bias, read orientation bias
- mapping uncertainty

Model
\(\Pr(Z_i \mid \theta_c,\theta_h,\beta) =\)
⊕ Bayes' theorem
= posterior probability for events

Bayesian FDR control
Variant calling grammar
samples:
janedoe:
universe: "0.0 | 0.5 | 1.0"
events:
present: "janedoe:0.5 | janedoe:1.0"samples:
normal:
resolution: 5
universe: "0.0 | 0.5 | 1.0 | ]0.0,0.5["
tumor:
resolution: 100
universe: "[0.0,1.0]"
contamination:
by: normal
fraction: 0.25
events:
germline: "normal:0.5 | normal:1.0"
somatic_normal: "normal:]0.0,0.5["
somatic_tumor: "normal:0.0 & tumor:]0.0,1.0]"Variant calling grammar
samples:
normal:
resolution: 5
universe: "0.0 | 0.5 | 1.0 | ]0.0,0.5["
tumor:
resolution: 100
universe: "[0.0,1.0]"
contamination:
by: normal
fraction: 0.25
relapse:
resolution: 100
universe: "[0.0,1.0]"
contamination:
by: normal
fraction: 0.53
events:
germline: "normal:0.5 | normal:1.0"
somatic_normal: "normal:]0.0,0.5["
somatic_tumor: "normal:0.0 & tumor:]0.0,1.0]"
somatic_relapse: "normal:0.0 & tumor:0.0 & relapse:]0.0,1.0]"Variant calling grammar
samples:
romeo:
universe:
all: "0.0 | 0.5 | 1.0"
X: "0.0 | 1.0"
Y: "0.0 | 1.0"
juliet:
universe:
all: "0.0 | 0.5 | 1.0"
Y: "0.0"
events:
both_het: "juliet:0.5 & romeo:0.5"
both_hom: "juliet:1.0 & romeo:1.0"
juliet_only: "(juliet:0.5 | juliet:1.0) & romeo:0.0"
romeo_only: "(romeo:0.5 | romeo:1.0) & juliet:0.0"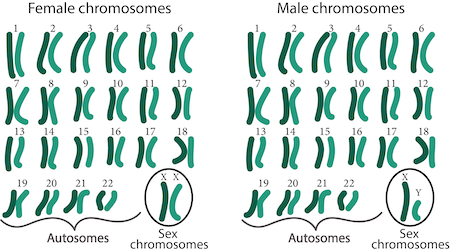
https://genetics.thetech.org
Variant calling grammar
White box model
chr1 1200 . GAAGCCATCTGCCCCATGGGAAAGTGACTTAAT G . .
SVLEN=-32;PROB_SOMATIC_TUMOR=0.110292;PROB_ABSENT=202.181;PROB_GERMLINE_HET=61.7704;
PROB_SOMATIC_NORMAL=16.0075;PROB_GERMLINE_HOM=4346.23;PROB_ARTIFACT=103.395
DP:AF:OBS:SB:ROB
36:0:11Np+<10Np+>8Np-<7Np->:.:.
48:0.102431:14Np+<13Np-<11Np->8Np+>1Vp-<1Vp+>:.:.- posterior allele frequency estimate
- observations (count, posterior odds, type, strand, read orientation)
- strand bias estimate
- read orientation estimate
- event probabilities (PHRED scaled)





https://varlociraptor.github.io
Towards a unified theory of variant calling
By Johannes Köster
Towards a unified theory of variant calling
Talk at Genome Informatics 2020
- 1,712



