chewBBACA's
Nomenclature Server
Lab Meeting 31/10/2019
Pedro Cerqueira

Summary
1. Gene-by-Gene based methods
2. chewBBACA
3. Data Privacy
4. Nomenclature Server
5. Nomenclature Server - Roadmap
6. Nomenclature Server - At the moment
7. Final remarks
Is my strain the same as theirs?


Patient A


Patient B
1. Gene-by-Gene based methods
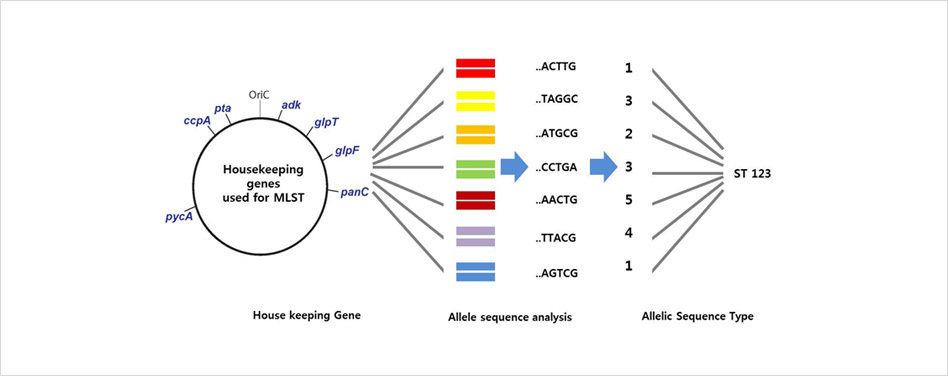
Multilocus Sequence Typing (MLST)
- Defined scheme of typically 7 housekeeping gene fragments
- Robust, portable and unified method for characterizing isolates at a molecular level
- Not enough resolution to perform high resolution typing.
-
Whole-Genome Multilocus Sequence Typing (wgMLST)
- Extend MLST to whole-genome level
- Set of genes that are present across a set of genomes representing a species, akin to a pan-genome.
-
Core-Genome Multilocus Sequence Typing (cgMLST)
- Gene-by-gene allelic profiling of core genome genes in a set of same species isolates
PCR & Sanger Sequecing
whole genome shotgun sequencing
1. Gene-by-Gene based methods
- BLAST Score Ratio (BSR) Based Allele Calling Algorithm
- Open source solution for the creation of whole genome and core genome MultiLocus Sequence Typing (wg/cgMLST) schemas
- Performs allele calls on complete or draft genomes resulting from de novo assemblers

2. chewBBACA
- Requirements

- Installation
git clone https://github.com/B-UMMI/chewBBACA.gitpip install chewbbacaconda install -c bioconda chewbbacaProdigal # for CDS detection
BLAST
Python >= 3.0.0 with numpy>=1.14.0 scipy>=0.13.3 biopython>=1.70 plotly>=1.12.9 SPARQLWrapper>=1.8.0 pandas>=0.22.0
ClustalW2 # optional
mafft # optional
2. chewBBACA
- Profiles
| File | Locus 1 | Locus 2 |
|---|---|---|
| genome1.fasta | Allele 2 | Allele 5 |
| genome2.fasta | Allele 6 | Allele 9 |
- Profile Visualization
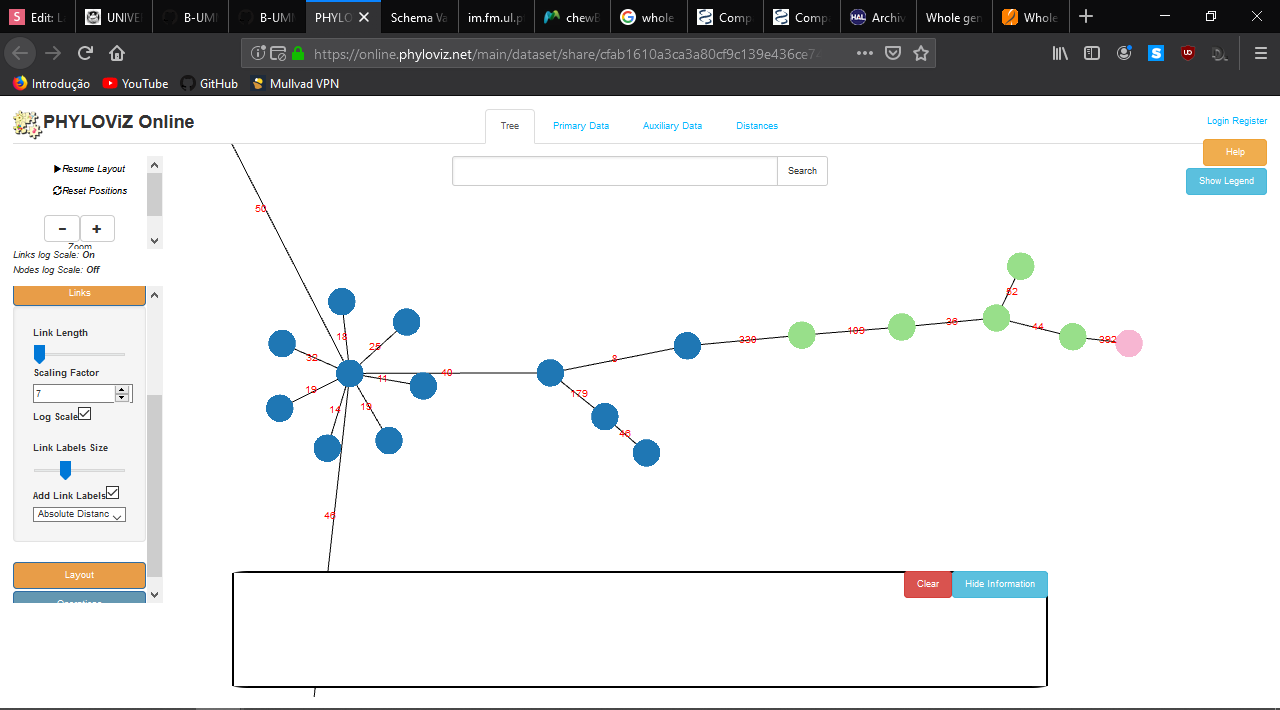
https://online2.phyloviz.net
2. chewBBACA
Output
What do people share?
- Allelic profiles?
- Raw data (reads or assemblies)?
Strict privacy laws may prevent the users from sharing raw data. Unpublished data is also a matter of concern.



- Schemas?
Schemas can be created with different parameters, requiring the users to share their configurations.
Profiles are generated based on the schema, which may contain private data, making it hard to share and obtain the same results.
3. Data Privacy
Is my strain the same as theirs?
Goals:
- Share and compare allele calls within the community at a global level
4. Nomenclature Server

-
Provide a public and centralized web service enabling users to:
- Download the necessary data for the wg/cgMLST schemas
- Query/submit results to the database
- Allow users to perform analyses in their local machines

Server
Client
chewBBACA NS
Allele call results
- Schemas
- Profiles
Query/Submit

4. Nomenclature Server
What data does it store?
- Password hash
- Sequence hash
- Allelic Profiles (also hash)
4. Nomenclature Server
-
Server Front end grooming


5. Nomenclature Server - Roadmap
-
Server API Review and Documentation
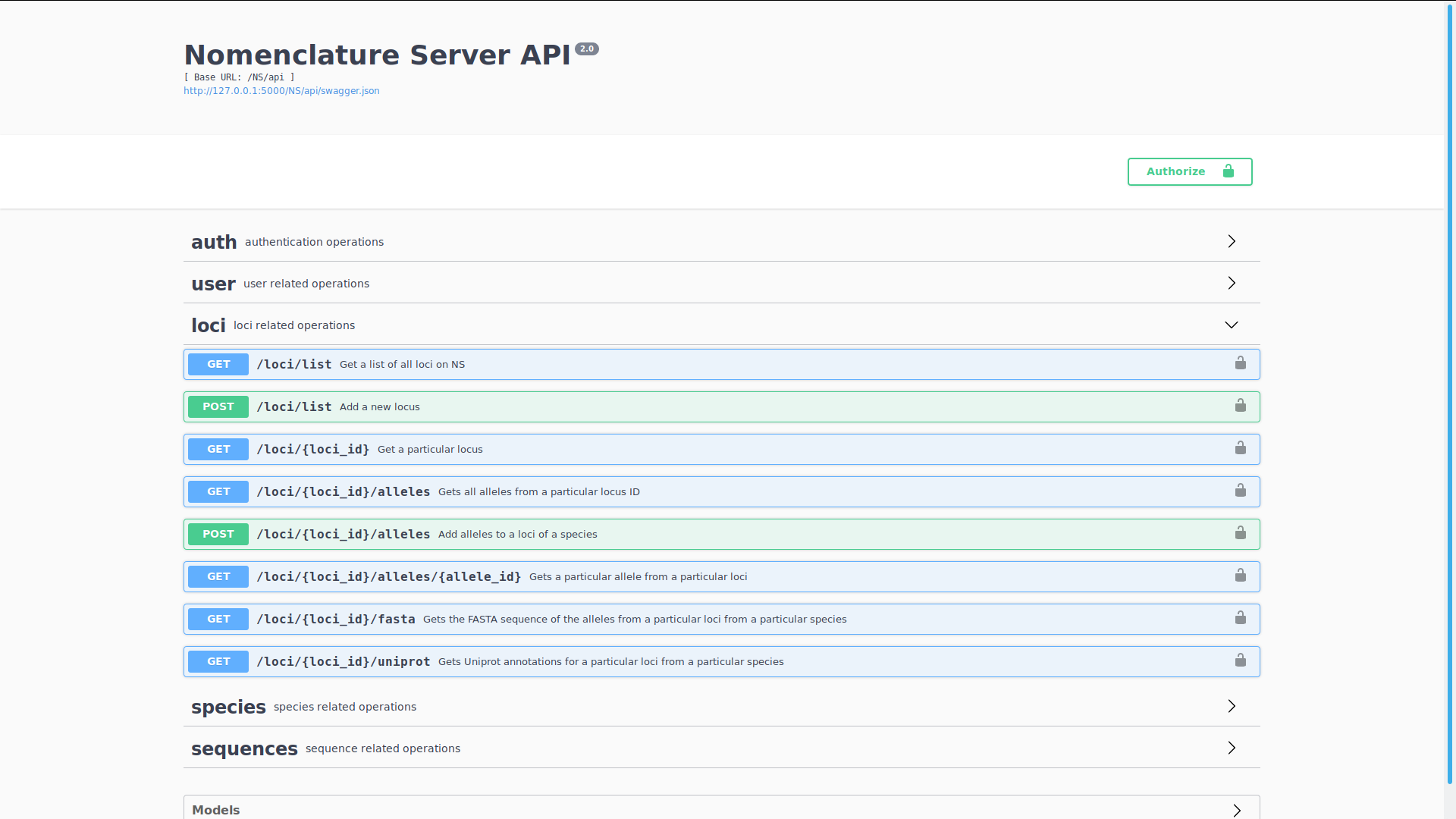
5. Nomenclature Server - Roadmap
-
Client Refactorization
-
Download Schemas
-
Download Profiles
-
Synchronize Schemas
-
Upload Schemas
-
Upload metadata
5. Nomenclature Server - Roadmap

[
{
"schemas": {
"type": "uri",
"value": "http://127.0.0.1:5000/NS/api/species/13/schemas/6"
},
"locus": {
"type": "uri",
"value": "http://127.0.0.1:5000/NS/api/loci/11358"
},
"alleles": {
"type": "uri",
"value": "http://127.0.0.1:5000/NS/api/loci/11358/alleles/3"
},
"uniprot": {
"type": "uri",
"value": "http://purl.uniprot.org/uniprot/Q70EW3"
},
"label": {
"type": "typed-literal",
"datatype": "http://www.w3.org/2001/XMLSchema#string",
"value": "Exotoxin L"
}- Is this sequence in NS?
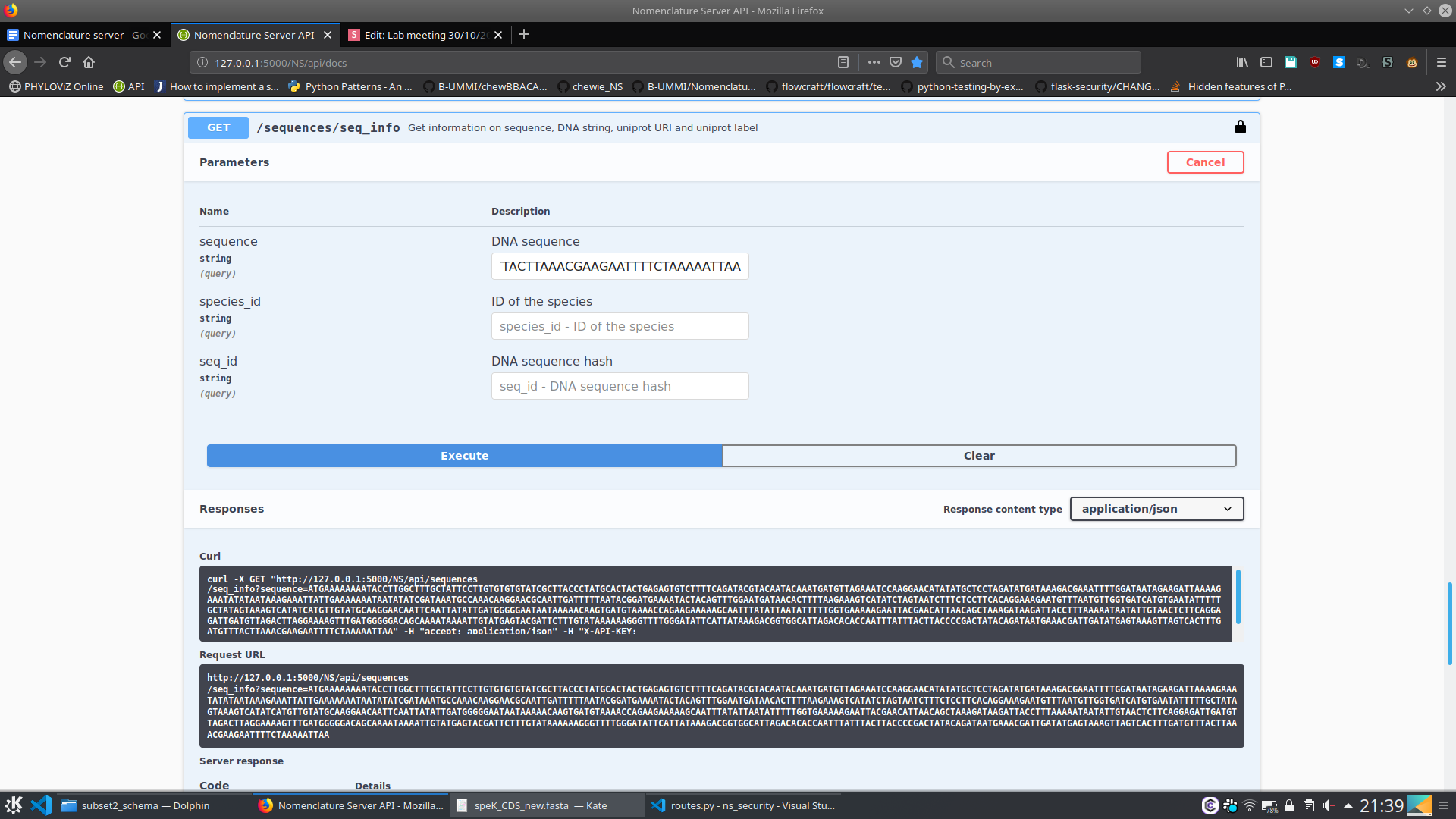
6. Nomenclature Server - At the moment
- What are the fasta sequences of the alleles belonging to this loci?
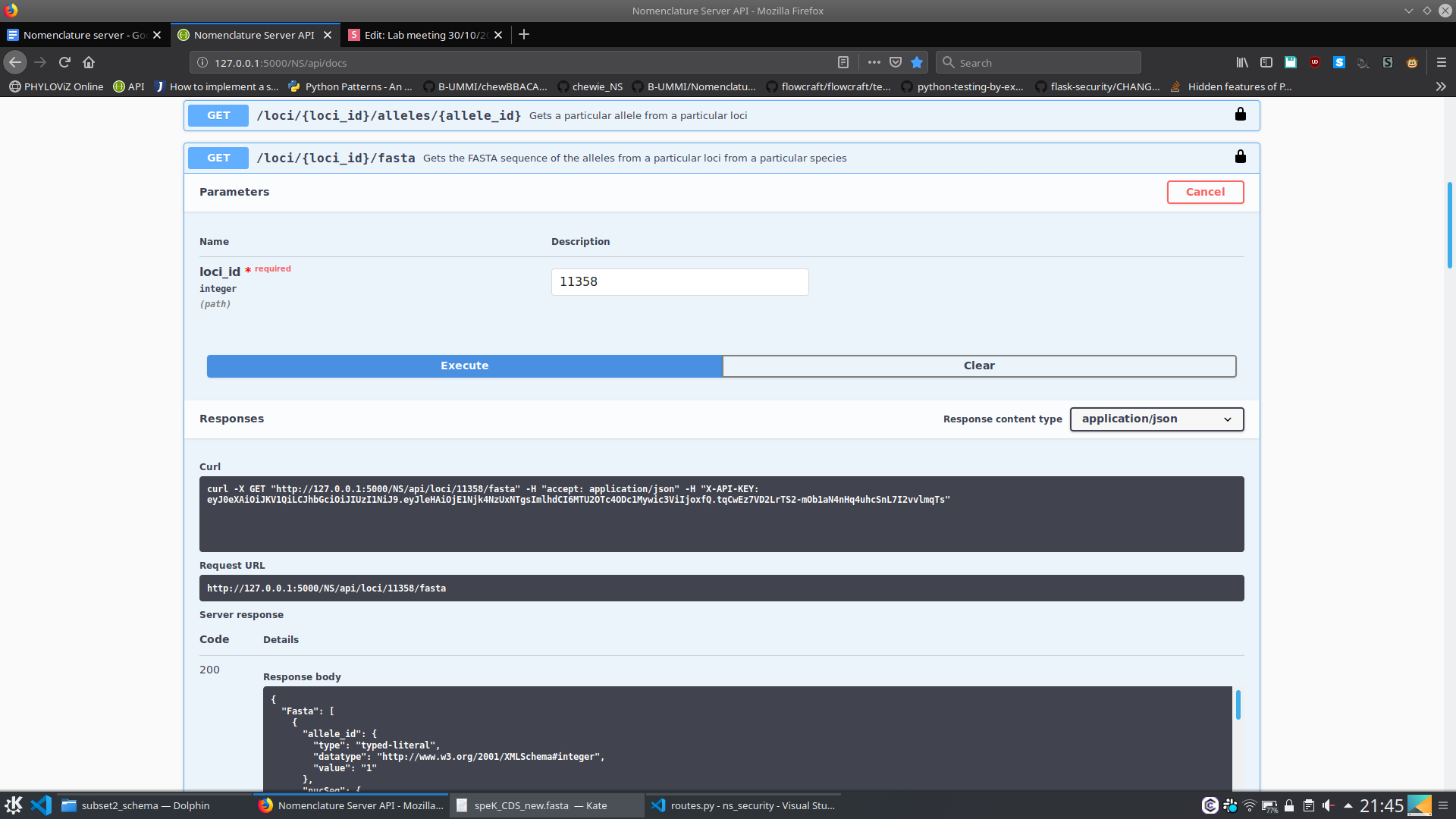
{
"Fasta": [
{
"allele_id": {
"type": "typed-literal",
"datatype": "http://www.w3.org/2001/XMLSchema#integer",
"value": "1"
},
"nucSeq": {
"type": "literal",
"value": "ATGAAAAAAAATACCTTGACTTTGTTATTCCTTGTGTGTGTATCGCTTGCTCTATACACTACTGAGAGTGTCTTTTCAGATACGTACAATACAAATGATGTTAGAAATCCAAGGAACATATATGCTCCTAGATATGATAAAGACGAAATTTTGGATAATAGAAGATTAAAAGAAATATATAATAAAGAAATTATTGAAAAAAATAATATATCGATAAATGCCAAACAAGGAACGCAATTGATTTTTAATACGGATGAAAATACTACAGTTTGGAATGATAACACTTTTAAGAAAGTCATATCTAGTAATCTTTCTCCTTCACAGGAAAGAATGTTTAATGTTGGTGATCATGTGAATATTTTTGCTATAGTAAAGTCATATCATGTTGTATGCAAGGAACAATTCAATTATAGTGATGGGGGAATAATAAAAACAAGTGATGTAAAACCAGAAGAAAAAGCAATTTATATTAATATTTTTGGTGAAAAAGAATTACGAACATTAACAGCTAAAGATAAGATTACCTTTAAAAATAATATTGTAACTCTTCAGGAGATTGATGTTAGACTTAGGAAAAGTTTGATGGGGGACAGCAAAATAAAATTGTATGAGTACGATTCTTTGTATAAAAAAGGGTTTTGGGATATTCATTATAAAGACGGTGGCATTAGACACACCAATTTATTTACTTACCCCGACTATACAGATAATGAAACGATTGATATGAGTAAAGTTAGTCACTTTGATGTTCACTTAAACGAAGATTTTTCTAAAGATTAG"
}
}6. Nomenclature Server - At the moment
- What are the Uniprot annotations of the alleles belonging to this loci?
{
"UniprotInfo": [
{
"UniprotLabel": {
"type": "literal",
"value": "Pyrogenic exotoxin SpeK"
},
"UniprotURI": {
"type": "literal",
"value": "http://purl.uniprot.org/uniprot/A0A0E1ESR1"
}
},
{
"UniprotLabel": {
"type": "literal",
"value": "Exotoxin L"
},
"UniprotURI": {
"type": "literal",
"value": "http://purl.uniprot.org/uniprot/Q70EW3"
}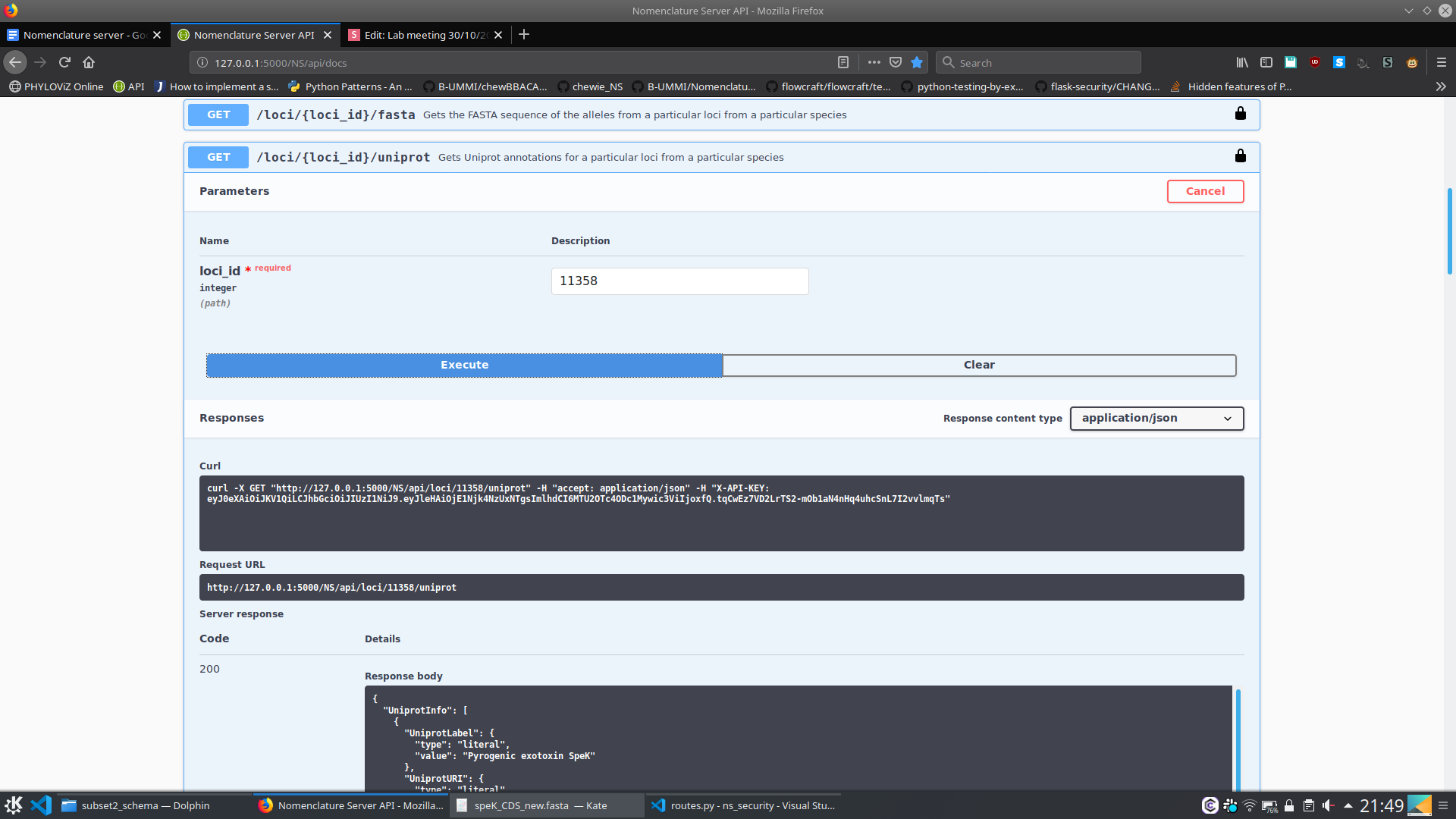
6. Nomenclature Server - At the moment
- What kind of metatada is associated with a particular isolate from this species?
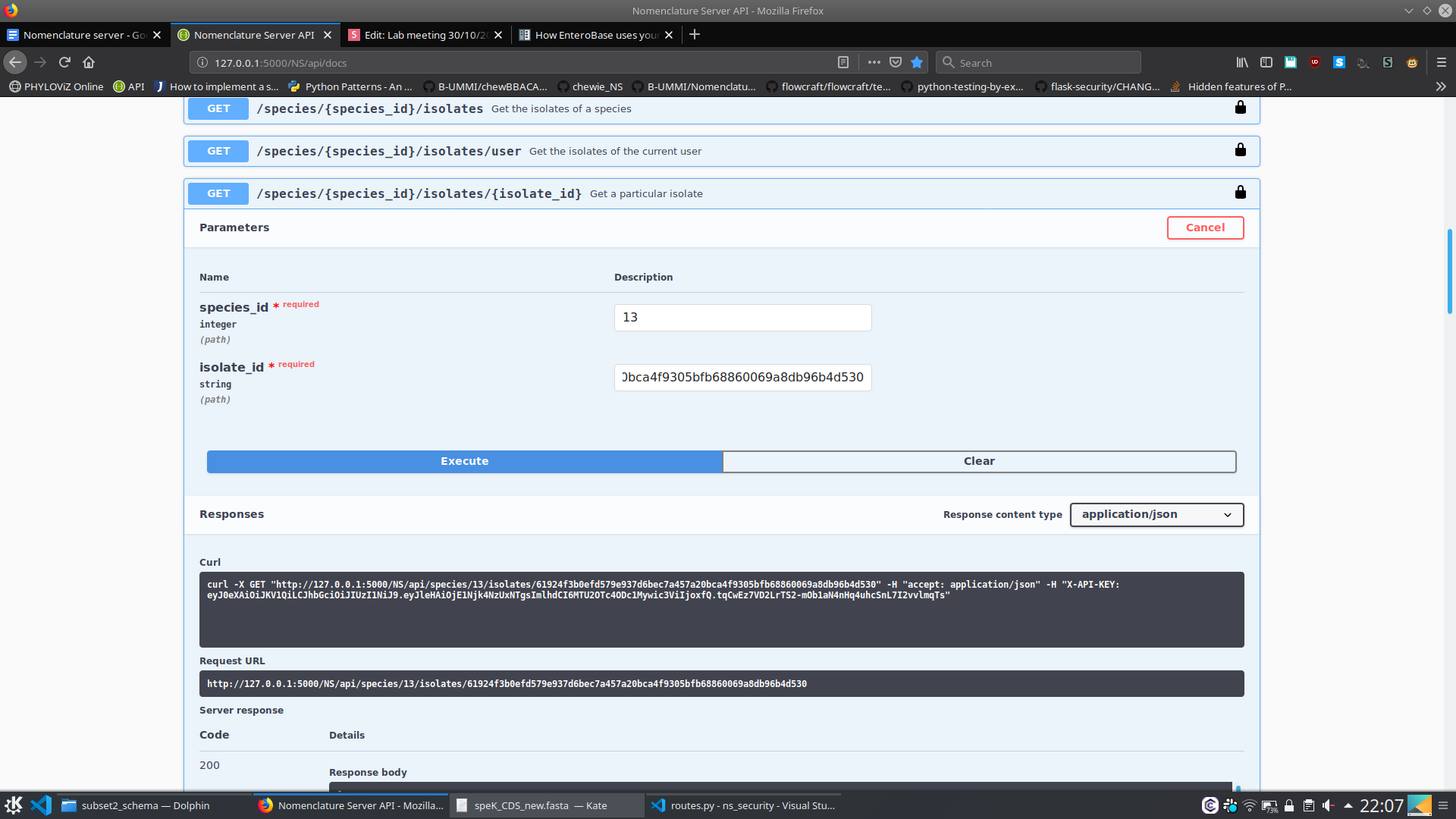
6. Nomenclature Server - At the moment
- What kind of metatada is associated with a particular isolate from this species?
[
{
"name": {
"type": "typed-literal",
"datatype": "http://www.w3.org/2001/XMLSchema#string",
"value": "SH5857.contigs.length_GCcontent_kmerCov.mappingCov.polished.fasta"
},
"country": {
"type": "uri",
"value": "http://dbpedia.org/resource/France"
},
"country_name": {
"type": "literal",
"xml:lang": "en",
"value": "France"
},
"accession": {
"type": "uri",
"value": "https://www.ncbi.nlm.nih.gov/sra/ERR1221234"
},
"st": {
"type": "typed-literal",
"datatype": "http://www.w3.org/2001/XMLSchema#integer",
"value": "1"
},
"date_entered": {
"type": "typed-literal",
"datatype": "http://www.w3.org/2001/XMLSchema#dateTime",
"value": "2019-09-20T15:30:38.801202+01:00"
},
"host": {
"type": "uri",
"value": "http://purl.uniprot.org/taxonomy/9823"
},
"lat": {
"type": "typed-literal",
"datatype": "http://www.w3.org/2001/XMLSchema#long",
"value": "-35.5"
},
"long": {
"type": "typed-literal",
"datatype": "http://www.w3.org/2001/XMLSchema#long",
"value": "36.2222"
},
"isol_source": {
"type": "typed-literal",
"datatype": "http://www.w3.org/2001/XMLSchema#string",
"value": "armpit"
}
}
]6. Nomenclature Server - At the moment
7. Final Remarks
- chewBBACA NS (client) aims to provide the all the necessary functions to download, call alleles and send data to the server in a flexible way.

- Users can work with their data locally, avoiding some concerns over data privacy in sharing data.
- Possibility to synchronize schemas without sending any data to the server
- More wookie!
Lab meeting 31/10/2019
By Pedro Cerqueira
Lab meeting 31/10/2019
chewBBACA NS
- 390



